Abstract
CRISPR advances genome engineering by directing endonuclease sequence specificity with a guide RNA molecule (gRNA). For precisely targeting a gene for modification, each genetic construct requires a unique gRNA. By generating a gRNA against the flippase recognition target (FRT) site, a common genetic element shared by multiple genetic collections, CRISPR-FRT circumvents this design constraint to provide a broad platform for fast, scarless, off-the-shelf genome engineering.
Similar content being viewed by others
Introduction
In E. coli, efficient genome editing using CRISPR and homology-directed repair requires induction of the CRISPR components (Cas9 and gRNA), induction of λ phage recombinase genes1 and a rescue DNA template with the desired mutation. Even though the mechanism of Cas9-based gene editing is still incompletely understood2, the incorporation of a rescue template into the genome by homologous recombination likely prevents Cas9-gRNA from cutting its target sequence, thereby providing a selection wherein engineered clones survive by preventing a futile cycle of lethal double-strand DNA breakage and repair. Each engineered mutation requires a rescue template and unique gRNA that must be designed, cloned and sequence confirmed3,4,5,6. A recently developed CRISPR-based method (CREATE—CRISPR enabled trackable genome engineering7) simplifies and automates the design procedure with an algorithm that incorporates the gRNA and rescue template into a 200 nucleotide oligo that can be directly cloned into a plasmid. However, even with a 70–90% success rate of CREATE, positive clones must be screened by sequencing the modified locus. Here we make CRISPR more accessible and standardized with a simple solution that simultaneously avoids cloning of new gRNAs, circumvents complex design of rescue templates and provides an easy phenotypic screen for positive clones.
We demonstrate a method, CRISPR-FRT (Fig. 1), which directs a gRNA to a FRT (flippase recognition target) sequence present in each knockout mutant of the E. coli Keio collection8. In the arrayed Keio collection of 3884 deletion mutants, each non-essential gene of E. coli has been replaced by a kanamycin-resistance (KanR) cassette flanked on each end by FRT sites7. Instead of designing and cloning a unique gRNA for each gene, a single gRNA-FRT can target any gene that is part of the Keio collection. A Keio strain transformed with plasmids encoding the endonuclease Cas9 and gRNA-FRT experiences a lethal double-strand DNA break at the FRT sites. Since E. coli naturally lacks non-homologous end joining, survival depends upon escaping a futile cycle of homology-directed repair and re-cutting by the Cas9/gRNA-FRT complex. Escape from the Cas9/gRNA-FRT complex can occur by recombination of a homologous rescue DNA template lacking a FRT site. Although any rescue DNA with homology outside the FRT-KanR-FRT cassette should be successful, in this paper we utilize mutated forms of the gene corresponding to the Keio knockout to produce specific (sometimes single nucleotide) changes to genes in their native locus. The mutated gene of interest, along with ~200–500 base pairs of homology flanking the FRT-KanR-FRT cassette, is amplified by PCR and supplied to cells with plasmids encoding gRNA-FRT and Cas9. The λ-red recombinase system1 is also expressed in these cells to promote recombination and avoid degradation of the rescue DNA template. Recombination of the rescue DNA template results in replacement of the FRT-flanked kanamycin-resistance cassette by the gene containing the mutation of interest. Consequently, this protocol allows one to easily introduce specific mutations in the ancestral Keio background (BW25113), without any scars. Other E. coli strains can be engineered by first transferring the KanR cassette using P1vir transduction and then proceeding with the CRISPR-FRT protocol. Likewise, multiple mutations can be constructed in the same strain by consecutive cycles of P1vir transduction of a new gene deletion from the Keio collection followed by another round of CRISPR-FRT. Alternatively, the λ-red recombinase genes encoded on the pKDsgRNA-FRT4 or the pCas3 plasmid can be used to precisely replace the new target gene by a FRT-flanked KanR cassette. In this approach, a PCR-amplified oligo from the appropriate Keio clone is used as a template for homologous recombination in the mutant background. Either approach allows for consecutive rounds of mutant construction.
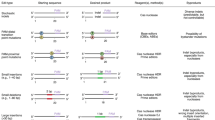
Overview of the CRISPR-FRT protocol. CRISPR-FRT makes use of the arrayed collection of Keio knockout mutants having a FRT-flanked kanamycin-resistance (KanR) cassette replacing each non-essential E. coli gene. CRISPR-FRT includes a gRNA-FRT that directs the Cas9 nuclease to bind and cut the two FRT sites. A convenient rescue template (e.g., a mutated gene from an evolved E. coli strain amplified by PCR, a plasmid-encoded gene variant, etc) recombines (dashed lines) over the homologous regions flanking the KanR cassette. Survivors are screened to separate KanR false positives (red X) from the kanamycin sensitive (KanS) true positives (green check) that replaced KanR cassette with the mutated gene
Results
Convenience of CRISPR-FRT
CRISPR-FRT (Fig. 1), directs a gRNA to a FRT (flippase recognition target) sequence present in each knockout mutant of the E. coli Keio collection8 (Supplementary Fig. 1). There are several advantages of CRISPR-FRT. Using a single gRNA to target any non-essential E. coli gene obviates the need to design, clone and sequence confirm individual gRNAs for each gene or each mutation in the same gene. A static set of Cas9 and gRNA delivery plasmids can be used for each desired mutant. Also, the rescue DNA template can be easily designed. Normally, mismatches, within or adjacent to the gRNA target site are required to avoid cutting. CRISPR-FRT circumvents this aspect of rescue DNA template design by removing all traces of the FRT sites targeted by the gRNA. With this system, constructing a mutation only requires designing a set of primers to amplify the mutated gene and the corresponding Keio knockout strain transformed with the CRISPR-FRT plasmids. Furthermore, successful incorporation of the rescue DNA template results in the replacement of the KanR cassette, thereby providing a kanamycin sensitive phenotype that can be screened. For CRISPR experiments utilizing plasmids already possessing a KanR gene, expression of flippase can remove the KanR cassette and leave a single FRT scar that is to be targeted by CRISPR-FRT (Supplementary Fig. 2). Without the KanR cassette resistance phenotype, screening for true positive clones requires colony PCR. In either condition thus far, all clones producing the appropriately sized PCR DNA fragment have been confirmed by sequencing to have undergone homology-directed repair with the rescue DNA.
Single-nucleotide editing with CRISPR-FRT
To assess robustness and reproducibility, we cloned a gRNA targeting FRT into two independent CRISPR plasmid systems: the pTarget series3 and the noSCAR system4. CRISPR-FRT was used to reconstruct mutations observed in previously conducted evolution experiments to higher ethanol tolerance9, 10, higher persistence11, ciprofloxacin resistance and colistin resistance in E. coli. In each case, rescue DNA was generated using primers designed to amplify the mutated gene with homology of 220 base pairs or more on each side of the KanR cassette (Fig. 1). The amplified product was recombined into its native locus in either the Keio background (E. coli BW25113) or E. coli MG1655 transduced with the FRT-flanked KanR cassette from the appropriate Keio knockout clone. The implementation of CRISPR-FRT in the pTarget system3 first required flippase-mediated removal of the KanR cassette to leave a single FRT target followed by introduction of the pTarget plasmids and rescue template. Clones that recombined the rescue template were then identified by colony PCR (Table 1, acrA → yigI). In contrast, the noSCAR4 system plasmids were compatible with the FRT-KanR-FRT cassette and positive clones were identified by screening for kanamycin sensitivity (Table 1, basR → mutL). We also used CRISPR-FRT to separate several acrR mutants present as subpopulations within a culture of E. coli adapted for ciprofloxacin resistance (Fig. 2a). From a single CRISPR-FRT reaction, we recovered four unique acrR single substitution mutants (M1I, V29frame-shift, K55E and W63R). Finally, in order to introduce mutations to two genes, we used consecutive cycles of KanR cassette P1vir transduction and CRISPR-FRT of basR G53E or basS C84R into either yejM P126S or yejM R165C single mutants, respectively. Overall, we found that CRISPR-FRT functions well in either of the two CRISPR platforms tested and is adaptable to the production of multiple mutations within or between separate genes. Moreover, CRISPR-FRT proved successful at three independent labs, demonstrating the robustness and reproducibility and making it particularly well-suited for both research and educational purposes.
Extended applications of CRISPR-FRT. a Members of a plasmid-encoded mutagenesis library (1–4…n) or PCR-amplified linear fragments containing multiple mutated versions of one gene can serve as rescue DNA and thereby be transferred to the chromosome. This allows for single-step transfer of a mutagenesis library to the genome or reconstruction of multiple mutations (red X) in the same gene by only using a single pair of PCR primers to generate the different rescuing templates that contain the desired mutations. b Mutations in essential genes can be delivered to Keio strains with KanR cassettes inserted into the closest neighboring non-essential gene if recombination (dashed lines) occurs beyond the mutation
In order to provide a useful demonstration of using CRISPR-FRT, we assayed several of the constructed mutants for their impact on phenotype (Fig. 3). First, the basR and basS mutants showed significantly increased colistin minimal inhibitory concentration (MIC) values and all acrR mutants showed significantly increased ciprofloxacin MIC values, demonstrating their effect on colistin and ciprofloxacin resistance, respectively. As expected, the mutations in nadC and vacJ which were identified as founder mutations already present in the non-resistant ancestral strain, did not result in higher ciprofloxacin MIC compared to the MIC for the wild type. Second, we showed that the single oppB mutation, which was previously identified in an evolution experiment to higher persistence10, confers significantly higher persister levels both when treated with amikacin or ciprofloxacin, suggesting a putative role for this ATP-dependent oligopeptide uptake system in persistence. Next, we tested the mutation rate of mutants harboring single mutations in DNA replication and repair genes mutL, mutH, uvrD and mfd. While all mutations caused the mutation rate to increase, only the mutation rate of mutL (S101R), mutL (H270R), mutH (W106R) and mfd (V864A) was significantly increased compared to the wild type. Finally, a previously identified mutation in the envZ gene was assayed for ethanol tolerance9. Both the growth rate and final cell density increased in the envZ mutant compared to the wild type when exposed to a near-lethal concentration of 5% (v/v) ethanol. In general, these assays demonstrate that CRISPR-FRT allows rapid testing of different phenotypes by enabling rapid and easy introduction of multiple, separate mutations (Fig. 3).
CRISPR-FRT allows rapid reconstruction and phenotypic characterization of generated mutants. a Mean colistin minimum inhibitory concentrations (MIC) for wild-type MG1655 and otherwise isogenic basR and basS mutants (n = 3; error bars represent the s.d.; two-sample unpaired t-test; ***p < 0.001 vs. wild type). b Mean ciprofloxacin minimum inhibitory concentrations (MIC) for wild-type BW25113 and otherwise isogenic nadC, vacJ and acrR mutants (n ≥ 3; error bars represent standard error of the mean; two-sample unpaired t-test **p < 0.01, ***p < 0.001 vs. wild type). c When treated with amikacin or ciprofloxacin the oppB (A180E) mutant shows a significantly increased surviving persister fraction compared to the wild type (n = 3; error bars represent the s.d.; two-sided t-test; ***p < 0.001). d Mutations in mutL, mutH, uvrD and mfd can change the genomic mutation rate. While all mutants show a higher mutation rate, only the mutL (S101R), mutL (H270R), mutH (W106R) and mfd (V864A) mutants show a significantly higher mutation rate compared to the wild type (n = 24; error bars represent upper and lower limits of the 95% confidence intervals; ShinyFlan R package built-in two-sample comparison [30]; *p < 0.05; ***p < 0.001). e The envZ (L116P) mutant exhibits enhanced growth characteristics in the presence of 5% (v/v) ethanol compared to the wild type (left panel). Both the maximal final density (right top panel) and the growth rate (right bottom panel) significantly improved compared to the wild type (n = 5; error bars represent s.d.; unpaired two-sided t-test; ***p < 0.001). n.s. non-significant
Targeting essential genes
CRISPR-FRT can also be used to construct mutations in essential genes that are not part of the Keio collection. For this, the Keio knockout that is closest to the mutation in the essential gene is targeted by the gRNA-FRT. The majority of essential genes (80%) have a directly adjacent non-essential gene available in the Keio collection (Fig. 4). The rescue DNA is then extended to include homology that reaches beyond the mutation in the neighboring essential gene (Fig. 2b). Successful engineering of the essential gene thereby depends upon the homologous recombination cross-over point occurring outside the mutation in the rescue DNA. To test this feature, we used Keio knockouts of non-essential genes (rmf or yejL) to reconstruct a point mutation in neighboring, essential genes, fabA or yejM (Table 1). In each case, the wild-type non-essential gene was properly delivered while the desired mutation in the essential genes occurred in 40–44% of sequenced colonies. One of these mutations, V367A in yejM, is 1120 nucleotides from the yejL KanR cassette, indicating that even relatively distant mutations can be constructed with this method. The 40–44% efficiency could be influenced by genetic distance between the mutation and the FRT site or, in specific cases, deleterious effects of mutations in the targeted essential genes. Although lower in efficiency than directly targeting a non-essential Keio knockout, the ability of CRISPR-FRT to create mutations in essential genes greatly expands its utility.
Transferring a plasmid library to the chromosome
In addition to being a versatile and efficient way to rapidly reconstruct single point mutations, our CRISPR-FRT method can also generate larger libraries of these point mutants in a gene’s native locus (Fig. 2a). Normally, site-directed mutagenesis libraries are constructed on plasmids that can have copy number effects and require antibiotic supplementation; an important consideration for commercial scaling of engineered strains. As a proof of principle, we transferred an existing pool of 68 recA mutants from a plasmid to the recA chromosomal locus. The existing KanR recA plasmid library was transformed into the Keio recA knockout mutant that had been rendered kanamycin sensitive by expression of flippase and removal of the chromosomal KanR cassette. The cells containing the library were then transformed with CRISPR-FRT plasmids and 100 out of 350,000 survivors were screened by colony PCR (Table 1). The 77 clones yielding DNA of the appropriate size were sequenced and found to have delivered the recA gene from the plasmid library. The plasmid-encoded recA library was collected just prior to transforming the CRISPR-FRT plasmids and sequenced to determine the library’s initial sequence diversity (Fig. 5). We found no difference in diversity between the plasmid and chromosomal libraries (Multinomial goodness-of-fit test by Monte-Carlo simulation, p value ± s.d. = 0.504 ± 0.002). The similar distribution of recA mutants between the libraries and the large number of successful gene conversion events suggests all members of the small library were delivered to the chromosome. These results suggest other existing plasmid-based libraries could likewise be transferred to the chromosome if a homologous DNA sequence between the plasmid and target DNA locus is present.
The distribution of recA alleles encoded by the plasmid pool before CRISPR-FRT is similar to the distribution of recA mutants transferred to the chromosome after CRISPR-FRT. a Column scatter graph showing the distribution of recA mutants encoded by the plasmid pool before CRISPR-FRT and recA mutants on the chromosome after CRISPR-FRT. Mean proportion of mutants shown with heavy bars and standard deviation shown with light bars (n = 31). b Distribution of individual recA mutants before and after CRISPR-FRT. Plotting previously reported12 relative recombination function of recA mutants (circles, second y-axis) suggests deleterious mutants with <1% recombination activity are underrepresented in the original plasmid library pool
Engineering a protein tag into an F plasmid gene
Finally, CRISPR-FRT can be used to engineer protein tags onto genes in their native locus. As a proof of principle, we added a FLAG-tag to the traT gene in an F plasmid derivative. As no equivalent of a Keio collection exists for the F plasmid, we first replaced the traT gene with the FRT-KanR-FRT cassette using an E. coli strain encoding λ phage recombineering genes on the chromosome13. Having created an F plasmid derivative with ΔtraT::FRT-KanR-FRT, we next supplied the CRISPR-FRT/noSCAR plasmid set along with a traT-FLAG rescue DNA generated via a PCR reaction using primers designed to append the FLAG tag to the 3’ end of the traT gene. In contrast to editing the genome, a double-stranded DNA break caused by CRISPR-mediated cleavage of the F plasmid does not provide a direct selection for engineered clones because cells can also eliminate the target FRT sites through plasmid loss. We find plasmid loss predominates (92 of 99 clones screened) relative to the number of successful recombination events (traT-FLAG, 5 of 99 clones) or escape from CRISPR-FRT cutting (ΔtraT::KanR, 1 of 99). Inclusion of tetracycline in the CRISPR-FRT agar plates reduced loss of the tetracycline-resistant F plasmid derivative and improved recovery of traT-FLAG clones (Table 1). Although less efficient than making chromosomal point mutants, CRISPR-FRT is able to deliver FLAG-tagged traT to its native locus on a large plasmid. This shows the flexibility of CRISPR-FRT as a platform to engineer genes in their native locus beyond making point mutants on the chromosome.
Discussion
CRISPR-FRT targets a single sequence that is found at a unique genetic locus for each member of a genetic library to allow for fast, robust, and scar-less genomic engineering in E. coli. It eliminates both the cloning of new gRNAs and the design of specific rescue DNA constructs for each desired mutation. When using current CRISPR methods, attempts to modify genes with custom gRNAs often result in unintended mutations occurring in or near the gRNA-binding site. We found that a rescue template with the desired mutation and multiple synonymous mutations in or near the gRNA-binding site could avoid this undesired targeting of the rescue DNA. However, previous work has shown that even synonymous mutations can have phenotypic effects on their own in some cases14, 15. In contrast to the current methods, CRISPR-FRT permits precise single-nucleotide editing, anywhere in the gene, and avoids this specific type of off-target effect by complete ablation of the FRT target site.
This approach not only provides a ready-to-use method for reconstruction of mutations in the E. coli Keio collection, but is applicable to other collections with insertion sequences that can be targeted using a similar strategy. Genome library collections of 94 intergenic insertions16 and 140 sRNA/small protein knockouts17 in E. coli and 1,052 Salmonella enterica gene knockouts18 also employ a FRT-flanked KanR cassette that can be directly used with the CRISPR-FRT system. Gene knockout collections containing FRT sites are also available for other bacteria, including Acinetobacter baylyi19 and Burkholderia thailandensis20. Collections of gene replacements or transposon insertions, some with FRT sequences, also exist in higher organisms including Saccharomyces cerevisiae21, Drosophila melanogaster22, Homo sapiens cell lines and many more23. However, in organisms where non-homologous end joining strongly dominates homology-directed repair, effectiveness of CRISPR-FRT will be compromised until methods are developed to block NHEJ24, 25. In general, the concept of using CRISPR to target a shared sequence among insertion elements can be applied to any organism with an arrayed library and the ability to deliver a functional CRISPR system.
Methods
Bacterial strains and culture conditions
To reconstruct adaptive mutations with CRISPR-FRT, we used the knockout mutants from the Keio collection derived from E. coli BW251138 or E. coli MG1655 that had been transduced with the FRT-KanR-FRT cassette from a member of the Keio collection. These strains and all other strains and plasmids are listed in Supplementary Data 1. When the target gene was essential and no KO mutant was available, we selected the closest neighboring gene to target with CRISPR-FRT. All strains were grown in lysogeny broth (LB) medium in an orbital shaker at 200 r.p.m. and 30–32 °C or 37 °C, depending on the sensitivity of the CRISPR delivery plasmids. The HME45 strains harboring pOX38-Tc plasmid variants were grown at 30 °C.
Plasmid and strain construction
We obtained two plasmid sets to test the CRISPR-FRT method: the pTarget system3 and the noSCAR system4. The protospacer is a 20 nt DNA stretch that defines the specificity and directs Cas9 to the position of interest. Cas9 cleaves the DNA near a PAM (-NGG) site in the DNA. Therefore, protospacers need to be designed as such that they flank a PAM site. The 48 nt-long FRT site has 4 possible PAM sites. We first used the ATUM gRNA design tool for E. coli K12 MG1655 (online available from https://www.atum.bio/eCommerce/cas9/input) to determine the best PAM-site with the least chance for off-target cleavage (Supplementary Fig. 1). This 20 nt sequence was used as overhangs in primers to amplify the pKDsgRNA gRNA delivery plasmid. Next, we used ligation-independent cloning or Gibson assembly (New England Biolabs, NEB) to include the FRT protospacer sequence into the pKDsgRNA gRNA delivery plasmid. This created the plasmid pKDsgRNA-FRT. For cloning the FRT-gRNA into pTargetF, we performed an inverse-PCR type reaction applied to the whole plasmid3 using oligonucleotide primers gRNA:FRT and gRNA:LexA-G85-2 (Supplementary Data 2). However, in the latter case the resulting PCR product was cut with SpeI (NEB), the ends self-ligated with T4 DNA ligase (New England Biolabs), phenol:chloroform extracted, ethanol precipitated and transformed into DH5α cells. This created pTargetF-FRT. Due to kanamycin resistance on pCas, each Keio strain used with the pTargetF-FRT/pCas was first rendered kanamycin sensitive by flipping out the KanR cassette and leaving a single FRT scar. This was achieved using flippase expressed from pCP201. The genes targeted with the pTargetF-FRT/pCas based system included the following: acrA, acrR, deoC, malZ, nadC, vacJ, ygiD, ygiV, yigI and recA. For delivery of the pool of 68 recA mutants plus wild-type from a kanamycin-resistant F plasmid derivative12, the kanamycin-resistance gene on pCas3 was replaced by the chloramphenicol gene by recombineering. DH5α[pCas] cells were grown overnight, induced with arabinose for 3 h to express λ Red genes from pCas, made electro-competent and transformed at 1.4 kV, 25 μF, 200 Ω with overlap PCR product of the chloramphenicol gene having homology to the origin of replication, oriR101, and the tracrRNA region of the Cas9 gene. Template for the overlap PCR included pCas, for oriR101 sequence, and pACYC184-lexA26, for the chloramphenicol gene. Primers used in the overlap PCR reactions included repA101-1, ori101-CmR-2, ori101-CmR-2-complement and CmR-tracr-2-complement. Surviving clones on chloramphenicol (12.5 μg/ml) were mini-prepped, re-transformed into DH5α cells and then plated onto chloramphenicol plates again to remove unmodified pCas. The resulting plasmid is named pCas-CmR(+). Construction of the pool of 68 recA mutants on the pGE591 plasmid was facilitated by the QuikChange II XL Site-Directed Mutagenesis Kit (Agilent) and the primers listed in Supplementary Data 2. The yehL S20P rescue template originates from a genomic DNA library constructed by ligating partially Sau3AI digested DNA into BamHI digested pTargetF-FRT.
Construction of the rescue oligo
The rescue oligos needed to repair the double-stranded DNA breaks caused by Cas9 cleavage were constructed using PCR amplification. As a template, we used adapted strains resulting from previously run evolution experiment that harbored the specific mutation of interest9,10,11. For each mutation, we designed a primer pair that targeted a region including the entire mutated gene and 220 bp or more homology overhangs on each side (Fig. 1) (Supplementary Data 2). For the PCR reaction, we used either high-fidelity Universe polymerase (Bimake), high-fidelity Q5 polymerase (NEB) or phusion polymerase (NEB) to limit the risk for additional DNA changes during amplification. The resulting rescue oligo was purified using the DNA Clean & Concentrator™-5 (Zymo Research) and eluted in 8 µl elution buffer to obtain a highly concentrated rescue oligo (100–1000 ng/µl) for subsequent DNA transformation. For constructing yehL and recA mutants, the rescuing template was actually plasmid encoded. For the traT-FLAG rescue template, 499 bp upstream of the traT gene and the traT open reading frame was amplified from pOX38-Tc plasmid DNA using oligonucleotide primers orb194 and orb195. The primers were designed to incorporate a 1 × FLAG tag on the 3’ end of the traT gene. 501 base pairs downstream of the traT gene were amplified from pOX38-Tc using oligonucleotide primers orb196 and orb197. The upstream and downstream PCR products were joined by fusion PCR using oligonucleotide primers orb194 and orb197. For each PCR product rescue template, the appropriately sized product was gel purified (Qiagen or Zymogen).
CRISPR-FRT
To reconstruct mutations from previously performed evolution experiments, we adapted two earlier described protocols3, 4 and included the common FRT targeting protospacer in its gRNA delivery plasmid. To apply CRISPR-FRT in the two CRISPR platforms we followed instructions as previously described3, 4. In brief, plasmids carrying the cas9 gene, FRT-gRNA and λ-red recombinase genes are separately transformed to the cell. Next, the recombinase genes and CRISPR genes are induced while the rescue oligo, containing the mutated gene and homologous flanking sequences, is transformed into the cell. The Cas9 nuclease will make dsDNA breaks at the FRT sites and cells only survive when the mutated genes recombined into the genome to remove the FRT sites. Resulting colonies were tested by colony PCR or streaking on a non-selective agar plate and an agar plate containing kanamycin (40 µg/ml). Colonies that showed growth on kanamycin were false positive. In most cases, the kanamycin sensitive colonies were true positives. Finally, a subset of sensitive colonies was sequenced (GATC Biotech, Germany; Genewiz, Plainfield, NJ, USA; or Baylor College of Medicine sequencing core facility, Houston, TX, USA) to confirm correct reconstruction of the adaptive mutation.
To reconstruct multiple mutations in the same background (e.g., fabF:acrB and fabA:fabR), we performed consecutive rounds of CRISPR-FRT. In case of fabF:acrB, after construction of the fabF mutation, we introduced a new FRT-flanked KanR cassette to replace the native acrB gene. To this end, we PCR amplified the region containing the FRT-flanked KanR cassette and overlapping ends on both sides. Next, the mutant harboring the first mutation was induced with arabinose (0.2%) to express the λ-red recombinase genes encoded on the pKDsgRNA-FRT3 or the pCas92 plasmid. After an induction period of approximately 3 h, the PCR-amplified oligo was electroporated to the induced cells. After 1 h of incubation, transformed cells were plated on Kan plates to select for positive clones. Proper introduction of the cassette was verified by colony PCR. The positive clones were then used for a consecutive round of CRISPR-FRT to introduce the second mutation. Both Cas9 and the sgRNA delivery plasmid can be retained in the cell while performing consecutive rounds of CRISPR-FRT, shortening the time to construct multiple mutants in the same background.
Adjacency of essential genes to non-essential genes
CRISPR-FRT of essential genes requires the presence of a nearby Keio knockout of a non-essential gene. For the purposes of the method presented here, we define essentially as the inability to generate a clean knock-out in the Keio collection8 without a duplication event occurring27. In order to determine how many essential genes lie adjacent to a non-essential Keio knockout clone, we first determined how essential genes are grouped on the E. coli chromosome (as singles, doubles, triples, etc.) and how many genes belonged to each class. With this information, the number of genes occurring between an essential gene and a non-essential gene was determined using a Pascal’s triangle.
CRISPR-FRT of recA library
The small library of plasmid-encoded recA mutants was transferred to the chromosome using pTarget-FRT and pCas-CmR(+). To permit selection for kanamycin-resistant transformants of recA library plasmids, the kanamycin-resistance cassette was first removed from ∆recA::FRT-KanR-FRT Keio strain JW2669 by expression of flippase from the pCP20 and simultaneous curing of the temperature-sensitive pCP20 plasmid1. This generated strain DCM207 harboring the ΔrecA::FRT scar. The small recA library was then transferred as a pool into DCM207. Transformants were selected on kanamycin plates, pooled, made electro-competent, transformed with pCas-CmR( + ) and again selected on plates containing both chloramphenicol and kanamycin at 32 °C. The resulting colonies were again pooled, induced with arabinose to express λ-red recombinase genes from pCas-CmR( + ), made electro-competent, transformed with pTarget-FRT and selected on plates containing both spectinomycin and chloramphenicol. In order to cure pTarget-FRT from these cells, the resulting colonies were pooled, diluted and plated on chloramphenicol plates containing 1 mM IPTG3. Individual clones were screened for replacement of the ΔrecA::FRT scar with a recA gene from the plasmid pool by colony PCR (primers mltB-1 and alaS-1, Supplementary Data 2). In order to avoid amplification of the recA locus from the plasmid, one primer, mltB-1, was designed to specifically bind the chromosome; upstream from the plasmid’s cloning junction. The downstream primer, alaS-1, binds downstream of the recA gene encoded on either the plasmid or chromosome. Although not implemented here, curing of the plasmid library could be performed by directing a gRNA against a unique site in the recA plasmid. A new pTarget plasmid encoding an sgRNA targeting a unique sequence on the recA plasmid could be transformed into the chromosomal recA library pool and plated on chloramphenicol + spectinomycin plates to select for pCas-CmR(+)/pTarget and cure the recA plasmid pool. Surviving colonies would then be pooled and again plated on IPTG + chloramphenicol to cure the new pTarget. Finally, the surviving clones would be pooled, plated onto plates without antibiotic, and incubated at 42 °C to cure pCas-CmR(+).
To determine whether the recA chromosomal library reflects the diversity of the plasmid library, we performed sequencing before and after conducting CRISPR-FRT. Colony PCR products from clones surviving the CRISPR reaction (see above) were Sanger sequenced (Baylor College of Medicine sequencing core) and aligned to the recA gene using SnapGene v4.1.5. In order to determine the distribution of mutants in the recA plasmid library pool just prior to CRISPR-FRT, DCM207 [pCas-CmR(+), pGE591 library pool] cells were grown overnight, plasmids were isolated by mini-prep (Qiagen) and then used as template in a PCR reaction using primers RecA-FP and alaS-1. The PCR product was then prepared for deep sequencing using a Nextera XT DNA sample preparation kit and an Illumina Miseq v3, 600 cycle sequencing cartridge. The data were aligned to the recA sequence and SNPs were reported with their frequency using breseq software28 with a cut off of 0.1%. A multinomial goodness-of-fit test by Monte–Carlo Simulation was performed to compare the mutation distribution in the genome with that in the plasmid using the XNomial package in R. A p value of 0.504 ± 0.002 was reported, suggesting that the mutations in the genome sample were a good representation of the plasmid sample.
For CRISPR-FRT of the traT gene on the F plasmid derivative, pOX38-Tc (gift of Laura Frost)29, the traT gene was replaced with an FRT-KanR-FRT cassette from pKD13 using recombineering in E. coli strain HME45. E. coli HME45 has a defective λ prophage that retains the gam-bet-exo recombination genes (and others) under control of a temperature-sensitive cI repressor13. Oligonucleotide primers orb177 and orb178 were used to amplify the FRT-KanR-FRT cassette from pKD13 and provide homology for recombineering to generate pOX38-Tc-∆traT::FRT-KanR-FRT. The CRISPR-FRT plasmids were introduced into HME45 [pOX38-Tc-∆traT::FRT-KanR-FRT] and CRISPR was performed as described above with a few exceptions. Cells were grown to an OD600 of 0.4–0.6 and then the λ-red recombinase genes encoded in the HME45 chromosome were induced by shaking in a 42 °C water bath for 15 min. As temperature induction could result in loss of the temperature-sensitive pKDsgRNA-FRT plasmid, both the traT-FLAG rescue DNA (810 ng) and the pKDsgRNA-FRT plasmid (370 ng) were introduced by electroporation. Cells were selected on LB agar plates supplemented with anhydrotetracycline (1000 ng/µL, Sigma), chloramphenicol, and spectinomycin and screened for plasmid retention and loss of kanamycin resistance as described above. In order to provide a selection against plasmid loss events, CRISPR-FRT was repeated as above but with the inclusion of tetracycline along with chloramphenicol, spectinomycin and anhydrotetracycline in the CRISPR/recombineering plate. KanS colonies were screened for traT-FLAG using oligonucleotide primers orb179 and orb180. Three representative PCR products were verified by DNA sequencing (Macrogen, USA) with oligonucleotide primers orb179 and/or orb180.
MIC determination
Mutations originating from ciprofloxacin and colistin adaptive laboratory evolution experiments were tested for conferring resistance to their respective selective antibiotics by using Etest strips (bioMérieux). Overnight cultures inoculated from an isolated colony (at least three biological replicates) were diluted 100-fold into fresh lysogeny broth (LB) and allowed to grow for 2 h before being transferred to an LB plate with a sterile swab. Once the plates were dry, a single Etest strip was applied and the plate was incubated overnight at 37 °C.
Persistence assay
We determined the persistence fraction of wild type and oppB mutants in triplicate. All strains were grown for 24 h in liquid Mueller Hinton Broth (MHB) medium in an orbital shaker at 200 rpm and 37 °C. First, these cultures were diluted 100-fold in 100 mL MHB containing flasks and incubated for 16 h in an orbital shaker at 200 rpm and 37 °C. Next, the initial cell number in each flask was determined by making a dilution series and plating the 10−6 dilution on solid LB agar plates (control plates). To determine the level of persisters, 1 mL of each culture was treated with ciprofloxacin (5 µg/mL) or amikacin (400 µg/mL) for 5 h in an orbital shaker at 37 °C and 200 r.p.m. After the antibiotic treatment, samples were centrifuged for 5 min at 4.032 × g and the pellets were resuspended in 10 mM MgSO4 to wash away the antibiotic. A 10-fold dilution series was generated starting from these resuspended cultures and dilution 10−2 and 10−4 were plated on solid LB agar plates to determine the number of surviving persister cells. Finally, the number of persister cells and the total number of cells (control plates) were used to determine the persister fraction for each tested strain11. Since log-transformed persister fractions are normally distributed statistical significance between the log-transformed persister fractions of wild-type and mutant cells was determined using an unpaired two-sided Student’s t-test with unequal variances (based on an F-test).
Fluctuation analysis
We determined the genomic mutation rate of selected mutant and wild-type strains by using the Luria-Delbruck fluctuation assay9. In brief, overnight cultures of the strains were grown until mid-exponential phase and diluted in LB-medium to a density of 5000 cells per mL. Next, these diluted cultures were divided in at least 30 replicate cultures of 200 µL each in a 96-well plate or Eppendorf tubes and grown individually for 24 h in an orbital shaker at 200 rpm and 37 °C. After 24 h, at least 4 replicate cultures were used to determine the total cell count for each individual culture by making a 10-fold dilution series and plating on solid LB agar plates. The remaining individual cultures were entirely plated on solid LB agar plates supplemented with 100 µg/mL rifampicin to determine the number of spontaneous resistant mutants. Acquiring rifampicin resistance occurs through mutations in the rpoB gene. Therefore, the number of rifampicin-resistant colonies is directly related to the frequency of mutations occurring in rpoB and hence the mutation rate of the strain. The data were analyzed by using flan, a recently developed R package for inference of mutation models. The software uses the number of resistant colonies in multiple individual cultures and the average number of total cells per culture to estimate the mutation rate with the Maximum Likelihood method30. The mutation rates of two samples were statistically compared by using the built-in two-sample test.
Growth analysis
To quantify the ethanol tolerance level of wild-type and mutant strains growth dynamics were monitored in the presence of 5% (v/v) ethanol. Overnight cultures were diluted 100-fold in flasks containing 50 mL LB supplemented with 5% (v/v) ethanol. The flasks were closed with rubber sealed caps to prevent ethanol evaporation. Growth of each strain was monitored in 5-fold by measuring optical density (OD595nm) at various time points during growth. The resulting growth curves were fitted using the widely accepted Gompertz equation31. This fitting allowed for determination of growth rate, lag time and maximal density of wild-type and mutant strains in the presence of ethanol. Statistical significance of the difference in growth rate and maximal density was calculated using an unpaired two-sided Student’s t-test with equal variances (based on an F-test).
Data availability
All data and plasmids are available upon request from the authors.
References
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. USA 97, 6640–6645 (2000).
Cui, L. & Bikard, D. Consequences of Cas9 cleavage in the chromosome of Escherichia coli. Nucl. Acids Res. 44, 4243–4251 (2016).
Jiang, Y. et al. Multigene editing in the Escherichia coli genome via the CRISPR-Cas9 system. Appl. Environ. Microbiol. 81, 2506–2514 (2015).
Reisch, C. R. & Prather, K. L. The no-SCAR (scarless Cas9 assisted recombineering) system for genome editing in Escherichia coli. Sci. Rep. 5, 15096 (2015).
Jiang, W., Bikard, D., Cox, D., Zhang, F. & Marraffini, L. A. RNA-guided editing of bacterial genomes using CRISPR-Cas systems. Nat. Biotech. 31, 233–239 (2013).
Zhao, D. et al. Development of a fast and easy method for Escherichia coli genome editing with CRISPR/Cas9. Microb. Cell. Fact. 15, 205 (2016).
Garst, A. D. et al. Genome-wide mapping of mutations at single-nucleotide resolution for protein, metabolic and genome engineering. Nat. Biotech. 35, 48–55 (2016).
Baba, T. et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol. Syst. Biol. 2, 2006.0008 (2006).
Swings, T. et al. Adaptive tuning of mutation rates allows fast response to lethal stress in Escherichia coli. eLife 6, e22939 (2017).
Swings, T. et al. Network-based identification of adaptive pathways in evolved ethanol-tolerant bacterial populations. Mol. Biol. Evol. 34, 2927–2943 (2017).
Van den Bergh, B. et al. Frequency of antibiotic application drives rapid evolutionary adaptation of Escherichia coli persistence. Nat. Microbiol. 1, 16020 (2016).
Adikesavan, A. et al. Separation of recombination and SOS response in Escherichia coli RecA suggests LexA interaction sites. PLoS Genet. 7, e1002244 (2011).
Yu, D. G. et al. An efficient recombination system for chromosome engineering in Escherichia coli. Proc. Natl Acad. Sci. USA 97, 5978–5983 (2000).
Spencer, P. S. et al. Silent substitutions predictably alter translation elongation rates and protein folding efficiencies. J. Mol. Biol. 422, 328–335 (2012).
Agashe, D. et al. Good codons, bad transcript: large reductions in gene expression and fitness arising from synonymous mutations in a key enzyme. Mol. Biol. Evol. 33, 1542–1553 (2016).
Nehring, R. B. et al. An ultra-dense library resource for rapid deconvolution of mutations that cause phenotypes in Escherichia coli. Nucl. Acids Res. 44, e41 (2016).
Hobbs, E. C., Astarita, J. L. & Storz, G. Small RNAs and small proteins involved in resistance to cell envelope stress and acid shock in Escherichia coli: analysis of a bar-coded mutant collection. J. Bacteriol. 192, 59–67 (2010).
Porwollik, S. et al. Defined single-gene and multi-gene deletion mutant collections in Salmonella enterica sv Typhimurium. PLoS ONE 9L, e99820 (2014).
De Berardinis, V. et al. A complete collection of single-gene deletion mutants of Acinetobacter baylyi ADP1. Mol. Syst. Biol. 4, 174 (2008).
Gallagher, L. et al. Sequence-defined transposon mutant library of Burkholderia thailandensis. mBio 4, e00604–e00613 (2013).
Winzeler, E. et al. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 285, 901–906 (1999).
Venken, K. J. T. & Bellen, H. J. Chemical mutagens, transposons, and transgenes to interrogate gene function in Drosophila melanogaster. Methods 68, 15–28 (2014).
Yusa, K. piggyBac transposon. Microbiol. Spectr. 3, MDNA3–0028–2014 (2015).
Maruyama, T. et al. Increasing the efficiency of precise genome editing with CRISPR-Cas9 by inhibition of nonhomologous end joining. Nat. Biotech. 33, 538–542 (2015).
Bernheim, A. et al. Inhibition of NHEJ repair by type II-A CRISPR-Cas systems in bacteria. Nat. Commun. 8, 2094 (2017).
Marciano, D. C. et al. Negative feedback in genetic circuits confers evolutionary resilience and capacitance. Cell Rep. 7, 1789–1795 (2014).
Yamamoto, N. et al. Update on the Keio collection of Escherichia coli single-gene deletion mutants. Mol. Syst. Biol. 5, 335 (2009).
Deatherage, D. & Barrick, J. Identification of mutations in laboratory-evolved microbes from next-generation sequencing data using breseq. Methods Mol. Biol. 1151, 165–188 (2014).
Arutyunov, D., Arenson, B., Manchak, J. & Frost, L. S. F plasmid TraF and TraH are components of an outer membrane complex involved in conjugation. J. Bacteriol. 192, 1730–1734 (2010).
Mazoyer, A. et al. flan: an R package for inference on mutation models. R. J. 9, 334–351 (2017).
Zwietering et al. Modeling of the bacterial growth curve. Appl. Environ. Microbiol. 56, 1875–1881 (1990).
Acknowledgements
The research was supported by the KU Leuven Research Council (PF/10/010, PDM/17/130, C1/17 3E170455), FWO (G047112N, G055517N, G0B2515N) and the VIB. The authors also gratefully acknowledge support from the National Institutes of Health (R01GM48746 to P.J.C.; R01GM088653 to C.H.; NIH-GM079656 and NIH-GM066099 to O.L.) and the National Science Foundation (NSF DBI-1356569 to O.L.). We thank Herman Dierick for helpful discussions and suggestions.
Author information
Authors and Affiliations
Contributions
T.S. designed CRISPR-FRT and its applications, performed and discussed the experiments and wrote and edited the manuscript. D.C.M. designed CRISPR-FRT and its applications, performed and discussed the experiments and wrote and edited the manuscript. B.A. generated the recA mutant library. R.E.B. helped in performing the experiments. C.W. helped in performing the experiments. M.L. helped in performing the experiments. C.B. helped in performing the experiments. T.S. helped in performing the experiments. B.V.d.B. helped in performing the experiments. I.F. helped in performing the experiments. N.V. discussed the results and edited the manuscript. P.J.C. discussed the results and edited the manuscript. C.H. discussed the results and edited the manuscript. O.L. discussed the results and edited the manuscript. J.M. discussed the results and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The funding sources were not involved in study design, data collection and interpretation, or the decision to submit the work for publication.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Swings, T., Marciano, D.C., Atri, B. et al. CRISPR-FRT targets shared sites in a knock-out collection for off-the-shelf genome editing. Nat Commun 9, 2231 (2018). https://doi.org/10.1038/s41467-018-04651-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-018-04651-5
This article is cited by
-
Deep mutational scanning of essential bacterial proteins can guide antibiotic development
Nature Communications (2023)
-
Strategies to Enhance the Biosynthesis of Monounsaturated Fatty Acids in Escherichia coli
Biotechnology and Bioprocess Engineering (2023)
-
Evolutionary action of mutations reveals antimicrobial resistance genes in Escherichia coli
Nature Communications (2022)
-
Structure of a type IV secretion system core complex encoded by multi-drug resistance F plasmids
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.