Abstract
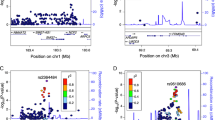
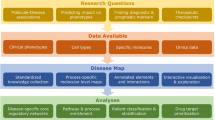
Genome-wide association studies have identified ~170 loci associated with Crohn’s disease (CD) and defining which genes drive these association signals is a major challenge. The primary aim of this study was to define which CD locus genes are most likely to be disease related. We developed a gene prioritization regression model (GPRM) by integrating complementary mRNA expression datasets, including bulk RNA-Seq from the terminal ileum of 302 newly diagnosed, untreated CD patients and controls, and in stimulated monocytes. Transcriptome-wide association and co-expression network analyses were performed on the ileal RNA-Seq datasets, identifying 40 genome-wide significant genes. Co-expression network analysis identified a single gene module, which was substantially enriched for CD locus genes and most highly expressed in monocytes. By including expression-based and epigenetic information, we refined likely CD genes to 2.5 prioritized genes per locus from an average of 7.8 total genes. We validated our model structure using cross-validation and our prioritization results by protein-association network analyses, which demonstrated significantly higher CD gene interactions for prioritized compared with non-prioritized genes. Although individual datasets cannot convey all of the information relevant to a disease, combining data from multiple relevant expression-based datasets improves prediction of disease genes and helps to further understanding of disease pathogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Jostins L, Ripke S, Weersma RK, Duerr RH, McGovern DP, Hui KY, et al. Host-microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature. 2012;491:119–24.
Liu JZ, van Sommeren S, Huang H, Ng SC, Alberts R, Takahashi, A et al. Association analyses identify 38 susceptibility loci for inflammatory bowel disease and highlight shared genetic risk across populations. Nat Genet 2015;47:979–86.
Di Meglio P, Di Cesare A, Laggner U, Chu CC, Napolitano L, Villanova F, et al. The IL23R R381Q gene variant protects against immune-mediated diseases by impairing IL-23-induced Th17 effector response in humans. PLoS ONE. 2011;6:e17160.
Pidasheva S, Trifari S, Phillips A, Hackney JA, Ma Y, Smith A, et al. Functional studies on the IBD susceptibility gene IL23R implicate reduced receptor function in the protective genetic variant R381Q. PLoS ONE. 2011;6:e25038.
Sarin R, Wu X, Abraham C. Inflammatory disease protective R381Q IL23 receptor polymorphism results in decreased primary CD4+and CD8+human T-cell functional responses. Proc Natl Acad Sci USA. 2011;108:9560–5.
Lassen KG, McKenzie CI, Mari M, Murano T, Begun J, Baxt LA, et al. Genetic coding variant in GPR65 alters lysosomal pH and links lysosomal dysfunction with colitis risk. Immunity. 2016;44:1392–405.
Murthy A, Li Y, Peng I, Reichelt M, Katakam AK, Noubade R, et al. A Crohn’s disease variant in Atg16l1 enhances its degradation by caspase 3. Nature. 2014;506:456–62.
Ogura Y, Bonen DK, Inohara N, Nicolae DL, Chen FF, Ramos R, et al. A frameshift mutation in NOD2 associated with susceptibility to Crohn’s disease. Nature. 2001;411:603–6.
Bonen DK, Ogura Y, Nicolae DL, Inohara N, Saab L, Tanabe T, et al. Crohn’s disease-associated NOD2 variants share a signaling defect in response to lipopolysaccharide and peptidoglycan. Gastroenterology. 2003;124:140–6.
van Heel DA, Ghosh S, Butler M, Hunt KA, Lundberg AM, Ahmad T, et al. Muramyl dipeptide and toll-like receptor sensitivity in NOD2-associated Crohn’s disease. Lancet. 2005;365:1794–6.
Lahiri A, Hedl M, Yan J, Abraham C. Human LACC1 increases innate receptor-induced responses and a LACC1 disease-risk variant modulates these outcomes. Nat Commun. 2017;8:15614.
Glocker EO, Kotlarz D, Boztug K, Gertz EM, Schaffer AA, Noyan F, et al. Inflammatory bowel disease and mutations affecting the interleukin-10 receptor. N Engl J Med. 2009;361:2033–45.
Huang H, Fang M, Jostins L, Umicevic Mirkov M, Boucher G, Anderson CA, et al. Fine-mapping inflammatory bowel disease loci to single-variant resolution. Nature. 2017;547:173–8.
Fairfax BP, Humburg P, Makino S, Naranbhai V, Wong D, Lau E, et al. Innate immune activity conditions the effect of regulatory variants upon monocyte gene expression. Science. 2014;343:1246949.
Consortium GT. Human genomics. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science. 2015;348:648–60.
Pierson E, Consortium GT, Koller D, Battle A, Mostafavi S, Ardlie KG, et al. Sharing and specificity of co-expression networks across 35 human tissues. PLoS Comput Biol. 2015;11:e1004220.
Holtta V, Sipponen T, Westerholm-Ormio M, Salo HM, Kolho KL, Farkkila M, et al. In Crohn’s disease, anti-TNF-alpha treatment changes the balance between mucosal IL-17, FOXP3, and CD4 cells. ISRN Gastroenterol. 2012;2012:505432.
Eriksson C, Rantapaa-Dahlqvist S, Sundqvist KG. Changes in chemokines and their receptors in blood during treatment with the TNF inhibitor infliximab in patients with rheumatoid arthritis. Scand J Rheumatol. 2013;42:260–5.
Eder P, Lykowska-Szuber L, Krela-Kazmierczak I, Stawczyk-Eder K, Zabel M, Linke K. The influence of infliximab and adalimumab on the expression of apoptosis-related proteins in lamina propria mononuclear cells and enterocytes in Crohn’s disease - an immunohistochemical study. J Crohn Colitis. 2013;7:706–16.
Hvas CL, Kelsen J, Agnholt J, Dige A, Christensen LA, Dahlerup JF. Discrete changes in circulating regulatory T cells during infliximab treatment of Crohn’s disease. Autoimmunity. 2010;43:325–33.
Haberman Y, Tickle TL, Dexheimer PJ, Kim MO, Tang D, Karns R, et al. Pediatric Crohn disease patients exhibit specific ileal transcriptome and microbiome signature. J Clin Invest. 2014;124:3617–33.
Kugathasan S, Denson LA, Walters TD, Kim MO, Marigorta UM, Schirmer M, et al. Prediction of complicated disease course for children newly diagnosed with Crohn’s disease: a multicentre inception cohort study. Lancet. 2017;38:1710–8.
Giambartolomei C, Vukcevic D, Schadt EE, Franke L, Hingorani AD, Wallace C, et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 2014;10:e1004383.
Plagnol V, Smyth DJ, Todd JA, Clayton DG. Statistical independence of the colocalized association signals for type 1 diabetes and RPS26 gene expression on chromosome 12q13. Biostatistics. 2009;10:327–34.
Howie BN, Donnelly P, Marchini J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 2009;5:e1000529.
Delaneau O, Zagury JF, Marchini J. Improved whole-chromosome phasing for disease and population genetic studies. Nat Methods. 2013;10:5–6.
Delaneau O, Marchini J, Zagury JF. A linear complexity phasing method for thousands of genomes. Nat Methods. 2011;9:179–81.
Parts L, Stegle O, Winn J, Durbin R. Joint genetic analysis of gene expression data with inferred cellular phenotypes. PLoS Genet. 2011;7:e1001276.
Stegle O, Parts L, Durbin R, Winn J. A Bayesian framework to account for complex non-genetic factors in gene expression levels greatly increases power in eQTL studies. PLoS Comput Biol. 2010;6:e1000770.
Lassen KG, Kuballa P, Conway KL, Patel KK, Becker CE, Peloquin JM, et al. Atg16L1 T300A variant decreases selective autophagy resulting in altered cytokine signaling and decreased antibacterial defense. Proc Natl Acad Sci USA. 2014;111:7741–6.
Gusev A, Ko A, Shi H, Bhatia G, Chung W, Penninx BW, et al. Integrative approaches for large-scale transcriptome-wide association studies. Nat Genet. 2016;48:245–52.
Pasaniuc B, Zaitlen N, Shi H, Bhatia G, Gusev A, Pickrell J, et al. Fast and accurate imputation of summary statistics enhances evidence of functional enrichment. Bioinformatics (Oxf, Engl). 2014;30:2906–14.
Yang J, Benyamin B, McEvoy BP, Gordon S, Henders AK, Nyholt DR, et al. Common SNPs explain a large proportion of the heritability for human height. Nat Genet. 2010;42:565–9.
Cutler DJ, Zwick ME, Okou DT, Prahalad S, Walters T, Guthery SL, et al. Dissecting allele architecture of early onset IBD using high-density genotyping. PLoS ONE. 2015;10:e0128074.
de Lange KM, Moutsianas L, Lee JC, Lamb CA, Luo Y, Kennedy NA, et al. Genome-wide association study implicates immune activation of multiple integrin genes in inflammatory bowel disease. Nat Genet. 2017;49:256–61.
Ching KH, Kisailus AE, Burbelo PD. Biochemical characterization of distinct regions of SPEC molecules and their role in phagocytosis. Exp Cell Res. 2007;313:10–21.
Rioux JD, Daly MJ, Silverberg MS, Lindblad K, Steinhart H, Cohen Z, et al. Genetic variation in the 5q31 cytokine gene cluster confers susceptibility to Crohn disease. Nat Genet. 2001;29:223–8.
Langfelder P, Horvath S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics. 2008;9:559.
Eden E, Navon R, Steinfeld I, Lipson D, Yakhini Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinformatics. 2009;10:48.
Zheng GX, Terry JM, Belgrader P, Ryvkin P, Bent ZW, Wilson R, et al. Massively parallel digital transcriptional profiling of single cells. Nat Commun. 2017;8:14049.
Wong KL, Tai JJ, Wong WC, Han H, Sem X, Yeap WH, et al. Gene expression profiling reveals the defining features of the classical, intermediate, and nonclassical human monocyte subsets. Blood. 2011;118:e16–31.
Villani AC, Satija R, Reynolds G, Sarkizova S, Shekhar K, Fletcher J, et al. Single-cell RNA-seq reveals new types of human blood dendritic cells, monocytes, and progenitors. Science 2017;356:pii: eaah4573.
Ning K, Gettler K, Zhang W, Ng SM, Bowen BM, Hyams J, et al. Improved integrative framework combining association data with gene expression features to prioritize Crohn’s disease genes. Hum Mol Genet. 2015;24:4147–57.
Farh KK, Marson A, Zhu J, Kleinewietfeld M, Housley WJ, Beik S, et al. Genetic and epigenetic fine mapping of causal autoimmune disease variants. Nature. 2015;518:337–43.
Szklarczyk D, Morris JH, Cook H, Kuhn M, Wyder S, Simonovic M, et al. The STRING database in 2017: quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res. 2017;45(D1):D362–D368.
Bain CC, Mowat AM. Macrophages in intestinal homeostasis and inflammation. Immunol Rev. 2014;260:102–17.
Chuang LS, Villaverde N, Hui KY, Mortha A, Rahman A, Levine AP, et al. A frameshift in CSF2RB predominant among Ashkenazi Jews increases risk for Crohn’s disease and reduces monocyte signaling via GM-CSF. Gastroenterology. 2016;151:710–23 e2.
Levine AP, Pontikos N, Schiff ER, Jostins L, Speed D, Consortium NIBDG. et al. Genetic complexity of Crohn’s disease in two large Ashkenazi Jewish families. Gastroenterology. 2016;151:698–709.
Brant SR, Okou DT, Simpson CL, Cutler DJ, Haritunians T, Bradfield JP, et al. Genome-wide association study identifies African-specific susceptibility loci in African Americans with inflammatory bowel disease. Gastroenterology. 2016;152:206–17.
Cros J, Cagnard N, Woollard K, Patey N, Zhang SY, Senechal B, et al. Human CD14dim monocytes patrol and sense nucleic acids and viruses via TLR7 and TLR8 receptors. Immunity. 2010;33:375–86.
Ziegler-Heitbrock L. The CD14+CD16+blood monocytes: their role in infection and inflammation. J Leukoc Biol. 2007;81:584–92.
Rosen MJ, Karns R, Vallance JE, Bezold R, Waddell A, Collins MH, et al. Mucosal expression of type 2 and type 17 immune response genes distinguishes ulcerative colitis from colon-only Crohn’s disease in treatment-naive pediatric patients. Gastroenterology. 2017;152:1345–57 e7.
Trapnell C, Pachter L, Salzberg SL. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics (Oxf, Engl). 2009;25:1105–11.
Trapnell C, Roberts A, Goff L, Pertea G, Kim D, Kelley DR, et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc. 2012;7:562–78.
Roberts A, Trapnell C, Donaghey J, Rinn JL, Pachter L. Improving RNA-Seq expression estimates by correcting for fragment bias. Genome Biol. 2011;12:R22.
Anders S, Pyl PT, Huber W. HTSeq--a Python framework to work with high-throughput sequencing data. Bioinformatics (Oxf, Engl). 2015;31:166–9.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet. 2007;81:559–75.
Chang CC, Chow CC, Tellier LC, Vattikuti S, Purcell SM, Lee JJ. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience. 2015;4:7.
Shabalin AA. Matrix eQTL: ultra fast eQTL analysis via large matrix operations. Bioinformatics (Oxf, Engl). 2012;28:1353–8.
Yang J, Lee SH, Goddard ME, Visscher PM. GCTA: a tool for genome-wide complex trait analysis. Am J Hum Genet. 2011;88:76–82.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Franke A, McGovern DP, Barrett JC, Wang K, Radford-Smith GL, Ahmad T, et al. Genome-wide meta-analysis increases to 71 the number of confirmed Crohn’s disease susceptibility loci. Nat Genet. 2010;42:1118–25.
Devlin B, Roeder K. Genomic control for association studies. Biometrics. 1999;55:997–1004.
Liu JZ, McRae AF, Nyholt DR, Medland SE, Wray NR, Brown KM, et al. A versatile gene-based test for genome-wide association studies. Am J Hum Genet. 2010;87:139–45.
Pena OM, Pistolic J, Raj D, Fjell CD, Hancock RE. Endotoxin tolerance represents a distinctive state of alternative polarization (M2) in human mononuclear cells. J Immunol. 2011;186:7243–54.
Du P, Kibbe WA, Lin SM. lumi: a pipeline for processing Illumina microarray. Bioinformatics (Oxf, Engl). 2008;24:1547–8.
Jaitin DA, Kenigsberg E, Keren-Shaul H, Elefant N, Paul F, Zaretsky I, et al. Massively parallel single-cell RNA-seq for marker-free decomposition of tissues into cell types. Science. 2014;343:776–9.
Acknowledgements
This work was supported by the HPC facilities operated by, and the staffs of, the Yale Center for Research Computing and the Yale Center for Genome Analysis, as well as NIH grant 1S10OD018521–01, which helped fund the cluster.
Funding
This study was supported by NIH research grants (R01 DK092235, U01 DK062429, U01 DK062422, R01 DK106593, and P30 DK078392), as well as the Crohn’s and Colitis Foundation, the Helmsley Charitable Trust, and the Sanford Grossman Charitable Trust.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Gettler, K., Giri, M., Kenigsberg, E. et al. Prioritizing Crohn’s disease genes by integrating association signals with gene expression implicates monocyte subsets. Genes Immun 20, 577–588 (2019). https://doi.org/10.1038/s41435-019-0059-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41435-019-0059-y
This article is cited by
-
Transcriptome-wide association studies associated with Crohn’s disease: challenges and perspectives
Cell & Bioscience (2024)
-
The relationship between extreme inter-individual variation in macrophage gene expression and genetic susceptibility to inflammatory bowel disease
Human Genetics (2024)
-
Macrophages in intestinal homeostasis and inflammatory bowel disease
Nature Reviews Gastroenterology & Hepatology (2023)
-
Identifying high-impact variants and genes in exomes of Ashkenazi Jewish inflammatory bowel disease patients
Nature Communications (2023)
-
Interpretation of the role of germline and somatic non-coding mutations in cancer: expression and chromatin conformation informed analysis
Clinical Epigenetics (2022)