Abstract
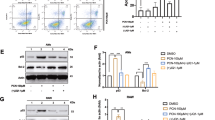
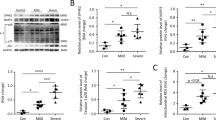
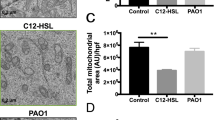
The metabolic reprogramming underlying the generation of regulatory B cells during infectious diseases remains unknown. Using a Pseudomonas aeruginosa-induced pneumonia model, we reported that IL-10-producing B cells (IL-10+ B cells) play a key role in spontaneously resolving infection-mediated inflammation. Accumulated cytosolic reactive oxygen species (ROS) during inflammation were shown to drive IL-10+ B-cell generation by remodeling one-carbon metabolism. Depletion of the enzyme serine hydroxymethyltransferase 1 (Shmt1) led to inadequate one-carbon metabolism and decreased IL-10+ B-cell production. Furthermore, increased one-carbon flux elevated the levels of the methyl donor S-adenosylmethionine (SAM), altering histone H3 lysine 4 methylation (H3K4me) at the Il10 gene to promote chromatin accessibility and upregulate Il10 expression in B cells. Therefore, the one-carbon metabolism-associated compound ethacrynic acid (EA) was screened and found to potentially treat infectious pneumonia by boosting IL-10+ B-cell generation. Overall, these findings reveal that ROS serve as modulators to resolve inflammation by reprogramming one-carbon metabolism pathways in B cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The raw ATAC sequencing and RNA sequencing data have been submitted to the GEO repository (GSE211326, GSE211327). The untargeted metabolomic data have been submitted to the MetaboLights database with the identifier MTBLS5664. Any additional information needed to reanalyze the data reported in this paper is available from the lead contact upon request.
References
Rosser EC, Piper C, Matei DE, Blair PA, Rendeiro AF, Orford M, et al. Microbiota-derived metabolites suppress arthritis by amplifying aryl-hydrocarbon receptor activation in regulatory B cells. Cell Metab. 2020;31:837–851.e10.
Jansen K, Cevhertas L, Ma S, Satitsuksanoa P, Akdis M, van de Veen W. Regulatory B cells, A to Z. Allergy. 2021;76:2699–715.
Wang Z, Lu Z, Lin S, Xia J, Zhong Z, Xie Z, et al. Leucine-tRNA-synthase-2-expressing B cells contribute to colorectal cancer immunoevasion. Immunity. 2022;55:1067–1081.e8.
Baker RE, Mahmud AS, Miller IF, Rajeev M, Rasambainarivo F, Rice BL, et al. Infectious disease in an era of global change. Nat Rev Microbiol. 2022;20:193–205.
Edwards AM, Baric RS, Saphire EO, Ulmer JB. Stopping pandemics before they start: lessons learned from SARS-CoV-2. Science. 2022;375:1133–9.
Moon HG, Kim SJ, Jeong JJ, Han SS, Jarjour NN, Lee H, et al. Airway epithelial cell-derived colony stimulating factor-1 promotes allergen sensitization. Immunity. 2018;49:275–287.e5.
Rosser EC, Mauri C. Regulatory B cells: origin, phenotype, and function. Immunity. 2015;42:607–12.
Wang L, Fu Y, Chu Y. Regulatory B cells. Adv Exp Med Biol. 2020;1254:87–103.
Roux C, Jafari SM, Shinde R, Duncan G, Cescon DW, Silvester J, et al. Reactive oxygen species modulate macrophage immunosuppressive phenotype through the up-regulation of PD-L1. Proc Natl Acad Sci USA. 2019;116:4326–35.
Ohl K, Tenbrock K. Reactive oxygen species as regulators of MDSC-mediated immune suppression. Front Immunol. 2018;9:2499.
Zhang D, Chia C, Jiao X, Jin W, Kasagi S, Wu R, et al. D-mannose induces regulatory T cells and suppresses immunopathology. Nat Med. 2017;23:1036–45.
Olenchock BA, Rathmell JC, Vander Heiden MG. Biochemical underpinnings of immune cell metabolic phenotypes. Immunity. 2017;46:703–13.
Kedia-Mehta N, Finlay DK. Competition for nutrients and its role in controlling immune responses. Nat Commun. 2019;10:2123.
Caro-Maldonado A, Wang R, Nichols AG, Kuraoka M, Milasta S, Sun LD, et al. Metabolic reprogramming is required for antibody production that is suppressed in anergic but exaggerated in chronically BAFF-exposed B cells. J Immunol. 2014;192:3626–36.
Cyster JG, Allen CDC. B cell responses: cell interaction dynamics and decisions. Cell. 2019;177:524–40.
Locasale JW. Serine, glycine and one-carbon units: cancer metabolism in full circle. Nat Rev Cancer. 2013;13:572–83.
Laforge M, Elbim C, Frère C, Hémadi M, Massaad C, Nuss P, et al. Tissue damage from neutrophil-induced oxidative stress in COVID-19. Nat Rev Immunol. 2020;20:515–6.
Piper CJM, Rosser EC, Oleinika K, Nistala K, Krausgruber T, Rendeiro AF, et al. Aryl hydrocarbon receptor contributes to the transcriptional program of IL-10-producing regulatory B cells. Cell Rep. 2019;29:1878–1892.e7.
Sugiura A, Andrejeva G, Voss K, Heintzman DR, Xu X, Madden MZ, et al. MTHFD2 is a metabolic checkpoint controlling effector and regulatory T cell fate and function. Immunity. 2022;55:65–81.e9.
Ho HJ, Aoki N, Wu Y-J, Gao M-C, Sekine K, Sakurai T, et al. A pacific oyster-derived antioxidant, DHMBA, protects renal tubular HK-2 cells against oxidative stress via reduction of mitochondrial ROS production and fragmentation. Int J Mol Sci. 2023;24:10061.
Jin P, Zhou Q, Xi S. Low-dose arsenite causes overexpression of EGF, TGFalpha, and HSP90 through Trx1-TXNIP-NLRP3 axis mediated signaling pathways in the human bladder epithelial cells. Ecotoxicol Environ Saf. 2022;247:114263.
Li X, Egervari G, Wang Y, Berger SL, Lu Z. Regulation of chromatin and gene expression by metabolic enzymes and metabolites. Nat Rev Mol Cell Biol. 2018;19:563–78.
Yang WS, Kim JH, Jeong D, Hong YH, Park SH, Yang Y, et al. 3-Deazaadenosine, an S-adenosylhomocysteine hydrolase inhibitor, attenuates lipopolysaccharide-induced inflammatory responses via inhibition of AP-1 and NF-kappaB signaling. Biochem Pharm. 2020;182:114264.
Yu W, Wang Z, Zhang K, Chi Z, Xu T, Jiang D, et al. One-carbon metabolism supports S-adenosylmethionine and histone methylation to drive inflammatory macrophages. Mol Cell. 2019;75:1147–1160.e5.
Raza IGA, Clarke AJ. B cell metabolism and autophagy in autoimmunity. Front Immunol. 2021;12:681105.
Liu X, Jiang X, Liu R, Wang L, Qian T, Zheng Y, et al. B cells expressing CD11b effectively inhibit CD4 T cell responses and ameliorate experimental autoimmune hepatitis. Hepatol Nov. 2015;62:1563–75.
Wang L, Ray A, Jiang X, Wang J, Basu S, Liu X, et al. T regulatory cells and B cells cooperate to form a regulatory loop that maintains gut homeostasis and suppresses dextran sulfate sodium-induced colitis. Mucosal Immunol Nov. 2015;8:1297–312.
Stark AK, Chandra A, Chakraborty K, Alam R, Carbonaro V, Clark J, et al. PI3Kdelta hyper-activation promotes development of B cells that exacerbate Streptococcus pneumoniae infection in an antibody-independent manner. Nat Commun. 2018;9:3174.
Rong HM, Li T, Zhang C, Wang D, Hu Y, Zhai K, et al. IL-10-producing B cells regulate Th1/Th17-cell immune responses in Pneumocystis pneumonia. Am J Physiol Lung Cell Mol Physiol. 2019;316:L291–301.
Song H, Xi J, Li GG, Xu S, Wang C, Cheng T, et al. Upregulation of CD19(+)CD24(hi)CD38(hi) regulatory B cells is associated with a reduced risk of acute lung injury in elderly pneumonia patients. Intern Emerg Med. 2016;11:415–23.
Fanucchi S, Dominguez-Andres J, Joosten LAB, Netea MG, Mhlanga MM. The intersection of epigenetics and metabolism in trained immunity. Immunity. 2021;54:32–43.
Hultqvist M, Olofsson P, Gelderman KA, Holmberg J, Holmdahl R. A new arthritis therapy with oxidative burst inducers. PLoS Med. 2006;3:e348.
de Alencar JCG, Moreira CL, Müller AD, Chaves CE, Fukuhara MA, da Silva EA, et al. Double-blind, randomized, placebo-controlled trial with N-acetylcysteine for treatment of severe acute respiratory syndrome caused by coronavirus disease 2019 (COVID-19). Clin Infect Dis. 2021;72:e736–e741.
Taher A, Lashgari M, Sedighi L, Rahimi-Bashar F, Poorolajal J, Mehrpooya M. A pilot study on intravenous N-Acetylcysteine treatment in patients with mild-to-moderate COVID19-associated acute respiratory distress syndrome. Pharm Rep. 2021;73:1650–9.
Alfonso H, Franklin P, Ching S, Croft K, Burcham P, Olsen N, et al. Effect of N-acetylcysteine supplementation on oxidative stress status and alveolar inflammation in people exposed to asbestos: a double-blind, randomized clinical trial. Respirology. 2015;20:1102–7.
Muri J, Kopf M. Redox regulation of immunometabolism. Nat Rev Immunol. 2021;21:363–81.
Yang HY, Kim J, Lee KY, Jang YS. Rac/ROS-related protein kinase C and phosphatidylinositol-3-kinase signaling are involved in a negative regulating cascade in B cell activation by antibody-mediated cross-linking of MHC class II molecules. Mol Immunol. 2010;47:706–12.
Luo W, Weisel F, Shlomchik MJ. B cell receptor and CD40 signaling are rewired for synergistic induction of the c-Myc transcription factor in germinal center B cells. Immunity. 2018;48:313–326.e5.
Zeng JD, Wu WKK, Wang HY, Li XX. Serine and one-carbon metabolism, a bridge that links mTOR signaling and DNA methylation in cancer. Pharm Res. 2019;149:104352.
Anastasiou D, Poulogiannis G, Asara JM, Boxer MB, Jiang JK, Shen M, et al. Inhibition of pyruvate kinase M2 by reactive oxygen species contributes to cellular antioxidant responses. Science. 2011;334:1278–83.
Sadhu MJ, Guan Q, Li F, Sales-Lee J, Iavarone AT, Hammond MC, et al. Nutritional control of epigenetic processes in yeast and human cells. Genetics. 2013;195:831–44.
Su X, Wellen KE, Rabinowitz JD. Metabolic control of methylation and acetylation. Curr Opin Chem Biol. 2016;30:52–60.
Ara AI, Xia M, Ramani K, Mato JM, Lu SC. S-adenosylmethionine inhibits lipopolysaccharide-induced gene expression via modulation of histone methylation. Hepatology. 2008;47:1655–66.
van de Wouw M, Walsh CJ, Vigano G, Lyte JM, Boehme M, Gual-Grau A, et al. Kefir ameliorates specific microbiota-gut-brain axis impairments in a mouse model relevant to autism spectrum disorder. Brain Behav Immun. 2021;97:119–34.
Bian Y, Li W, Kremer DM, Sajjakulnukit P, Li S, Crespo J, et al. Cancer SLC43A2 alters T cell methionine metabolism and histone methylation. Nature. 2020;585:277–82.
Sahin E, Sahin M. Epigenetical targeting of the FOXP3 gene by S-adenosylmethionine diminishes the suppressive capacity of regulatory T cells ex vivo and alters the expression profiles. J Immunother. 2019;42:11–22.
Borck PC, Guo LW, Plutzky J. BET epigenetic reader proteins in cardiovascular transcriptional programs. Circ Res. 2020;126:1190–208.
Millan-Zambrano G, Burton A, Bannister AJ, Schneider R. Histone post-translational modifications—cause and consequence of genome function. Nat Rev Genet. 2022;23:563–80.
Stephen-Victor E, Das M, Karnam A, Pitard B, Gautier J-F, Bayry J. Potential of regulatory T-cell-based therapies in the management of severe COVID-19. Eur Respir J. 2020;56:2002182.
Matute-Bello G, Downey G, Moore BB, Groshong SD, Matthay MA, Slutsky AS, et al. An official American Thoracic Society workshop report: features and measurements of experimental acute lung injury in animals. Am J Respir Cell Mol Biol. 2011;44:725–38.
Bodogai M, Lee Chang C, Wejksza K, Lai J, Merino M, Wersto RP, et al. Anti-CD20 antibody promotes cancer escape via enrichment of tumor-evoked regulatory B cells expressing low levels of CD20 and CD137L. Cancer Res. 2013;73:2127–38.
Sarikonda G, Sachithanantham S, Manenkova Y, Kupfer T, Posgai A, Wasserfall C, et al. Transient B-cell depletion with anti-CD20 in combination with proinsulin DNA vaccine or oral insulin: immunologic effects and efficacy in NOD mice. PLoS ONE. 2013;8:e54712.
Peng J, Peng J, Huang L, Enkhjargal B, Zhang T, Mo J. LRP1 activation attenuates white matter injury by modulating microglial polarization through Shc1/PI3K/Akt pathway after subarachnoid hemorrhage in rats. Redox Biol. 2019;21:101121.
Buenrostro JD, Giresi PG, Zaba LC, Chang HY, Greenleaf WJ. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat Methods. 2013;10:1213–8.
Kerur N, Fukuda S, Banerjee D. cGAS drives noncanonical-inflammasome activation in age-related macular degeneration. Nat Med. 2018;24:50–61.
Zelena E, Dunn WB, Broadhurst D, Francis-McIntyre S, Carroll KM, Begley P, et al. Development of a robust and repeatable UPLC-MS method for the long-term metabolomic study of human serum. Anal Chem. 2009;81:1357–64.
Want EJ, Masson P, Michopoulos F, Wilson ID, Theodoridis G, Plumb RS, et al. Global metabolic profiling of animal and human tissues via UPLC-MS. Nat Protoc. 2013;8:17–32.
Acknowledgements
This work was supported by the General Program of the National Natural Science Foundation of China (81971493, 81771736) and the Science Fund for Creative Research Groups of the National Natural Science Foundation of China (82121002). Major Program of National Natural Science Foundation of China (821300501, 82330053), Shanghai Rising-Star Program (20QA1407900), and Innovative Research Team of High-level Local Universities in Shanghai.
Author information
Authors and Affiliations
Contributions
YF performed the experiments, discussed and analyzed the data and wrote the paper. BY and QW performed the statistical analysis. HZ assisted in the experiment and provided reagent or technical support. DZ performed the CyTOF experiments. FL and RL discussed and wrote the paper. YC and LW conceived and supervised the study, designed the research, interpreted the results, and wrote the paper. YC is the primary contact for communication.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fu, Y., Yu, B., Wang, Q. et al. Oxidative stress-initiated one-carbon metabolism drives the generation of interleukin-10-producing B cells to resolve pneumonia. Cell Mol Immunol 21, 19–32 (2024). https://doi.org/10.1038/s41423-023-01109-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41423-023-01109-7