Abstract
Background
The clinical value and molecular characteristics of tumor differentiation in oral squamous cell carcinoma (OSCC) remain unclear. There is a lack of a related molecular classification prediction system based on pathological images for precision medicine.
Methods
Integration of epidemiology, genomics, experiments, and deep learning to clarify the clinical value and molecular characteristics, and develop a novel OSCC molecular classification prediction system.
Results
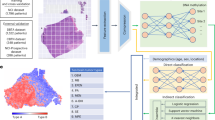
Large-scale epidemiology data (n = 118,817) demonstrated OSCC differentiation was a significant prognosis indicator (p < 0.001), and well-differentiated OSCC was more chemo-resistant than poorly differentiated OSCC. These results were confirmed in the TCGA database and in vitro. Furthermore, we found chemo-resistant related pathways and cell cycle-related pathways were up-regulated in well- and poorly differentiated OSCC, respectively. Based on the characteristics of OSCC differentiation, a molecular grade of OSCC was obtained and combined with pathological images to establish a novel prediction system through deep learning, named ShuffleNetV2-based Molecular Grade of OSCC (SMGO). Importantly, our independent multi-center cohort of OSCC (n = 340) confirmed the high accuracy of SMGO.
Conclusions
OSCC differentiation was a significant indicator of prognosis and chemotherapy selection. Importantly, SMGO could be an indispensable reference for OSCC differentiation and assist the decision-making of chemotherapy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 24 print issues and online access
$259.00 per year
only $10.79 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The large-scale epidemiological information is available from the SEER database (https://seer.cancer.gov/) from 1975 to 2016 using SEER*Stat 8.3.6. The clinical information, somatic mutation profiles, RNA-seq profiles, and pathological images of the TCGA database were downloaded from https://portal.gdc.cancer.gov/. CNV profiles in the TCGA database were obtained from UCSC Xena (https://xenabrowser.net/). The RNA-seq profiles of OSCC cell lines were obtained from the GEO database (https://www.ncbi.nlm.nih.gov/geo/; GEO accession numbers: GSE98942 and GSE146483). The clinical information and pathological images of the IMC cohort are available within the article, its supplementary information, or from the corresponding first author upon reasonable request. Source data are provided in this study.
Code availability
The codes of this study are provided on GitHub (https://github.com/chxhstatis/SMGO).
References
Arnold M, Ferlay J, van Berge Henegouwen MI, Soerjomataram I. Global burden of oesophageal and gastric cancer by histology and subsite in 2018. Gut. 2020;69:1564–71.
Arun I, Maity N, Hameed S, Jain PV, Manikantan K, Sharan R, et al. Lymph node characteristics and their prognostic significance in oral squamous cell carcinoma. Head Neck. 2021;43:520–33.
Sawair FA, Irwin CR, Gordon DJ, Leonard AG, Stephenson M, Napier SS. Invasive front grading: reliability and usefulness in the management of oral squamous cell carcinoma. J Oral Pathol Med. 2003;32:1–9.
Wunschel M, Neumeier M, Utpatel K, Reichert TE, Ettl T, Spanier G. Staging more important than grading? Evaluation of malignancy grading, depth of invasion, and resection margins in oral squamous cell carcinoma. Clin Oral Investig. 2020;25:1169–82.
Thomas B, Stedman M, Davies L. Grade as a prognostic factor in oral squamous cell carcinoma: a population-based analysis of the data. Laryngoscope. 2014;124:688–94.
Xu QS, Wang C, Li B, Li JZ, Mao MH, Qin LZ, et al. Prognostic value of pathologic grade for patients with oral squamous cell carcinoma. Oral Dis. 2018;24:335–46.
EI-Naggar AK, Chan JKC, Grandis JR, Takata T, Slootweg PJ. (eds.) WHO Classification of Head and Neck Tumours (4th edition). IACR: Lyon; 2017. p. 109–111.
Anneroth G, Batsakis J, Luna M. Review of the literature and a recommended system of malignancy grading in oral squamous cell carcinomas. Scand J Dent Res. 1987;95:229–49.
Natrajan R, Sailem H, Mardakheh FK, Arias Garcia M, Tape CJ, Dowsett M, et al. Microenvironmental heterogeneity parallels breast cancer progression: a histology-genomic integration analysis. PLoS Med. 2016;13:e1001961.
Zhai H, Li D, Feng Q, Qian X, Li L, Yao J. Pancreatic neuroendocrine tumors: grade is superior to T, N, or M status in predicting outcome and selecting patients for chemotherapy: a retrospective cohort study in the SEER database. Int J Surg. 2019;66:103–9.
Metzger-Filho O, Ferreira AR, Jeselsohn R, Barry WT, Dillon DA, Brock JE, et al. Mixed invasive ductal and lobular carcinoma of the breast: prognosis and the importance of histologic grade. Oncologist. 2019;24:e441–e9.
Lin NC, Hsu JT, Tsai KY. Survival and clinicopathological characteristics of different histological grades of oral cavity squamous cell carcinoma: A single-center retrospective study. PLoS One. 2020;15:e0238103.
Bhattacharya A, Toth K, Mazurchuk R, Spernyak JA, Slocum HK, Pendyala L, et al. Lack of microvessels in well-differentiated regions of human head and neck squamous cell carcinoma A253 associated with functional magnetic resonance imaging detectable hypoxia, limited drug delivery, and resistance to irinotecan therapy. Clin Cancer Res. 2004;10:8005–17.
Fine RL, Gulati AP, Krantz BA, Moss RA, Schreibman S, Tsushima DA, et al. Capecitabine and temozolomide (CAPTEM) for metastatic, well-differentiated neuroendocrine cancers: the Pancreas Center at Columbia University experience. Cancer Chemother Pharmacol. 2013;71:663–70.
Sotiriou C, Wirapati P, Loi S, Harris A, Fox S, Smeds J, et al. Gene expression profiling in breast cancer: understanding the molecular basis of histologic grade to improve prognosis. J Natl Cancer Inst. 2006;98:262–72.
Zhang N, Zhang H, Wang Z, Dai Z, Zhang X, Cheng Q, et al. Immune infiltrating cells-derived risk signature based on large-scale analysis defines immune landscape and predicts immunotherapy responses in glioma tumor microenvironment. Front Immunol. 2021;12:691811.
Alakwaa FM, Savelieff MG. Bioinformatics analysis of metabolomics data unveils association of metabolic signatures with methylation in breast cancer. J Proteome Res. 2020;19:2879–89.
Yang K, Wu Y. A prognosis-related molecular subtype for early-stage non-small lung cell carcinoma by multi-omics integration analysis. BMC Cancer 2021;21:128.
Steigen SE, Soland TM, Nginamau ES, Laurvik H, Costea DE, Johannessen AC, et al. Grading of oral squamous cell carcinomas—Intra and interrater agreeability: simpler is better? J Oral Pathol Med. 2020;49:630–5.
Nagpal K, Foote D, Tan F, Liu Y, Chen PC, Steiner DF, et al. Development and validation of a deep learning algorithm for Gleason grading of prostate cancer from biopsy specimens. JAMA Oncol. 2020;6:1372–80.
Das N, Hussain E, Mahanta LB. Automated classification of cells into multiple classes in epithelial tissue of oral squamous cell carcinoma using transfer learning and convolutional neural network. Neural Netw. 2020;128:47–60.
Stephen BE, David RB, Carolyn CC, April GF, Frederick LG, Andy TI AJCC 7th Ed Cancer Staging Manual. the American Joint Committee on Cancer; 2009.
Kitamura R, Toyoshima T, Tanaka H, Kawano S, Kiyosue T, Matsubara R, et al. Association of cytokeratin 17 expression with differentiation in oral squamous cell carcinoma. J Cancer Res Clin Oncol. 2012;138:1299–310.
Momose F, Araida T, Negishi A, Ichijo H, Shioda S, Sasaki S. Variant sublines with different metastatic potentials selected in nude mice from human oral squamous cell carcinomas. J Oral Pathol Med. 1989;18:391–5.
Nakayama S, Sasaki A, Mese H, Alcalde RE, Matsumura T. Establishment of high and low metastasis cell lines derived from a human tongue squamous cell carcinoma. Invasion Metastasis 1998;18:219–28.
Kase Y, Uzawa K, Wagai S, Yoshimura S, Yamamoto JI, Toeda Y, et al. Engineered exosomes delivering specific tumor-suppressive RNAi attenuate oral cancer progression. Sci Rep. 2021;11:5897.
Zhou X, Wang A. Differential expression of oral cancer cell line UM1 and UM2 (data accessible at NCBI GEO database (Edgar et al., 2002), accession GSE98942).
Robinson MD, McCarthy DJ, Smyth GK. edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010;26:139–40.
Johnson DE, Burtness B, Leemans CR, Lui VWY, Bauman JE, Grandis JR. Head and neck squamous cell carcinoma. Nat Rev Dis Prim. 2020;6:92.
Katirachi SK, Grønlund MP, Jakobsen KK, Grønhøj C, von Buchwald C. The prevalence of HPV in oral cavity squamous cell carcinoma. Viruses. 2023;15:451.
Neena D, Shah S, Keyuri P, Munira J Histological grading of oral cancer: a comparison of different systems and their relation to lymph node metastasis. Nat J Commun Med. 2011;2:136–42.
Hawthorne VS, Huang WC, Neal CL, Tseng LM, Hung MC, Yu D. ErbB2-mediated Src and signal transducer and activator of transcription 3 activation leads to transcriptional up-regulation of p21Cip1 and chemoresistance in breast cancer cells. Mol Cancer Res. 2009;7:592–600.
Yin Y, Wang X, Li T, Ren Q, Li L, Sun X, et al. MicroRNA-221 promotes breast cancer resistance to adriamycin via modulation of PTEN/Akt/mTOR signaling. Cancer Med 2020;9:1544–52.
Perez-Sayans M, Suarez-Penaranda JM, Iruegas EP, de Almeida MR, Barros-Angueira F, Torreira MG, et al. p21Waf1/CIP1 is a poor diagnostic and prognostic marker for OSCC although its expression increases in patients with N1 regional metastasis. Cancer Biomark. 2015;15:19–26.
Goel S, Bergholz JS, Zhao JJ. Targeting CDK4 and CDK6 in cancer. Nat Rev Cancer. 2022;22:356–72.
Oh DY, Bang YJ. HER2-targeted therapies - a role beyond breast cancer. Nat Rev Clin Oncol. 2020;17:33–48.
Campanella G, Hanna MG, Geneslaw L, Miraflor A, Werneck Krauss Silva V, Busam KJ, et al. Clinical-grade computational pathology using weakly supervised deep learning on whole slide images. Nat Med. 2019;25:1301–9.
Weijers M, Snow GB, Bezemer PD, van der Waal I. Malignancy grading is no better than conventional histopathological grading in small squamous cell carcinoma of tongue and floor of mouth: retrospective study in 128 patients. J Oral Pathol Med. 2009;38:343–7.
Zhong LP, Zhang CP, Ren GX, Guo W, William WN Jr., Sun J, et al. Randomized phase III trial of induction chemotherapy with docetaxel, cisplatin, and fluorouracil followed by surgery versus up-front surgery in locally advanced resectable oral squamous cell carcinoma. J Clin Oncol. 2013;31:744–51.
Licitra L, Grandi C, Guzzo M, Mariani L, Lo Vullo S, Valvo F, et al. Primary chemotherapy in resectable oral cavity squamous cell cancer: a randomized controlled trial. J Clin Oncol. 2003;21:327–33.
Bossi P, Lo Vullo S, Guzzo M, Mariani L, Granata R, Orlandi E, et al. Preoperative chemotherapy in advanced resectable OCSCC: long-term results of a randomized phase III trial. Ann Oncol. 2014;25:462–6.
Bulten W, Pinckaers H, van Boven H, Vink R, de Bel T, van Ginneken B, et al. Automated deep-learning system for Gleason grading of prostate cancer using biopsies: a diagnostic study. Lancet Oncol. 2020;21:233–41.
Dorofte L, Grélaud D, Fiorentino M, Giunchi F, Ricci C, Franceschini T, et al. Low level of interobserver concordance in assessing histological subtype and tumor grade in patients with penile cancer may impair patient care. Virchows Arch. 2022;480:879–86.
Chai AWY, Lim KP, Cheong SC. Translational genomics and recent advances in oral squamous cell carcinoma. Semin Cancer Biol. 2020;61:71–83.
Diao P, Jiang Y, Li Y, Wu X, Li J, Zhou C, et al. Immune landscape and subtypes in primary resectable oral squamous cell carcinoma: prognostic significance and predictive of therapeutic response. J Immunother Cancer. 2021;9:e002434.
Wang CI, Kao HK, Chen TW, Huang Y, Cheng HW, Yi JS, et al. Characterization of copy number variations in oral cavity squamous cell carcinoma reveals a novel role for MLLT3 in cell invasiveness. Oncologist. 2019;24:e1388–e400.
Martinez-Marti A, Felip E, Matito J, Mereu E, Navarro A, Cedrés S, et al. Dual MET and ERBB inhibition overcomes intratumor plasticity in osimertinib-resistant-advanced non-small-cell lung cancer (NSCLC). Ann Oncol. 2017;28:2451–7.
Kina S, Kawabata-Iwakawa R, Miyamoto S, Arasaki A, Sunakawa H, Kinjo T. A molecular signature of well-differentiated oral squamous cell carcinoma reveals a resistance mechanism to metronomic chemotherapy and novel therapeutic candidates. J Drug Target. 2021;29:1118–1127.
Wu Q, Ma J, Wei J, Meng W, Wang Y, Shi M. FOXD1-AS1 regulates FOXD1 translation and promotes gastric cancer progression and chemoresistance by activating the PI3K/AKT/mTOR pathway. Mol Oncol. 2021;15:299–316.
Wu M, Zhang P. EGFR-mediated autophagy in tumourigenesis and therapeutic resistance. Cancer Lett. 2020;469:207–16.
Kukcinaviciute E, Jonusiene V, Sasnauskiene A, Dabkeviciene D, Eidenaite E, Laurinavicius A. Significance of Notch and Wnt signaling for chemoresistance of colorectal cancer cells HCT116. J Cell Biochem. 2018;119:5913–20.
Dasari S, Tchounwou PB. Cisplatin in cancer therapy: molecular mechanisms of action. Eur J Pharmacol. 2014;740:364–78.
Longley DB, Harkin DP, Johnston PG. 5-fluorouracil: mechanisms of action and clinical strategies. Nat Rev Cancer. 2003;3:330–8.
LeCun Y, Bengio Y, Hinton G. Deep learning. Nature. 2015;521:436–44.
Song Z, Zou S, Zhou W, Huang Y, Shao L, Yuan J, et al. Clinically applicable histopathological diagnosis system for gastric cancer detection using deep learning. Nat Commun. 2020;11:4294.
Acknowledgements
The authors thank all patients for their participation and donation of samples. Thanks for the support of the West China Hospital of Stomatology, Guangdong Provincial Stomatological Hospital, and the General Hospital of the People’s Liberation Army.
Funding
This work was supported by grants from: National Natural Science Foundation of China grant 82001059 (to HX). National Natural Science Foundation of China grant 82273165 (to XX). Sichuan Science and Technology Program grant 2022NSFSC1495 (to XX).
Author information
Authors and Affiliations
Contributions
QS: contributed to acquisition, analysis, interpretation, drafted manuscript, and critically revised the manuscript. YJ: contributed to experiments, interpretation, and critically revised the manuscript. ZW: contributed to experiments and interpretation. JP: contributed to data acquisition, analysis and drafted the part of the manuscript. ZX: contributed to analysis and drafted the part of the manuscript. WL: contributed to interpretation. DY: contributed to acquisition. HZ: contributed to interpretation and critically revised the manuscript. XX: contributed to interpretation and critically revised the manuscript. YZ: contributed to interpretation and critically revised the manuscript. XZ: contributed to interpretation and critically revised the manuscript. QC: contributed to conception, design, acquisition, and critically revised the manuscript. HX: contributed to conception, design, acquisition, interpretation, and critically revised the manuscript. All authors gave final approval and agreed to be accountable for all aspects of the work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
This study was approved by the ethics committees of the West China Hospital of Stomatology, Guangdong Provincial Stomatological Hospital, and the General Hospital of the People’s Liberation Army [WCHSIRB-D-2014-004]. This study was conducted following the Declaration of Helsinki. All participants eligible for this study had completed a written informed consent form.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Shang, Q., Jiang, Y., Wan, Z. et al. The clinical implication and translational research of OSCC differentiation. Br J Cancer 130, 660–670 (2024). https://doi.org/10.1038/s41416-023-02566-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-023-02566-7