Abstract
Background
Triple-negative breast cancer (TNBC) is the most heterogeneous breast cancer subtype. Partly due to its heterogeneity, it is currently challenging to stratify TNBC patients and predict treatment outcomes.
Methods
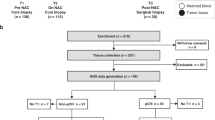
In this study, we examined blood cytokine profiles of TNBC patients throughout treatments (pre-treatment, during chemotherapy, pre-surgery, and 1 year after the surgery in a total of 294 samples). We analyzed the obtained cytokine datasets using weighted correlation network analyses, protein-protein interaction analyses, and logistic regression analyses.
Results
We identified five cytokines that correlate with good clinical outcomes: interleukin (IL)-1α, TNF-related apoptosis-inducing ligand (TRAIL), Stem Cell Factor (SCF), Chemokine ligand 5 (CCL5 also known as RANTES), and IL-16. The expression of these cytokines was decreased during chemotherapy and then restored after the treatment. Importantly, patients with good clinical outcomes had constitutively high expression of these cytokines during treatments. Protein-protein interaction analyses implicated that these five cytokines promote an immune response. Logistic regression analyses revealed that IL-1α and TRAIL expression levels at pre-treatment could predict treatment outcomes in our cohort.
Conclusion
We concluded that time-series cytokine profiles in breast cancer patients may be useful for understanding immune cell activity during treatment and for predicting treatment outcomes, supporting precision medicine.
Trial registration
The study has been registered with the University Hospital Medical Information Network Clinical Trials Registry (http://www.umin.ac.jp/ctr/index-j.htm) with the unique trial number UMIN000023162. The association Japan Breast Cancer Research Group trial number is JBCRG-22. The clinical outcome of the JBCRG-22 study was published in Breast Cancer Research and Treatment on 25 March 2021. https://doi.org/10.1007/s10549-021-06184-w.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 24 print issues and online access
$259.00 per year
only $10.79 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Clinical data and observed cytokine concentrations for all patients in the cohort at all time points for the 1st 48-plex and 40-plex measurements are given in Supplementary Table 3. Observed cytokine concentrations for all patients in the cohort at all time points for the 2nd 48-plex and 40-plex measurements are shown in Supplementary Table 4. Other raw data generated in this study are available upon request from the corresponding authors.
Code availability
All R and Python codes used in this study are available upon request.
References
Jabeen S, Espinoza JA, Torland LA, Zucknick M, Kumar S, Haakensen VD, et al. Noninvasive profiling of serum cytokines in breast cancer patients and clinicopathological characteristics. Oncoimmunology. 2019;8:e1537691.
Kawaguchi K, Sakurai M, Yamamoto Y, Suzuki E, Tsuda M, Kataoka TR, et al. Alteration of specific cytokine expression patterns in patients with breast cancer. Sci Rep. 2019;9:2924.
Garrone O, Michelotti A, Paccagnella M, Montemurro F, Vandone AM, Abbona A, et al. Exploratory analysis of circulating cytokines in patients with metastatic breast cancer treated with eribulin: the TRANSERI-GONO (Gruppo Oncologico del Nord Ovest) study. ESMO Open. 2020;5:e000876.
Carey LA, Dees EC, Sawyer L, Gatti L, Moore DT, Collichio F, et al. The triple negative paradox: primary tumor chemosensitivity of breast cancer subtypes. Clin Cancer Res. 2007;13:2329–34.
Lehmann BD, Bauer JA, Chen X, Sanders ME, Chakravarthy AB, Shyr Y, et al. Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J Clin Investig. 2011;121:2750–67.
Masuda H, Baggerly KA, Wang Y, Zhang Y, Gonzalez-Angulo AM, Meric-Bernstam F, et al. Differential response to neoadjuvant chemotherapy among 7 triple-negative breast cancer molecular subtypes. Clin Cancer Res. 2013;19:5533–40.
Birkbak NJ, Wang ZC, Kim J-Y, Eklund AC, Li Q, Tian R, et al. Telomeric allelic imbalance indicates defective DNA repair and sensitivity to DNA-damaging agents. Cancer Discov. 2012;2:366–75.
Chopra N, Tovey H, Pearson A, Cutts R, Toms C, Proszek P, et al. Homologous recombination DNA repair deficiency and PARP inhibition activity in primary triple negative breast cancer. Nat Commun. 2020;11:2662.
Sharma P, Barlow WE, Godwin AK, Pathak H, Isakova K, Williams D, et al. Impact of homologous recombination deficiency biomarkers on outcomes in patients with triple-negative breast cancer treated with adjuvant doxorubicin and cyclophosphamide (SWOG S9313). Ann Oncol. 2018;29:654–60.
Telli ML, Timms KM, Reid J, Hennessy B, Mills GB, Jensen KC, et al. Homologous recombination deficiency (HRD) score predicts response to platinum-containing neoadjuvant chemotherapy in patients with triple-negative breast cancer. Clin Cancer Res. 2016;22:3764–73.
Timms KM, Abkevich V, Hughes E, Neff C, Reid J, Morris B, et al. Association of BRCA1/2 defects with genomic scores predictive of DNA damage repair deficiency among breast cancer subtypes. Breast Cancer Res. 2014;16:475.
Masuda N, Bando H, Yamanaka T, Kadoya T, Takahashi M, Nagai S, et al. Eribulin-based neoadjuvant chemotherapy for triple-negative breast cancer patients stratified by homologous recombination deficiency status: a multicenter randomized phase II clinical trial. Breast Cancer Res Treat. 2021;188:117–31.
Langfelder P, Horvath S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinforma. 2008;9:559.
Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J Royal Stat Soc Ser B (Methodol). 1995;57:289–300.
von Mering C, Huynen M, Jaeggi D, Schmidt S, Bork P, Snel B. STRING: a database of predicted functional associations between proteins. Nucleic Acids Res. 2003;31:258–61.
Szklarczyk D, Gable AL, Lyon D, Junge A, Wyder S, Huerta-Cepas J, et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019;47:D607–13.
Szklarczyk D, Gable AL, Nastou KC, Lyon D, Kirsch R, Pyysalo S, et al. The STRING database in 2021: customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res. 2021;49:D605–12.
Pedregosa F, Varoquaux G, Gramfort A, Michel V, Thirion B, Grisel O, et al. Scikit-learn: machine learning in Python. J Machine Learn Res. 2011;12:2825–30.
Dranoff G. Cytokines in cancer pathogenesis and cancer therapy. Nat Rev Cancer. 2004;4:11–22.
Yu J, Lin C, Huang J, Hong J, Gao W, Zhu S, et al. Efficacy of adjuvant chemotherapy stratified by age and the 21-gene recurrence score in estrogen receptor-positive breast cancer. BMC Cancer. 2021;21:707.
Targowski T, Jahnz-Rozyk K, Plusa T, Glodzinska-Wyszogrodzka E. Influence of age and gender on serum eotaxin concentration in healthy and allergic people. J Investig Allergol Clin Immunol. 2005;15:277–82.
Araujo JM, Gomez AC, Aguilar A, Salgado R, Balko JM, Bravo L, et al. Effect of CCL5 expression in the recruitment of immune cells in triple negative breast cancer. Sci Rep. 2018;8:4899.
Baker KJ, Houston A, Brint E. IL-1 family members in cancer; two sides to every story. Front Immunol. 2019;10:1197.
Cardoso Alves L, Corazza N, Micheau O, Krebs P. The multifaceted role of TRAIL signaling in cancer and immunity. FEBS J. 2021;288:5530–54.
Moller C, Alfredsson J, Engstrom M, Wootz H, Xiang Z, Lennartsson J, et al. Stem cell factor promotes mast cell survival via inactivation of FOXO3a-mediated transcriptional induction and MEK-regulated phosphorylation of the proapoptotic protein Bim. Blood. 2005;106:1330–6.
Richmond J, Tuzova M, Cruikshank W, Center D. Regulation of cellular processes by interleukin-16 in homeostasis and cancer. J Cell Physiol. 2014;229:139–47.
Bartee E, McFadden G. Cytokine synergy: an underappreciated contributor to innate anti-viral immunity. Cytokine. 2013;63:237–40.
Mestan J, Brockhaus M, Kirchner H, Jacobsen H. Antiviral activity of tumour necrosis factor. Synergism with interferons and induction of oligo-2’,5’-adenylate synthetase. J Gen Virol. 1988;69:3113–20.
Bartee E, Mohamed MR, Lopez MC, Baker HV, McFadden G. The addition of tumor necrosis factor plus beta interferon induces a novel synergistic antiviral state against poxviruses in primary human fibroblasts. J Virol. 2009;83:498–511.
Modi S, Jacot W, Yamashita T, Sohn J, Vidal M, Tokunaga E, et al. Trastuzumab deruxtecan in previously treated HER2-low advanced breast cancer. New Engl J Med. 2022;387:9–20.
Schmid P, Cortes J, Dent R, Pusztai L, McArthur H, Kummel S, et al. Event-free survival with pembrolizumab in early triple-negative breast cancer. New Engl J Med. 2022;386:556–67.
Silver DP, Livingston DM. Mechanisms of BRCA1 tumor suppression. Cancer Discov. 2012;2:679–84.
Turner N, Tutt A, Ashworth A. Hallmarks of ‘BRCAness’ in sporadic cancers. Nat Rev Cancer. 2004;4:814–9.
Dunphy G, Flannery SM, Almine JF, Connolly DJ, Paulus C, Jonsson KL, et al. Non-canonical activation of the DNA sensing adaptor STING by ATM and IFI16 mediates NF-kappaB signaling after nuclear DNA damage. Mol Cell. 2018;71:745–60.e745.
Pantelidou C, Sonzogni O, De Oliveria Taveira M, Mehta AK, Kothari A, Wang D, et al. PARP inhibitor efficacy depends on CD8(+) T-cell recruitment via intratumoral STING pathway activation in BRCA-deficient models of triple-negative breast cancer. Cancer Discov. 2019;9:722–37.
Parkes EE, Walker SM, Taggart LE, McCabe N, Knight LA, Wilkinson R, et al. Activation of STING-dependent innate immune signaling by S-phase-specific DNA damage in breast cancer. J Natl Cancer Inst. 2017;109:djw199.
Min A, Kim K, Jeong K, Choi S, Kim S, Suh KJ, et al. Homologous repair deficiency score for identifying breast cancers with defective DNA damage response. Sci Rep. 2020;10:12506.
Mego M, Cholujova D, Minarik G, Sedlackova T, Gronesova P, Karaba M, et al. CXCR4-SDF-1 interaction potentially mediates trafficking of circulating tumor cells in primary breast cancer. BMC Cancer. 2016;16:127.
Acknowledgements
We gratefully acknowledge the work of the members of JBCRG and Kyoto University, who helped with this study. This study was based on a physician-led clinical trial funded by the Japan Breast Cancer Research Group and Eisai Co., Ltd.
Funding
None.
Author information
Authors and Affiliations
Contributions
KK, SK, NM and MToi designed the study. NM, HK, TY, SI, NT, TA, MTakada and ES provided specimens. KK, ST, YF, YM, TN and RMV performed sample preparation. DPS performed WGCNA statistical analyses, STRING DB analysis, and logistic regression analysis, constructed figures, and wrote the manuscript. SK, NM, TK, KI, TU, SOgawa, SOno, SM, MToi, and KK supervised the data analyses. SK and KK constructed figures and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
NM: grants from Chugai, Eli Lilly, AstraZeneca, Pfizer, Daiichi-Sankyo, MSD, Eisai, Novartis, Sanofi, Kyowa-Kirin, and Nippon-Kayaku; honoraria from Chugai, Pfizer, AstraZeneca, Eli Lilly, and Eisai; executive board and board of directors of JBCRG. TY: honoraria from AstraZeneca, Daiichi-Sankyo, Eli Lilly, Novartis, Chugai, Eisai, Kyowa-Kirin, and Pfizer. SI: honoraria from Eisai, Taiho, Chugai, Kyowa-Kirin, Daiichi-Sankyo, and Nippon-Kayaku. NT: grant from Eisai; honoraria from Chugai, Pfizer, Eli Lilly, Novartis, Eisai, and AstraZeneca KK. TA: honoraria from Chugai, Pfizer, Eli Lilly, and AstraZeneca KK. MT: grants from Daiichi-Sankyo, AstraZeneca KK, Yakult, KBCRN, ABCSG, JBCRG and Medbis; honoraria from Chugai, Eisai, Daiichi-Sankyo, Eli Lilly, AstraZeneca, Pfizer, Nippon-Kayaku, and Mitaka Koki. TU: honoraria from Eisai Co., Ltd., Chugai Pharmaceutical Co., Ltd., AstraZeneca, Novartis Pharma. SO: grants from Takeda Foundation, ChordiaTherapeutics, Inc., Sumitomo Dainippon Pharma Co., Ltd., Otsuka Pharmaceutical Co., Ltd., Eisai Co., Ltd.; Royalties from ASAHI Genomics Inc, Qiagen and Cordia Therapeutics; Consulting fees from KAN Research Institute, Inc., ChordiaTherapeutics, Inc., Eisai Co., Ltd., Novartis Pharmaceuticals; honoraria from MSD Japan, Kyowa Hakko Kirin Co., Ltd., Pfizer Inc; Stock or stock options from Stock of ASAHI Genomics Inc, Stock potion of Cordia Therapeutics Inc., and Stock of Rebirthell Inc. SO: honoraria from Eli Lilly, Chugai, Pfizer, and Eisai. SM: grants from Nippon Boehringer Ingelheim, Eisai; honoraria from AstraZeneca, Taiho Pharmaceutical Co. Ltd., Chugai Pharmaceutical, Nippon Boehringer Ingelheim, Bristol Myers Squibb Co. Ltd., Novartis Pharma KK, MSD KK, Kyowa-Kirin Co. Ltd., Astellas Pharma, Pfizer Japan, Ono Pharmaceutical Co. Ltd., Eli Lilly Japan. KK: grants from TERUMO, Astellas, Eli Lilly, Kyoto Breast Cancer Research Network; consulting fees from Becton Dickinson Japan; honoraria from Eisai, Chugai, and Takeda. MT: grants from Chugai, Takeda, Pfizer, Kyowa-Kirin, Taiho, JBCRG Associates, Eisai, Eli Lilly, Daiichi-Sankyo, AstraZeneca, Astellas, Shimadzu, Yakult, Nippon-Kayaku, AFI Technology, Luxonus, Shionogi, and GL Science; honoraria from Chugai, Takeda, Pfizer, Kyowa-Kirin, Taiho, Eisai, Daiichi-Sankyo, AstraZeneca, Eli Lilly, MSD, Exact Science, Novartis, Konica Minolta, Shimadzu, Yakult, and Nippon-Kayaku; advisory board of Kyowa-Kirin, Daiichi-Sankyo, Eli Lilly, Konica Minolta, BMS, Athenex Oncology, Bertis, Terumo, Kansai Medical Net; board of directors of JBCRG Associates, KBCRN, OOTR; associate editor of the British Journal of Cancer, Scientific Reports, Breast Cancer Research and Treatment, Cancer Science, Frontiers in Women’s Cancer, Asian Journal of Surgery, Asian Journal of Breast Surgery; deputy editor of International Journal of Oncology.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Saldajeno, D.P., Kawaoka, S., Masuda, N. et al. Time-series blood cytokine profiles correlate with treatment responses in triple-negative breast cancer patients. Br J Cancer 130, 1023–1035 (2024). https://doi.org/10.1038/s41416-023-02527-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-023-02527-0