Abstract
Cropping systems have fertilized soils for decades with undetermined consequences for the productivity and functioning of terrestrial ecosystems. One of the critical unknowns is the role of soil biodiversity in controlling crop production after decades of fertilization. This knowledge gap limits our capacity to assess how changes in soil biodiversity could alter crop production and soil health in changing environments. Here, we used multitrophic ecological networks to investigate the importance of soil biodiversity, in particular, the biodiversity of key-stone taxa in controlling soil functioning and wheat production in a 35-year field fertilization experiment. We found strong and positive associations between soil functional genes, crop production and the biodiversity of key-stone phylotypes; soils supporting a larger number of key-stone nematode, bacteria and fungi phylotypes yielded the highest wheat production. These key-stone phylotypes were also positively associated with plant growth (phototrophic bacteria, nitrogen fixers) and multiple functional genes related to nutrient cycling. The retrieved information on the genomes clustered with key-stone bacterial phylotypes indicated that the key-stone taxa had higher gene copies of oxidoreductases (participating most biogeochemical cycles of ecosystems and linking to microbial energetics) and 71 essential functional genes associated with carbon, nitrogen, phosphorus, and sulfur cycling. Altogether, our work highlights the fundamental role of the biodiversity of key-stone phylotypes in maintaining soil functioning and crop production after several decades of fertilization, and provides a list of key-stone phylotypes linking to crop production and soil nutrient cycling, which could give science-based guidance for sustainable food production.
Similar content being viewed by others
Introduction
A growing body of experimental and observational literature is providing evidence that soil biodiversity plays essential roles in regulating multiple ecosystem services, including nutrients cycling, organic matter decomposition, plant productivity, and pathogen control in natural terrestrial ecosystems [1, 2]. The contribution of soil biodiversity to crop production and soil health is much less clear, especially after decades of nutrient fertilization. In the agricultural ecosystem, fertilizers are widely used to promote crop production, potentially reducing the importance of soil biodiversity for maintaining crop production. Over the last decades, fertilizations have vastly increased to keep track with human’s growing demand for food [3]. We also know that fertilization can alter the diversity and composition of soil organisms [4], which might break the positive associations between soil biodiversity and plant production in long-term fertilized croplands. These effects also vary across different types of fertilizations [5]. Advancing our knowledge on the role and importance of soil biodiversity for controlling soil processes and plant productivity in croplands is integral to understand if changes in soil biodiversity with on-going global change can alter crop production limiting our capacity to feed a continuously growing human population.
Soil food-webs support a wide range of multitrophic communities with contrasting life styles, including predators, preys, probiotics, or pathogens [6]. These taxa thrive in the bulk soil, but critically important, also in the rhizosphere interface wherein soil, plant and microbes support a hotspot of biological activity [7]. These multitrophic communities includes bacteria, fungi, and soil invertebrates such as nematodes, which are essential bioindicators of soil processes in agroecosystem [8]. Ecological network provides critical information on the potential associations among thousands of soil organisms, and can be used to identify soil key-stone phylotypes associated with the highest levels of soil functioning and crop production [9, 10]. The importance of the biodiversity of soil key-stone phylotypes within specific ecological clusters has been recently highlighted as a major driver for ecosystem functioning in natural ecosystems across global biomes [1, 10]. To put it simple, in natural ecosystems, soils having a larger number of these fundamental taxa, also have higher nutrient availability, higher plant productivity, organic matter decomposition and pathogen control. Despite these recent discoveries, very little is known yet about how the biodiversity of key-stone phylotypes contributes to maintaining soil processes and crop production in agricultural ecosystems. This is especially important after decades of fertilization—a situation that is being faced by a large percentage of the terrestrial ecosystems across the globe. Surprisingly, we are still lacking a list of microbial key-stone phylotypes highly associated with the functioning and productivity of agroecosystems. Similarly, we are lacking information on the important functional genes associated with these soil phylotypes and their association with plant productivity. Learning more about the key-stone phylotypes supporting plant production is essential to maintain high crop production in a more managed world.
To advance our knowledge on the importance of the biodiversity of key-stone phylotypes for supporting crop production, here, we used multitrophic ecological networks to investigate the contributions of soil key-stone taxa to support soil functions and crop production in a 35-year-long fertilization experiment. Our work was based on wheat (Triticum asdtivum L.), which is one of the most commercially important cereal crop globally, and accounts for about a quarter of total food production in China [11]. We hypothesized that soil biodiversity, and especially the biodiversity of key-stone taxa is still a fundamental driver of plant and soil functioning after 4 decades of fertilization, and therefore, worth to be conserved. The reason is that soil biodiversity, not only plays a fundamental role in delivering available nutrients such as nitrogen (N) and phosphorus (P), but also in controlling organic matter decomposition, pathogen control, and the release of micro-nutrients and plant growth hormones.
Materials and methods
Experimental design and sample collection
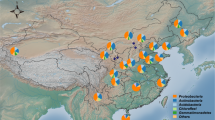
The fertilization experiment was established in 1982 with typical wheat–soybean crop rotations, in Mengcheng County (33°13′N, 116°35′E, 42 m elevation), Anhui Province, China. The soil type is the lime concretion black soil, annual precipitation is 872 mm and annual mean temperature is 14.8 °C. There are five treatments with four replicates (each plot area is 70 m2) in a completely randomized design (Supplementary Fig. S1): (1) Control: plots with no fertilizer added; (2) NPK: plots with chemical fertilizers comprising urea (180 kg N/ha/year), superphosphate (90 kg P2O5/ha/year) and potassium chloride (86 kg K2O/ha/year); (3) NPK+WS: plots with NPK chemical fertilizers plus wheat straw (about 7500 kg/hm2); (4) NPK+PM: plots with NPK chemical fertilizers plus pig manure (about 15,000 kg/hm2); (5) NPK+CM: NPK chemical fertilizers plus cow manure (about 30,000 kg/hm2). All the organic fertilizers and chemical fertilizers were annually added and fully mixed with soil before wheat planting. The wheat yield of recent 4 years (from 2014 to 2017) in this experiment was collected in Supplementary Table S1(a). Long-term NPK chemical fertilization increased the wheat yield by c. 10-fold when compared with the long-term unfertilized field (Control) (Supplementary Table S1). Consequently, wheat yield data in Control were detected as outliers in all analyses aiming to identify the association between microbial taxa and crop yield (Supplementary Fig. S2). Given that this is the main focus of our paper, we only used crop yield data (n = 32) of the NPK, NPK+WS, NPK+PM, and NPK + CM treatments in two soil compartments (bulk and rhizosphere soils) for further analyses.
During wheat filling stage (20th of the April, 2017), we randomly exacted 30–40 wheat individuals in each replicate filed, and gently shook off the loosely adhered soil, then collected rhizosphere soil (RS) by brushing the soils, which was tightly associated with wheat roots. Meanwhile, the topsoil (0–15 cm deep) was collected as bulk soil (BS) using an auger corer (~20 cm away from the plants). Then soils were passed through 2 mm meshes to remove the impurities (roots, stones etc.) before half of the samples were stored at −40 °C and 4 °C, prior to subsequent DNA extraction and physicochemical analysis.
Soil physicochemical analysis
Soil pH, moisture, total carbon (TC), total nitrogen (TN), total phosphorus (TP), total potassium (TK), dissolved organic carbon (DOC), nitrate (NO3−-N), ammonium (NH4+-N), dissolved organic nitrogen (DON), available phosphorus (AP) and available potassium (AK) were determined according to the methods described in Fan et al. [12]. Treatment effects and compartment variations (BS versus RS) of physicochemical properties were described in the Supplementary Table S2 and Table S3.
DNA extraction and high-throughput sequencing
Soil microbial DNA was extracted from 0.5 g of fresh soil collected from the long-term fertilized field by using the Fast DNA SPIN Kit (MP Biomedicals, Santa Ana, CA) following manufacturer’s instructions. Bacterial 16S rRNA genes targeting primer pairs were 515F (5′GTGCCAGCMGCCGCGGTAA-3′)/907R (5′-CCGTCAATTCCTTTGAGTTT-3′) [13]; the fungal ITS1-targeting primer pairs were ITS1F (5′-CTTGGTCATTTAGAGGAAGTAA-3′)/2043R (5′-GCTGCGTTCTTCATCGATGC-3′) [14]; and primer pairs AMV4.5NF (5′-AAGCTCGTAGTTGAATTTCG-3′)/AMDGR (5′-CCCAACTATCCCTATTAATCAT-3′) were used to amplify soil 18S rRNA gene fragments for the arbuscular mycorrhizal fungi (AMF) region [15]. Power Soil DNA kit (MO BIO Laboratories, Carlsbad, CA, USA, 10 g) was used to extract the total DNA from 10 g fresh soil for amplifying the Nematode 18S genes using the paired primers NF-1 (GGTGGTGCATGGCCGTTCTTAGTT)/18Sr2b (TACAAAGGGCAGGGACGTAAT) [16]. The sequences of bacteria and fungi have been submitted to the NCBI Sequence Read Archive (SRA) with accession number SRP126794 (https://www.ncbi.nlm.nih.gov/sra/SRP126794); the sequences of AMF have been submitted to the NCBI with accession number SRP218272 (https://trace.ncbi.nlm.nih.gov/Traces/sra/?study=SRP218272); the sequences of nematode have been submitted to the NCBI with accession number SRP218273 (https://trace.ncbi.nlm.nih.gov/Traces/sra/?study=SRP218273).
Sequence analysis
Quantitative Insight into Microbial Ecology (QIIME-1.9.1) pipeline (http://qiime.sourceforge.net/) were used to analyze the sequence data [17]. The high-quality sequences were acquired after removing the low-quality sequences (average quality score <25 and sequence length <200 bp or did not match the primer and barcode) and discarding the chimeric sequences. UCLUST was used to cluster the remaining high-quality sequences (bacteria: 1,320,400; fungi: 1,160,021; AMF: 620,620; nematode: 200,057) into operational taxonomic units (OTUs) at 97% similarity [18] by running pick_open_reference_otus.py work flow (http://qiime.org/scripts/pick_open_reference_otus.html), and then all singleton OTUs were deleted. The green genes 16S 13_8 database (http://greengenes.lbl.gov/) was used to assign the taxonomic identity of each phylotype of bacteria [19]; fungal taxonomic identity was determined using the UNITE database [20]; sequences of AMF were blasted against the MaarjAM Glomeromycota database [21]; the Silva_132 database was used to assign the taxonomic identity of phylotypes for nematode [22]. Finally, we got 36,773 OTUs for the bacteria, 1715 OTUs for the fungi, 685 OTUs for the AMF, and 938 OTUs for the nematode (Supplementary Tables S4, S5, S6, S7, S8, and S9).
The functional structures of fungal community in each treatment were further analyzed, the functional traits of fungal phylotypes were obtained from the online application FUNGuild (http://www.stbates.org/guilds/app.php) and literature [23, 24]. Of note, in order to reach the high accuracy, we kept the confidence ranking of “Highly Probable” and “Probable”, and only focused on the guilds of potential plant pathogen, given their relevance to plant health and potential risks of the agricultural ecosystems (Supplementary Fig. S3 and Supplementary Table S10).
High-throughput qPCR
Seventy-one functional genes [e.g., carbon (C), nitrogen (N), phosphorus (P), and sulfur (S) cycling] were analyzed by high-throughput qPCR on a Wafergen Smart Chip Real-Time qPCR platform as described by Wang et al. [25]. There were 90 DNA samples in total, and each DNA sample was amplified by PCR in triplicate at a threshold cycle of 31 as the detection limit. The same gene was regarded as only a single unique functional gene when targeted by multiple primer sets [26]. The relative abundances (gene copies normalized to the 16S rRNA gene) of functional genes for each sample were calculated as described by Pfaffl [27] (Supplementary Tables S11and S12).
Ecological clusters construction
“WGCNA” R package was used to construct the co-occurrence network based on the spearman correlation matrix [28]. We focused on microbial phylotypes (the operational taxonomic units or OTUs) that accounted for more than 80% of the relative abundance for bacteria, fungi, AMF, and nematode community, respectively. The bacterial, fungal, AMF, and nematode OTUs were merged into an abundance table, which includes 1452 bacterial OTUs, 383 fungal OTUs, 169 AMF OTUs, and 303 nematode OTUs (Supplementary Table S13). Then all pairwise spearman correlations between OTUs were calculated, and P-values were adjusted by Benjamini and Hochberg false discovery rate (FDR) test [29], then the adjusted P-values had a 0.001 cutoff. We focused on the microbial taxa that only presented high correlations with each other, which indicated that they are strongly co-occurring within communities, and therefore are more likely to interact with each other within soil food-webs [30]. Ecological clusters represent important ecological units that provide the opportunity to identify the highly connected and identifiable taxa [31, 32]. An interactive platform “Gephi” (default parameters set) was used to identify the modules (ecological clusters) of soil taxa strongly interacting with each other. We calculated the relative abundance of each module by averaging the standardized relative abundances (z-score) of the phylotypes within each ecological cluster (Supplementary Fig. S4 and Supplementary Appendix S1). Dominant phylotypes were chosen from the ecological clusters with relative abundance > 0.05, and strong spearman’s r > 0.8, significant P < 0.001 (Supplementary Appendix S1).
Bacterial whole-genomes downloading and annotation
We used BLAST to compare the representative sequences of chosen dominant bacterial OTUs (with >97% identity across the sequenced region of the 16S rRNA gene) to the 500 publicly available finished genomes (from rhizosphere; soil; and rhizoplane samples) in JGI-IMG website (https://img.jgi.doe.gov/). There were 47 genomes (Supplementary Table S14) with ssu rRNA 16S marker genes could be aligned and built in a phylogenetic tree, within which, 14, 19, 14 genomes could be clustered with the dominant OTUs from Module #0–2, respectively. These 47 whole genomes were downloaded from ftp://ftp.ncbi.nlm.nih.gov/genomes/genbank/bacteria/, “Prokka” pipeline was used for gene prediction and annotation (Supplementary Table S15).
Statistical analysis
ANOVA and Pairwise t-tests were used to compare the significant differences between fertilization treatments in the soil variables, the relative abundance of dominant soil microbial taxa, alpha diversity, the abundance of functional genes, the normalized gene copy numbers, and the relative abundance of ecological clusters, which were implemented in IBM SPSS 21 (Chicago, IL: Amos Development Corporation). Non-metric multidimensional scaling (NMDS) analysis and permutational multivariate analysis (ADONIS) were calculated in “vegan” R package. We calculated all pairwise spearman correlations between biodiversity, soil functional potentials [normalized abundance of 71 functional genes associated with C, N, P, S cycling (z-score)] and crop yield (Supplementary Table S16). The relationships between soil functional potentials, ecological clusters, and crop yield were then tested by linear regressions (Fig. 1 and Supplementary Figs. S5, S6, S7, S8, S9, and S10). The effects of the biodiversity of key-stone phylotypes (e.g., Chloroflexi, Nitrospirae, Mesorhizobium) and the abundance of functional genes [e.g., chiA (C degradation), amoB (N cycling), phoD (P cycling)] on crop yield in the long-term different fertilizations were also tested by linear regressions.
a Network diagram with nodes colored according to each of the four main ecological clusters (Modules #0–3); b Operational taxonomic unit (OTU) number properties of the dominant phylotypes in the main ecological clusters; c The regression relationships between the soil biodiversity of key-stone ecological cluster (Module #0) and soil functional potentials; d The regression relationships between the soil biodiversity of Module #0 and wheat crop yield. Crop yield data (n = 32) of NPK, NPK+WS, NPK+PM, and NPK+CM treatments in two soil compartments [bulk soil (BS) and rhizosphere soil (RS)] was used.
Structural equation modeling
The structural equation modeling (SEM) was used to evaluate the direct and indirect effects of prominent factors: fertilization treatments, edaphic variables (soil pH, TN, TP, and TC), rhizosphere effect, the diversity of total bacteria, fungi, AMF, nematode, the biodiversity of key-stone ecological cluster, and soil functional potentials on the crop production. Five fertilization treatments (Control; NPK; NPK+WS; NPK+PM; NPK+CM) were categorical variables with two levels: 1 (a particular treatment) and 0 (remaining considered treatments); the rhizosphere was categorical variables with two levels: 1 (rhizosphere soil samples) and 0 (bulk soil samples). As a few of variables were not normally distributed, bootstrapping was combined to test the probability that a path coefficient differs from zero. The standardized total effects (STEs) were calculated to aid interpretation of the SEM, for the effects of fertilization treatments, edaphic variables, rhizosphere effect, the diversity of total bacteria, fungi, AMF, nematode, the biodiversity of key-stone bacteria, fungi, AMF, nematode, and soil functional potentials on the crop production (Supplementary Fig. S11). SEM offers the ability to investigate the complex relationships among the biodiversity of different key-stone phylotypes, soil functional potentials, and crop yield in the long-term fertilization experiment. The biodiversity of key-stone bacteria, fungi, AMF, and Nematode was included as a composite variable in the SEM model, which together determined functional potentials and crop production. These composite variables did not alter the underlying SEM model, instead, they collapsed the effects of multiple conceptually related variables into a single composite effect, which could aid the interpretation of model results [1]. All the SEM analyses were conducted in IBM SPSS Amos 21 (Chicago, IL: Amos Development Corporation).
Results
Microbial communities and functional genes in long-term different fertilizations
Non-metric multidimensional scaling (NMDS) analysis (Supplementary Fig. S12) showed that the communities of bacteria, fungi, AMF, and nematode were significantly different among different fertilization regimes, which was confirmed by the permutational multivariate analysis (ADONIS) (Supplementary Table S17). The variations in bacteria, fungi, AMF, and nematode communities among different fertilization regimes were more significant than those between bulk soil and rhizosphere soil (Supplementary Table S18). For the bacterial community, the long-term application of NPK chemical fertilizers decreased the relative abundance of Actinobacteria, Bacteroidetes, Chloroflexi, Deltaproteobacteria, and the addition of organic matters (especially cow manure) increased their relative abundance. For the fungal community, the long-term application of chemical fertilizers decreased the relative abundance of Sordariomycetes, and the addition of pig manure increased the relative abundance of Pezizomycetes (Supplementary Table S4). Dothideomycetes was highly enriched in the rhizosphere soil in all the treatments (Supplementary Table S5). For the AMF community, the long-term fertilization increased the relative abundance of Claroideoglomeraceae, while decreased the relative abundance of Glomeraceae, particularly in the rhizosphere soil (Supplementary Table S4). For the nematode community, long-term fertilization increased the relative abundance of the dominant genera Caenorhabditis, while decreased most of the other genus belonging to Bacterivores, Fungivores, Herbivores, Omnivores-Predators (Supplementary Table S6). In addition, long-term application of chemical fertilizers decreased the diversity of bacteria and fungi, and the addition of cow manure increased their diversity. Meanwhile, the long-term application of different fertilizers decreased the diversity of AMF and nematodes (Supplementary Table S8).
For the functional genes, the long-term application of chemical fertilizers decreased the abundance of cdh and chiA genes associated with C degradation, and the abundance of nifH gene associated with N fixation. The abundance of most functional genes associated with C, N, P, and S cycling increased with the combination of NPK chemical fertilizers and cow manure (Supplementary Table S11). Meanwhile, multiple functional genes associated with the C (acsA, acsB, acsE, mct, rbcl), N (e.g., ureC, hao, nifH, nosZ1, nosZ2), P (phnk, phoD, phoX, ppx), S (yedZ, dsrA) cycles were enriched in the rhizosphere soil (Supplementary Table S12 and Supplementary Fig. S13). Soil functional potentials increased with the application of different fertilizers, particularly in the NPK+CM treatment (Supplementary Fig. S14), and the dissimilarity of the abundance of functional genes between rhizosphere and bulk soils was highly increased by the combined applications of NPK+organic matter (Supplementary Fig. S15). The soil functional potentials were significantly associated with the wheat crop production in both bulk and rhizosphere soils (Supplementary Figs. S5 and S6).
Key-stone ecological cluster linking to soil functions and crop production
We used ecological networks to identify clusters of microbial taxa highly correlated with each other, and therefore, potentially sharing environmental preferences and functional potentials. We detected four main ecological clusters (Module #0–3) divided from the multitrophic network (Fig. 1a, b). Over these modules, we found that the richness of phylotypes of bacteria, fungi, AMF, and nematode belonging to Module #0 was significantly and positively correlated with the abundance of soil functional genes, and more importantly, with the levels of crop production after four decades of fertilization (Fig. 1c and Supplementary Fig. S7). In other words, we found higher plant production and higher abundance of plant-growth promoting microbial functional genes in soils with more phylotypes belonging to Module #0. Given the functional importance of this module, we refer to Module#0 as the key-stone ecological cluster hereafter. The key-stone ecological cluster was dominated by bacteria (e.g., Acidobacteria; Alphaproteobacteria: Phenylobacterium, Bradyrhizobium; Betaproteobacteria: Burkholderia and Janthinobacterium; Chloroflexi) and fungi (Sordariomycetes: Lecythophora), and contained many plant-growth promoting phylotypes. A complete list with soil taxa within this module can be found in Supplementary Tables S13 and S19. In particular, there were many positive correlations (density = 0.32) among these key-stone phylotypes (Fig. 2a), and their relative abundance was highly and positively correlated with the abundance of multiple functional genes (Fig. 2b).
a A connection stands for a strong (spearman’s r > 0.8) and significant (P < 0.001) correlation. The co-occurrence network is colored by phylum, class, or genus. The size of each node is proportional to the relative abundance of each phylotype, and the thickness of each connection between two nodes (that is, edge) is proportional to the value of Spearman’s correlation coefficients. A blue edge indicates a positive correlation between two individual nodes, while a red edge indicates a negative correlation. The numbers inside the colored nodes are as follows: red1: the dominant bacterivores Caenorhabditis, red2: bacterivores Pratylenchus, red3: bacterivores Panagrolaimus; yellow1: fungivores Pristionchus, yellow2: fungivores Aphelenchoides, yellow3: fungivores Aphelenchus; green1: omnivores Aporcelaimellus, green2: omnivores Achromadora, green3: omnivores Ottolenchus. b Spearman correlations between the abundance of functional genes and the relative abundance of dominant phylotypes from Module #0–2 (z-score).
Further phylogenetic analysis showed that most of the key-stone phylotypes (dominant phylotypes in Module #0) tend to be phylogenetically clustered within specific bacterial phyla (e.g., Candidatus Solibacter and Candidatus Koribacter from phylum Acidobacteria and Rhodoplanes, Bradyrhizobium, and Rhizobium from phylum Alphaproteobacteria) when compared with those dominant taxa in other ecological clusters (Fig. 3a). We further obtained whole genomic information for available phylotypes in our ecological network and found that the normalized per-genome gene copy numbers of 71 functional genes associated with C, N, P, and S cycling were significantly higher in the genomes clustered within Module #0 than the genomes clustered within Module #1 or Module #2 (Fig. 3b). We also found that the normalized per-genome gene copy numbers of oxidoreductases, which participate in most biogeochemical cycles of ecosystems and link to microbial energetics [33] were higher in genomes clustered within Module #0 than those clustered within Module #1 or Module #2 (Fig. 3b).
a Phylogenetic tree of dominant OTUs from Module #0–2 and 47 whole genomes (with >97% identity across the sequenced region of the 16S rRNA gene) downloading from JGI-IMG websites (https://img.jgi.doe.gov/). Black bar: relative abundance of each OTU. Square (□) on the tree: ssu rRNA 16S marker genes of the whole genomes. Cyan ( ): Module #0; Magenta (
): Module #0; Magenta ( ): Module #1; Green (
): Module #1; Green ( ): Module #2. b The normalized per-genome gene copy numbers of 71 genes associated with carbon, nitrogen, phosphorus, sulfur cycling, and oxidoreductases genes. The normalized gene copy numbers are shown as the percentage (%) of genes per genome. Diamond symbol in each box plot represents the mean value.
): Module #2. b The normalized per-genome gene copy numbers of 71 genes associated with carbon, nitrogen, phosphorus, sulfur cycling, and oxidoreductases genes. The normalized gene copy numbers are shown as the percentage (%) of genes per genome. Diamond symbol in each box plot represents the mean value.
Unlike what we found for the key-stone ecological cluster, dominant phylotypes from Module #1 included more potential plant pathogens (Fig. 2a and Supplementary Fig. S3 and Supplementary Table S10), and there were more negative correlations within Module #1 when compared with other modules (Fig. 2a). The relative abundance of dominant phylotypes within Module #1 also presented less significant direct correlations with the abundance of functional genes (Fig. 2b). The sequence numbers of potential plant pathogens Acremonium significantly increased in the long-term NPK chemical fertilizations, but decreased in the NPK+PM and NPK+CM treatments; while the sequence numbers of pathogen-tolerance Flavisolibacter and pathogen-suppressing Pseudomonas significantly increased in the NPK+PM and NPK+CM treatments (Supplementary Table S20). In addition, most of the dominant phylotypes from Module #2 were attributed to the bacteria and AMF, within which, AMF taxa tended to be correlated with Betaproteobacteria (Burkholderia, Methylibium), Actinobacteria (Streptomyces, Amycolatopsis, and Frankia) (Fig. 2a and Supplementary Table S19); and these dominant taxa in Module #2 also presented less direct correlations with the abundance of functional genes (Fig. 2b). The associations between other modules and crop production and functional potential can be found in Supplementary Figs. S8, S9, and S10.
Ecological relationships between biodiversity of key-stone phylotypes, functional genes, and crop production
SEM analysis provided further statistical evidence that the diversity of key-stone phylotypes is essential to maintain plant production and the soil functional potentials. We found multiple direct and indirect associations between fertilizations, rhizosphere, soil properties, biodiversity (bacteria, fungi, AMF, nematode), soil functional potentials, and crop production (Fig. 4). SEM of biodiversity with total phylotypes, which explained 75% variations in wheat yield, showed that the diversity of bacteria was positively associated with crop production, while the diversity of fungi and AMF was negatively correlated with crop production (Supplementary Fig. S11A). SEM of the biodiversity of key-stone phylotypes, which explained 79% variations in crop production, showed that the diversity of key-stone bacteria, fungi, and AMF was directly and positively associated with wheat yield (Fig. 4 and Supplementary Fig. S11B). In particular, the diversity of key-stone nematode was indirectly associated with the crop production by positively correlating soil functional potentials (Fig. 4).
The direct and indirect effects of different fertilizations, rhizosphere effects, edaphic variables [soil pH, total carbon (TC), total nitrogen (TN), total phosphorous (TP)], diversity of total bacteria, fungi, AMF, nematode, biodiversity of key-stone ecological cluster, and soil functional potentials [normalized abundance of 71 functional genes associated with carbon, nitrogen, phosphorus, and sulfur cycling (z-score)] on crop production. Arrow width is proportional to the strength of the relationship. Solid line: positive correlation; dashed line: negative correlation. Numbers above/below the arrow lines are indicative of the correlations. The proportion of explained variance (R2) appears alongside wheat production in the model. χ2, Chi-square; df, degrees of freedom; P, probability level; RMSEA, root-mean squared error of approximation are the goodness-of-fit statistics for each model. Significance levels of each predictor are *P < 0.05, **P < 0.01. Control: non-fertilization; NPK: applications of nitrogen (N), phosphorous (P), and potassium (K); NPK+WS: NPK with wheat straw; NPK+PM: NPK with pig manure; NPK+CM: NPK with cow manure.
We further selected a handful of examples to illustrate the link between genetic function, key-stone taxa, crop yield and imposed treatments (Fig. 5). Specifically, we found that the abundance of chiA gene (endochitinase; bifunctional chitinase/lysozyme) was significantly corelated with the biodiversity of key-stone Chloroflexi in Module #0; the abundance of amoB gene significantly linked to the biodiversity of key-stone Nitrosospira in Module #0; and the abundance of phoD gene was significantly associated with the biodiversity of key-stone Mesorhizobium in Module #0 (Fig. 5). The biodiversity of these key-stone phylotypes (e.g., Chloroflexi, Nitrospirae, Mesorhizobium) and the abundance of functional genes (e.g., chiA, amoB, phoD) were significantly associated with crop production (Fig. 5).
Selected examples for the links between biodiversity of key-stone phylotypes (e.g., Chloroflexi, Nitrospirae, Mesorhizobium in Module #0) and the abundance of functional genes (e.g., chiA, amoB, phoD) on crop yield in the long-term different fertilizations. Data (n = 32) of NPK, NPK+WS, NPK+PM, and NPK+CM treatments in two soil compartments [bulk soil (BS) and rhizosphere soil (RS)] was used.
Discussion
Our work used multitrophic ecological networks to identify a list of soil key-stone phylotypes, and the biodiversity of key-stone phylotypes largely determines soil functions and crop production after almost four decades of different fertilization. Moreover, the retrieved information on the genomes clustered with key-stone bacterial phylotypes, provided evidence that these key-stone taxa had higher normalized per-genome gene copy numbers of oxidoreductases and 71 functional genes associated with C, N, P, and S cycling. These findings highlighted the importance of biodiversity of key-stone phylotypes within multitrophic communities in maintaining soil functioning and wheat production, providing the possibility to improve crop production by regulating microbial key-stone taxa in the agroecosystem. Moreover, this knowledge is important because it opens the door to isolate and apply these microbial communities aiming to improve crop production to feed a continue growing human population worldwide.
The biodiversity of key-stone phylotypes (e.g., Chloroflexi, Nitrospirae, Mesorhizobium) was positively associated with the abundance of functional genes (e.g., chiA, amoB, phoD) (Fig. 5). Chloroflexi have been found to be positively correlated with genes for amino sugars, sugar alcohols and simple carbohydrate metabolic pathways [34]; Nitrosospira belonged to one ammonia-oxidizing bacterial group harboring amoA and amoB genes [35]; and Mesorhizobium was found to be the main phylotypes containing phoD gene [36]. The solid relationships between the biodiversity of these key-stone phylotypes (e.g., Chloroflexi, Nitrospirae, Mesorhizobium) and the abundance of functional genes (e.g., chiA, amoB, phoD) indicated that the increase in functional gene abundance probably resulted from the increase in biodiversity of specific key-stone phylotypes. We further found that the biodiversity of multitrophic key-stone phylotypes could indirectly and directly affect crop production (Fig. 4). For example, the biodiversity of key-stone nematodes could indirectly affect crop production by regulating soil functional potentials through the entrance of new resources via litter decomposition into the soil food-webs [37]. The biodiversity of key-stone bacteria, fungi, and AMF could directly affect the crop production. The key-stone ecological cluster (Module #0) contained the least potential plant pathogens (Supplementary Fig. S13), and most of the key-stone phylotypes could be attributed to the multiple plant growth promoting microbes. For instance, the key-stone taxa Rhodoplanes and Chloroflexi belongs to phototrophic bacteria, which could participate in degrading plant compounds via pathways for the degradation of starch, pyrogallol, cellulose, and long-chain sugars [38]; Rhizobium, Bradyrhizobium and Burkholderia are N fixers [39], which are closely associated with N cycling; and Janthinobacterium are well-known for their antifungal effects [40]. Meanwhile, the normalized per-genome gene copy numbers of functional genes associated with C, N, P and S cycling were higher in genomes clustered with key-stone bacterial phylotypes, suggesting a higher nutrient cycling potential of these key-stone taxa. As redox-based characterization could participate in most biogeochemical cycles, and linking to the microbial energetics of ecosystems [33], the higher normalized per-genome gene copy numbers of oxidoreductases for the key-stone phylotypes, suggesting their relatively comprehensive higher functional strategies. NPK+cow manure fertilization treatment matched with the highest diversity of the key-stone phylotypes, suggesting that these key-stone microbial taxa can be promoted via adequate agricultural management practices.
Unlike the Module #0 (key-stone phylotypes), Module #1 included multiple potential fungal pathogens such as Acremonium (fungal endophytes which could infect the plants) [41], pathogen-tolerance Flavisolibacter (which could survive with indigenous pathogen) [42], and pathogen-suppressing Pseudomonas [43]. The diversity of Module #1 was directly and negatively associated with crop production. As with the diversity of Module #1, the total fungal diversity was directly and negatively associated with wheat production (Fig. 4), which probably resulted from the effect of potential fungal pathogens [44]. The decreasing potential pathogens Acremonium and increasing pathogen-suppressing Pseudomonas in the NPK+CM treatment indicated that certain long-term fertilization practices could suppress the pathogens by motivating more beneficial and tolerating microorganisms. In addition, Module #2 included many bacteria-AMF correlations, within which, most AMF taxa tended to be correlated with bacterial phylotypes of Betaproteobacteria (Burkholderia, Methylibium) [45] and Actinobacteria (Streptomyces, Amycolatopsis, and Frankia) [46], which could segment and form aerial hyphae. It has been found that Burkholderia and Methylibium could be associated with mycorrhizal roots [47, 48]; Streptomyces and Amycolatopsis could also form aerial hyphae and may also segment, and Frankia is the only N-fixing actinobacterium, which has the ability to enter into symbiotic associations with diverse plants. It is indicated that the bacterial communities living associated with extraradical hyphae or spores, and plant roots could positively affect the beneficial activity of AMF [47]. The decreasing diversity and abundance of these AMF and bacterial phylotypes under long-term different fertilization indicated that the enriched nutrients in the soil decreased the effectivity of AMF and their associated microorganisms.
For the functional genes, the abundance of nifH gene decreased with the long-term application of NPK chemical fertilizers, while increased with the addition of organic matter, particularly in the NPK+CM treatment. In a previous study [12], we found that the long-term application of NPK chemical fertilizers could suppress N fixation and the associated diazotrophic communities, while the addition of organic fertilizers (e.g., cow manure) could alleviate the suppression of chemical fertilizers on the abundance or diversity of N fixers and the associated genes. In addition, we identified a list of C fixation genes (e.g., acsA, acsB, acsE, mct, and rbcl) contributing to the accumulation of soil organic matter [26], multiple N cycling genes (e.g., nifH, ureC, hao, nosZ1, and nosZ2) regulating pivotal processes of N fixation or nitrification denitrification [49], and P cycling genes (e.g., phoD and phoX) participating in the process of organic phosphate mineralization [50], which were highly enriched in the rhizosphere soil, particularly in the NPK+CM treatment, suggesting that the improvement of soil fertility (e.g., soil organic matter) could provide a more suitable environment for microbial growth and activity, which could contribute to the availability of nutrient cycling [51] and further increase crop production [52, 53].
Conclusions
This study provides solid evidence that the biodiversity of key-stone phylotypes is essential for supporting plant production and soil functions after almost 4 decades of chemical and organic fertilizations—a scenario which affects agroecosystems from across the globe. In particular, the diversity of key-stone phylotypes could contribute to the crop yield by positively regulating multiple microbial functional genes [C (acsA, acsB, acsE, mct, rbcl), N (ureC, hao, nifH, nosZ1, nosZ2), P (phnk, phoD, phoX, ppx), S (yedz, dsrA) cycles] associated with plant growth and nutrient cycling. In addition, the combined application of chemical fertilizers and organic manure (e.g., cow manure) could suppress the pathogens and activate more beneficial and tolerant microorganisms, and the enriched nutrients decrease the effectivity of AMF and their associations with specific bacteria. These findings highlighted the importance of biodiversity of key-stone phylotypes in maintaining soil functional potentials and crop production, suggesting the possibility to improve crop production by mediating soil microbial key-stone taxa in intensified agricultural ecosystems.
References
Delgado-Baquerizo M, Reich PB, Trivedi C, Eldridge DJ, Abades S, Alfaro FD, et al. Multiple elements of soil biodiversity drive ecosystem functions across biomes. Nat Ecol Evol. 2020;4:210–20.
Delgado-Baquerizo M, Maestre FT, Reich PB, Jeffries TC, Gaitan JJ, Encinar D, et al. Microbial diversity drives multifunctionality in terrestrial ecosystems. Nat Commun. 2016;7:10541.
Shah F, Wu W. Soil and crop management strategies to ensure higher crop productivity within sustainable environments. Sustainability. 2019;11:1485.
Leff JW, Jones SE, Prober SM, Barberán A, Borer ET, Firn JL, et al. Consistent responses of soil microbial communities to elevated nutrient inputs in grasslands across the globe. Proc Natl Acad Sci USA. 2015;112:10967.
Sun R, Zhang X-X, Guo X, Wang D, Chu H. Bacterial diversity in soils subjected to long-term chemical fertilization can be more stably maintained with the addition of livestock manure than wheat straw. Soil Biol Biochem. 2015;88:9–18.
Mougi A, Kondoh M. Diversity of interaction types and ecological community stability. Science. 2012;337:349–51.
Kumar A, Patel JS, Meena VS. Rhizospheric microbes for sustainable agriculture: an overview. In: Meena VS, editor. Role of rhizospheric microbes in soil: volume 1: stress management and agricultural sustainability. Singapore: Springer Singapore; 2018. p. 1–31.
Yeates GW, Bongers T. Nematode diversity in agroecosystems. In: Paoletti MG, editor. Invertebrate biodiversity as bioindicators of sustainable landscapes. Amsterdam: Elsevier; 1999. p. 113–35.
Chaffron S, Rehrauer H, Pernthaler J, von Mering C. A global network of coexisting microbes from environmental and whole-genome sequence data. Genome Res. 2010;20:947–59.
Banerjee S, Schlaeppi K, van der Heijden MGA. Keystone taxa as drivers of microbiome structure and functioning. Nat Rev Microbiol. 2018;16:567–76.
Cai G, Chen D, Ding H, Pacholski A, Fan X, Zhu Z. Nitrogen losses from fertilizers applied to maize, wheat and rice in the North China Plain. Nutr Cycl Agroecosys. 2002;63:187–95.
Fan K, Delgado-Baquerizo M, Guo X, Wang D, Wu Y, Zhu M, et al. Suppressed N fixation and diazotrophs after four decades of fertilization. Microbiome. 2019;7:143.
Biddle JF, Fitz-Gibbon S, Schuster SC, Brenchley JE, House CH. Metagenomic signatures of the Peru Margin subseafloor biosphere show a genetically distinct environment. Proc Natl Acad Sci USA. 2008;105:10583–8.
Bokulich NA, Mills DA. Improved selection of internal transcribed spacer-specific primers enables quantitative, ultra-high-throughput profiling of fungal communities. Appl Environ Microbiol. 2013;79:2519–26.
Lumini E, Orgiazzi A, Borriello R, Bonfante P, Bianciotto V. Disclosing arbuscular mycorrhizal fungal biodiversity in soil through a land-use gradient using a pyrosequencing approach. Environ Microbiol. 2010;12:2165–79.
Porazinska D, Giblin-Davis, Robin M, Faller LF, William K, Natsumi M, et al. Evaluating high-throughput sequencing as a method for metagenomic analysis of nematode diversity. Mol Ecol Resour. 2009;9:1439–50.
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, et al. QIIME allows analysis of high-throughput community sequencing data. Nat Methods. 2010;7:335.
Edgar RC. Search and clustering orders of magnitude faster than BLAST. Bioinformatics. 2010;26:2460–1.
McDonald D, Price MN, Goodrich J, Nawrocki EP, DeSantis TZ, Probst A, et al. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J. 2011;6:610.
Kõljalg U, Larsson K-H, Abarenkov K, Nilsson RH, Alexander IJ, Eberhardt U, et al. UNITE: a database providing web-based methods for the molecular identification of ectomycorrhizal fungi. N. Phytol. 2005;166:1063–8.
Öpik M, Vanatoa A, Vanatoa E, Moora M, Davison J, Kalwij JM, et al. The online database MaarjAM reveals global and ecosystemic distribution patterns in arbuscular mycorrhizal fungi (Glomeromycota). N Phytol. 2010;188:223–41.
Quast C, Pruesse E, Gerken J, Peplies J, Yarza P, Yilmaz P, et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 2012;41:D590–D6.
Nguyen NH, Song Z, Bates ST, Branco S, Tedersoo L, Menke J, et al. FUNGuild: an open annotation tool for parsing fungal community datasets by ecological guild. Fungal Ecol. 2016;20:241–8.
Dean R, Kan JALV, Pretorius ZA, Hammond‐Kosack KE, Pietro AD, Spanu PD, et al. The Top 10 fungal pathogens in molecular plant pathology. Mol Plant Pathol. 2012;13:804.
Wang F-H, Qiao M, Su J-Q, Chen Z, Zhou X, Zhu Y-G. High throughput profiling of antibiotic resistance genes in urban park soils with reclaimed water irrigation. Environ Sci Technol. 2014;48:9079–85.
Zheng B, Zhu Y, Sardans J, Peñuelas J, Su J. QMEC: a tool for high-throughput quantitative assessment of microbial functional potential in C, N, P, and S biogeochemical cycling. Sci China Life Sci. 2018;61:1451–62.
Pfaffl MW. A new mathematical model for relative quantification in real-time RT–PCR. Nucleic Acids Res. 2001;29:e45–e45.
Langfelder P, Horvath S. Fast R functions for robust correlations and hierarchical clustering. J Stat Softw. 2012;46:11.
Benjamini Y, Krieger AM, Yekutieli D. Adaptive linear step-up procedures that control the false discovery rate. Biometrika. 2006;93:491–507.
Hines J, van der Putten WH, De Deyn GB, Wagg C, Voigt W, Mulder C, et al. Chapter four-Towards an integration of biodiversity–ecosystem functioning and food web theory to evaluate relationships between multiple ecosystem services. Adv Ecol Res. 2015;53:161–99.
Menezes AB, Prendergast-Miller MT, Richardson AE, Toscas P, Farrell M, Macdonald LM, et al. Network analysis reveals that bacteria and fungi form modules that correlate independently with soil parameters. Environ Microbiol. 2015;17:2677–89.
Heleno R, Devoto M, Pocock M. Connectance of species interaction networks and conservation value: Is it any good to be well connected? Ecol Indic. 2012;14:7–10.
Ramírez-Flandes S, González B, Ulloa O. Redox traits characterize the organization of global microbial communities. Proc Natl Acad Sci USA. 2019;116:3630.
Pérez Castro S, Cleland EE, Wagner R, Sawad RA, Lipson DA. Soil microbial responses to drought and exotic plants shift carbon metabolism. ISME J. 2019;13:1776–87.
Zhang C, Song Z, Zhuang D, Wang J, Xie S, Liu G. Urea fertilization decreases soil bacterial diversity, but improves microbial biomass, respiration, and N-cycling potential in a semiarid grassland. Biol Fert Soils. 2019;55:229–42.
Fraser TD, Lynch DH, Bent E, Entz MH, Dunfield KE. Soil bacterial phoD gene abundance and expression in response to applied phosphorus and long-term management. Soil Biol Biochem. 2015;88:137–47.
García-Palacios P, Shaw EA, Wall DH, Hättenschwiler S. Temporal dynamics of biotic and abiotic drivers of litter decomposition. Ecol Lett. 2016;19:554–63.
Hug LA, Castelle CJ, Wrighton KC, Thomas BC, Sharon I, Frischkorn KR, et al. Community genomic analyses constrain the distribution of metabolic traits across the Chloroflexi phylum and indicate roles in sediment carbon cycling. Microbiome. 2013;1:22.
Lu J, Yang F, Wang S, Ma H, Liang J, Chen Y. Co-existence of rhizobia and diverse non-rhizobial bacteria in the rhizosphere and nodules of dalbergia odorifera seedlings inoculated with Bradyrhizobium elkanii, Rhizobium multihospitium–like and Burkholderia pyrrocinia–like strains. Front Microbiol. 2017;8:2255.
Haack FS, Poehlein A, Kröger C, Voigt CA, Piepenbring M, Bode HB, et al. Molecular keys to the janthinobacterium and duganella spp. interaction with the plant pathogen Fusarium graminearum. Front Microbiol. 2016;7:1668.
Clay K, Leuchtmann A. Infection of woodland grasses by fungal endophytes. Mycologia. 1989;81:805–11.
Huang X, Liu L, Wen T, Zhang J, Wang F, Cai Z. Changes in the soil microbial community after reductive soil disinfestation and cucumber seedling cultivation. Appl Microbiol Biotechnol. 2016;100:5581–93.
Palleroni NJ. Pseudomonas. In: M.E. Trujillo, S. Dedysh, P. DeVos, B. Hedlund, P. Kämpfer, F.A. Rainey and W.B. Whitman, editors. Bergeyʼs Manual of Systematics of Archaea and Bacteria. John Wiley & Sons, Inc. in association with Bergey’s Manual Trust; 2015. p. 1–105.
Wei Z, Yang T, Friman V-P, Xu Y, Shen Q, Jousset A. Trophic network architecture of root-associated bacterial communities determines pathogen invasion and plant health. Nat Commun. 2015;6:8413.
Mao Y, Li X, Smyth EM, Yannarell AC, Mackie RI. Enrichment of specific bacterial and eukaryotic microbes in the rhizosphere of switchgrass (Panicum virgatum L.) through root exudates. Environ Microbiol Rep. 2014;6:293–306.
Barka EA, Vatsa P, Sanchez L, Gaveau-Vaillant N, Jacquard C, Klenk H-P, et al. Taxonomy, physiology, and natural products of actinobacteria. Microbiol Mol Biol Rev. 2016;80:1.
Agnolucci M, Battini F, Cristani C, Giovannetti M. Diverse bacterial communities are recruited on spores of different arbuscular mycorrhizal fungal isolates. Biol Fert Soils. 2015;51:379–89.
Levy A, Merritt AJ, Mayo MJ, Chang BJ, Abbott LK, Inglis TJJ. Association between Burkholderia species and arbuscular mycorrhizal fungus spores in soil. Soil Biol Biochem. 2009;41:1757–9.
Li X, Rui J, Xiong J, Li J, He Z, Zhou J, et al. Functional potential of soil microbial communities in the maize rhizosphere. PLoS ONE. 2014;9:e112609.
Ragot SA, Kertesz MA, Mészáros É, Frossard E, Bünemann EK. Soil phoD and phoX alkaline phosphatase gene diversity responds to multiple environmental factors. FEMS Microbiol Ecol. 2016;93:fiw212.
Gianfreda L. Enzymes of importance to rhizosphere processes. J Soil Sc Plant Nutr. 2015;15:283–306.
Su J-Q, Ding L-J, Xue K, Yao H-Y, Quensen J, Bai S-J, et al. Long-term balanced fertilization increases the soil microbial functional diversity in a phosphorus-limited paddy soil. Mol Ecol. 2015;24:136–50.
Ratliff TJ, Fisk MC. Phosphatase activity is related to N availability but not P availability across hardwood forests in the northeastern United States. Soil Biol Biochem. 2016;94:61–9.
Acknowledgements
We thank Ruibo Sun, Jianqiang Su, Xinyuan Zhou, Zhibin Guo, Keke Hua, Yingying Ni, Yuying Ma, Hongfei Wang, Liang Cheng, Yu Shi, Teng Yang, and Yuntao Li for their assistance in field management, soil sampling, and laboratory work. This work was funded by the National Natural Science Foundation of China (31870480), the Strategic Priority Research Program of the Chinese Academy of Sciences (XDB15010101), and the China Biodiversity Observation Networks (Sino BON). MD-B is supported by a Ramón y Cajal grant from the Spanish Government (agreement no. RYC2018-025483-I).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Fan, K., Delgado-Baquerizo, M., Guo, X. et al. Biodiversity of key-stone phylotypes determines crop production in a 4-decade fertilization experiment. ISME J 15, 550–561 (2021). https://doi.org/10.1038/s41396-020-00796-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-020-00796-8
This article is cited by
-
Meta-analysis reveals the effects of microbial inoculants on the biomass and diversity of soil microbial communities
Nature Ecology & Evolution (2024)
-
Vertical differences in carbon metabolic diversity and dominant flora of soil bacterial communities in farmlands
Scientific Reports (2024)
-
Dynamic root microbiome sustains soybean productivity under unbalanced fertilization
Nature Communications (2024)
-
Soil carbon storage and accessibility drive microbial carbon use efficiency by regulating microbial diversity and key taxa in intercropping ecosystems
Biology and Fertility of Soils (2024)
-
Green manuring relocates microbiomes in driving the soil functionality of nitrogen cycling to obtain preferable grain yields in thirty years
Science China Life Sciences (2024)