Abstract
Background
This study aimed to identify key microRNAs (miRNAs), pathways, and target genes mediating Hirschsprung’s disease (HSCR) pathogenesis and identify the diagnostic potential of miRNAs.
Methods
The Gene Expression Omnibus database and reverse transcription-quantitative PCR were used to compare miRNA expression between ganglionic and aganglionic colon tissues of children with HSCR, and the TAM 2.0 database was used to identify colon tissue-specific miRNAs. The StarBase database, TargetScan database, luciferase reporter, and western blot assays were used to analyze miRNA–messenger RNA interactions. OmicShare was used to perform functional and pathway enrichment analyses of the target genes. Migration assays were performed to validate the functions of the miRNAs.
Results
The TAM 2.0 database analysis and reverse transcription-quantitative PCR showed that hsa-miR-192-5p, hsa-miR-200a-3p, and hsa-miR-200b-3p were colon tissue-specific and upregulated in aganglionic colon tissue compared to paired ganglionic colon tissue. These three miRNAs effectively reduced cell viability and migration. Luciferase reporter and western blot assays verified the direct interaction between these three miRNAs and the target genes of ZEB2 and FNDC3B. Furthermore, the plasma levels of these miRNAs were higher in HSCR patients than in non-HSCR patients.
Conclusions
Three plasma miRNAs (hsa-miR-192-5p, hsa-miR-200a-3p, and hsa-miR-200b-3p) are potential peripheral HSCR biomarkers.
Impact
-
The molecular mechanisms underlying HSCR are unclear. HSCR is most accurately diagnosed using rectal biopsy samples, and no consensus has been reached on the use of blood-based tests for HSCR diagnosis. Circulating miRNAs may be candidate diagnostic HSCR biomarkers because they are typically easily detectable, stable, and tissue-specific. Three plasma miRNAs (miR-200a-3p, miR-192-5p, and miR-200b-3p) are potential peripheral HSCR biomarkers.
Similar content being viewed by others
Introduction
Hirschsprung’s disease (HSCR) is characterized by a deficit of ganglion cells in the distal bowel1 and is the most common enteric neuropathy. The annual incidence of HSCR is almost 1 in 5000 children, with a male sex ratio bias of 4:1.2 HSCR can be categorized into three types based on the length of the aganglionic segment: short-segment HSCR, long-segment HSCR, or total colon aganglionosis.3 The clinical symptoms of HSCR in children usually include abdominal distension and delayed meconium excretion.4 The treatment of children with HSCR is surgical extraction of the aganglionic bowel (AB); however, the long-term outcome remains poor.5 Although the specific pathogenesis of HSCR has not been elucidated, in the past few decades, some studies have confirmed that the cause of HSCR is associated with many genes, including RET, GDNF, NRG1, SIIP1, and PHOX2B.6,7,8,9,10,11 The diagnosis or exclusion of HSCR is typically dependent on clinical manifestations, anorectal manometry, contrast enema followed by radiology, and rectal biopsy.12 Even though these methods are highly accurate, misdiagnosis may occur during the neonatal period.13 Therefore, less invasive, high-throughput, blood-based tests are required to improve diagnostic accuracy. Previous findings have proven that microRNAs (miRNAs) are associated with the pathogenesis of HSCR.14,15
MiRNAs are noncoding endogenous RNAs (18–24 bp in length) that regulate target messenger RNA (mRNA) expression.16,17 Recent research has indicated that miRNAs are closely related to various cellular biological processes (BPs), including growth, cell death, inflammation, development, and differentiation.18 Other studies have shown that miRNAs may serve as biomarkers of pathological processes in different diseases,19,20 and several miRNAs, including miR-488-3p,21 miR-141,22 and miR-637,23 are associated with HSCR.
To elucidate the pathogenesis of HSCR and to identify potential new biomarkers, we screened miRNA expression profiles of paired aganglionic and ganglionic colon samples from children with HSCR obtained from the Gene Expression Omnibus (GEO) database.24 The miRNA expression profiles of HSCR samples obtained from the GEO database were then used as a query for a TAM 2.0 database search.25 The TAM 2.0 database contains several miRNAs that are colon tissue-specific. Based on the miRNA expression profiles, the BPs, cellular components (CCs), molecular functions (MFs), and enriched pathways were explored via enrichment analyses. Then, migration assays and reverse transcription-quantitative PCR (RT-qPCR), luciferase reporter, and western blot assays were used to validate the functions of the miRNAs and evaluate the diagnostic value of dysregulated miRNAs in the plasma of HSCR patients.
Materials and methods
Availability of miRNA expression profiles
MiRNA expression profiles deposited in a publicly available dataset (GSE77296) were obtained from the GEO database, a public networking tool that was used to compare the miRNA profiles of aganglionic and ganglionic colon samples from three children. The differentially expressed miRNAs between the aganglionic and ganglionic colon samples were analyzed according to the following criteria: P value < 0.01 and an absolute expression log 2 fold change (FC) > 1.5.
Analysis of miRNA–mRNA target networks
Based on TAM 2.0 database analysis, hsa-miR-192-5p, hsa-miR-200a-3p, and hsa-miR-200b-3p were considered colon tissue-specific genes. Identifying mRNA target genes is critical for determining the crucial functions and pathways associated with miRNAs. Thus, StarBase v2.0 was used to predict the target genes of these three miRNAs.26 In addition, miRNA–mRNA interaction networks were analyzed using Cytoscape and the TargetScan database.
Gene functional enrichment analyses
Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO) enrichment analyses of the target genes of these three miRNAs were carried out using the OmicShare datasets (enrichment was considered significant at P < 0.01) to identify the BPs, CCs, MFs, and pathways associated with the differentially expressed miRNAs.
Cell culture and transfection
HSCR pathogenesis is driven by defects in ganglion cells that are derived from neural crest cells.27 Neuroblastomas are a group of sympathetic ganglia tumors derived from primitive neural crest cells.28 Separation of primary enteric neurons from HSCR patients is difficult, and few neurons can be obtained.29 Due to the limitations of obtaining the primary enteric neurons of patients with HSCR and in accordance with previous research,30,31,32 human SH-SY5Y and SK-N-BE(2) cells, which are derived from neuroblastoma, were used in this study. Both cell lines were obtained from the American Type Culture Collection (Rockville, MD), grown in Dulbecco’s modified Eagle’s medium (Gibco, Carlsbad) supplemented with 10% fetal bovine serum and incubated at 37 °C with 5% CO2. In the transfection experiments, synthetic hsa-miR-200a-3p, hsa-miR-192-5p, and hsa-miR-200b-3p mimics (Tsingke Biotechnology, Wuhan, China; the sequences are shown in Table 1) were transfected using a RiboBio FECT™ CP Transfection Kit following the manufacturer’s guidelines.
Cell proliferation assay
Cell growth was assessed with CCK-8 Kit (Dojindo, Tokyo, Japan) following the manufacturer’s guidelines. Cell viability assays were conducted according to our previously described method.33 (The OD was detected at 450 nm.)
Migration assays
Scratch wound assay
Cells were seeded into 6-well plates (2 × 105 cells per well) and incubated until confluent. The middle of the well was scratched using a 100-μL pipette tip. The two cell lines were then cultured in a complete medium with or without miRNA mimics. Cells between the two borders of the migrating cell layer were photographed after 0, 24, and 48 h, and the width of the scratch was measured using the ImageJ software.34 The ratio of migration area was calculated as follows: migration area rate (%) = (A0 − An)/(A0 × 100), where A0 is the wound area at the 0 h time point and An is the wound area at the respective measurement time.
Transwell assay
The two cell lines were transfected with or without miRNA mimics. After transfection for 48 h, both cell lines were seeded in the upper chamber of the Transwell plate (Corning-Costar, Shanghai, China) containing a filter with 8-μm pore size. The assay was conducted according to our previously described method.33 The migrated cells on the bottom of each chamber membrane were counted manually using the ImageJ software.
Luciferase reporter assay
The TargetScan and StarBase databases were used to analyze the 3′-untranslated regions (UTRs) of ZEB2 and FNDC3B as potential binding sites of miR-192-5p, miR-200a-3p, and miR-200b-3p. The 3′-UTR sequences of ZEB2 and FNDC3B containing the miR-192-5p, miR-200a-3p, and miR-200b-3p and mutant 3′-UTR sequences were cloned into the psiCHECKTM-2 vector (Tsingke Biotechnology, Wuhan, China). SH-SY5Y and SK-N-BE(2) cells were transfected with a mixture of ZEB2 and FNDC3B wild type or mutant type and mimic NC, miR-192 mimics, miR-200a mimics, or miR-200b mimics for approximately 48 h. Then, the cells were harvested using a luminometer to measure the firefly and Renilla luciferase activities.
Western blot
Tissue or cellular proteins were extracted from 1× cell radioimmunoprecipitation assay lysis buffer (Beyotime, Wuhan, China). Western blotting was conducted according to our previously described method33 with antibodies against ZEB2 (A5705, ABclonal, Wuhan, China), FNDC3B (22605-1-AP, Proteintech, Wuhan, China), and β-actin (AC026, ABclonal, Wuhan, China).
Study design, participants, and sample material
The human tissue study was approved by the Ethics Committee of Tongji Medical College, Huazhong University of Science and Technology (approval number IORG0003571) and carried out in accordance with the Declaration of Helsinki. Whole-blood samples from 40 patients with diagnosed HSCR (30 boys and 10 girls) and 40 sex- and age-matched controls with no congenital disease were obtained from the Department of Pediatric Surgery, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, China. Written informed consent was obtained from the guardians of all participants. After surgery, HSCR diagnoses were confirmed by pathological examination and bowel samples were obtained. Exclusion criteria were established to prevent confounding effects of other congenital disorders. Blood samples were collected and placed in EDTA tubes before surgery. Serum was obtained by centrifugation and stored in RNase-free tubes in liquid nitrogen.
RT-qPCR
RT-qPCR was conducted according to our previously described method33 with the following PCR forward primers: hsa-miR-192-5p, 5′-CCTGACCTATGAATTGACAGCC-3′; hsa-miR-200a-3p, 5′-GGCTAACACTGTCTGGTAACGATGT-3′; hsa-miR-200b-3p, 5′-GGCTAATACTGCCTGGTAATGATGA-3′; U6: 5′-CTCGCTTCGGCAGCACA-3′; and the common reverse primer 5′-GTGCAGGGTCCGAGGT-3′. ZEB2 and FNDC3B primer sets were as follows: for human ZEB2, 5′-CCTCTGTAGATGGTCCAGTGA-3′ (forward) and 5′-GTCACTGCGCTGAAGGTACT-5′ (reverse); FNDC3B, 5′-ATAGCCAAGAGGTGGTGTGC-3′ (forward) and 5′-TACTCCACTGCAACGTGACC-3′ (reverse); beta-actin, 5′-GCACTCTTCCAGCCTTCCT-3′ (forward) and 5′-AGGTCTTTGCGGATGTCCAC-3′ (reverse).
Statistical analyses
The information was analyzed by using the χ2 test, unpaired Student’s t test, or repeated-measures analysis of variance. The data were conducted by GraphPad Prism 8.0 software. All experiments were performed in triplicate, and the results are displayed as the mean ± SD. Asterisk indicates P < 0.05 and NS indicates no significance. Exact P values are shown in Supplementary Table S2.
Results
Screening and identification of differentially expressed miRNAs and colon tissue-specific miRNAs in HSCR
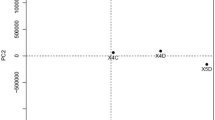
Mining of the RNA-sequencing data deposited in the GEO database (GSE77296) was performed to identify differential miRNA expression in aganglionic and GB tissue from three children. In total, 725 miRNAs with a log 2 FC > 1 and P < 0.01 were identified, of which 55 were downregulated and 106 were upregulated. A volcano plot was generated showing the expression of all downregulated and upregulated genes (Fig. 1a). To identify molecular markers relevant to HSCR, we screened colon tissue-specific miRNAs using the TAM 2.0 database, and hsa-miR-192-5p, hsa-miR-200a-3p, and hsa-miR-200b-3p were identified as colon tissue-specific. In addition, we further searched the GEO database (GSE77296), and the results showed that these three miRNAs were significantly increased in AB compared with paired GB (Fig. 1b).
a MiRNA expression profiles in paired normal and HSCR bowel samples are shown in the volcano plot; 55 miRNAs were downregulated and 106 were upregulated in aganglionic bowel tissue. b Mining of the public GEO database (GSE77296) revealing the miR-192-5p, miR-200a-3p, and miR-200b-3p transcription levels in HSCR ganglionic bowels and aganglionic bowels.
MiRNA–target gene regulatory network analysis
MiRNAs play major roles in regulating mRNA expression. The StarBase database was used to predict the target genes of these miRNAs, and Cytoscape was used to visualize the regulation of miRNA–mRNA interactions (Fig. 2). The target genes of these three miRNAs are listed in Appendix Table S1.
KEGG pathway and GO functional enrichment analysis of target genes
KEGG pathway and GO functional annotation analyses were performed with the target genes of the three miRNAs. After GO term enrichment analysis, 55 GO terms, including CC, MF, and BP terms, and KEGG pathways were identified, and we obtained the top 20 most significantly enriched terms (Fig. 3a–d). The following CC terms were mainly enriched in the target genes: cell, cell part, organelle part, and membrane (Fig. 3a). MF terms mainly enriched in the target genes included binding, catalytic activity, transcription regulator activity, and MF regulators (Fig. 3b). BP terms enriched in the target genes included positive regulation of cellular processes, biological regulation, regulation of BPs, and metabolic process (Fig. 3c). KEGG pathway analysis indicated that the pathways most enriched with the targets of the three upregulated miRNAs were microRNAs in cancer, axon guidance, ErbB signaling pathway, and focal adhesion (Fig. 3d).
MiRNA mimics inhibit cell proliferation and migration
We tested the effects of overexpressing the three miRNAs on SH-SY5Y and SK-N-BE(2) cell growth and migration. Stable transfection of the three miRNA mimics resulted in overexpression of the corresponding miRNAs in these two cell lines. In these two cell lines transfected with these three miRNA mimics, the CCK-8 assay (Fig. 4a) and Matrigel invasion (Fig. 4b) and scratch wound (Fig. 4c) assays revealed decreased cell proliferation and migration, respectively. These data showed that these three miRNAs play major roles in regulating cell proliferation and migration.
a CCK-8 assays indicating changes in the viability of cells transfected with mimic NC, the miR-192 mimic, the miR-200a mimic, and the miR-200b mimic after culture for 24, 48, 72, and 96 h. b Representative Transwell assays (left panel) and quantitative analysis of migrated cell numbers (right panel) after 24 h showing the migration capability of neuroblastoma cell lines transfected as indicated. c Scratch wound assays (left panel) and measurement of the wounded area for quantitative analysis of the migration of cells transfected as indicated; *P < 0.05 vs. mimic NC. Exact P values are shown in Supplementary Table S2.
MiR-192, miR-200a, and miR-200b modulate functions by inhibiting the target genes of ZEB2 and FNDC3B
To investigate how these three miRNAs regulate proliferation and migration, we first analyzed these three miRNA–target genes from the StarBase database. Based on the overlapping analysis of these three miRNA–target genes, we focused on ZEB2 and FNDC3B, both of which can interact with miR-192, miR-200a, and miR-200b (Fig. 5a). Next, we used the TargetScan database to predict binding sites of the 3′-UTR between these three miRNAs and two target genes (Supplementary Fig. S1). Dual-luciferase assays indicated that the relative activity was decreased in SH-SY5Y (Fig. 5b) and SK-N-BE(2) (Supplementary Fig. S2) cells transfected with miR-192-5p, miR-200a-3p, and miR-200b-3p mimics in wild-type 3′-UTR plasmids of ZEB2 and FNDC3B compared with the miRNA NC group. However, when transfected with the 3′-UTR mutant vector of ZEB2 and FNDC3B and these three miRNA mimics, there was no significant change in the dual-luciferase activity compared with the control group. Then, qRT-PCR (Fig. 5c) and western blot (Fig. 5d) assays revealed that transfection of these three miRNA mimics led to both the transcription and protein levels of ZEB2 and FNDC3B being downregulated in these two cell lines compared with miRNA NC. These findings indicated that ZEB2 and FNDC3B were direct target genes of miR-192-5p, miR-200a-3p, and miR-200b-3p.
a Venn diagram representing target genes of miR-192-5p, miR-200a-3p, and miR-200b-3p from the StarBase database, and overlapping analysis of results identified ZEB2 and FNDC3B target genes. b Dual-luciferase reporter assays showing the luciferase activity of wild-type or mutant ZEB2 and FNDC3B after transfection with miR-192-5p, miR-200a-3p, and miR-200b-3p mimics for 48 h in SH-SY5Y cells. c, d Real-time qRT-PCR and western blot assays showing the transcription and protein levels of ZEB2 and FNDC3B in these two cell lines transfected with mimic NC, miR-192-5p, miR-200a-3p, or miR-200b-3p. P < 0.05 vs. mimic NC. (Normalized to β-actin.) Exact P values are shown in Supplementary Table S2.
Upregulation of miRNA-192/-200a/-200b in AB segments and plasma of HSCR patients
To verify the microarray analysis results, we investigated the mRNA levels of these three mRNAs in 40 AB samples and paired GB samples of HSCR patients using RT-qPCR. The clinical information of the participants is summarized in Table 2. Consistent with the microarray results, RT-qPCR indicated that the three miRNAs were significantly upregulated in aganglionic tissue (Fig. 6a). Moreover, in HSCR tissue specimens, the transcription levels of ZEB2 and FNDC3B were negatively correlated with the expression of these three miRNAs (Fig. 6b). We further detected the expression of these three miRNA–target genes in Hirschsprung tissue specimens by western blot (Fig. 6c and Supplementary Fig. S3), and the results showed that ZEB2 and FNDC3B were significantly reduced in HSCR AB tissue compared with paired GB tissue. To determine the potential peripheral HSCR biomarkers, we analyzed the transcript levels of the three miRNAs in plasma from HSCR patients (n = 40) and controls (non-HSCR children, n = 40). The levels of these three miRNAs in plasma samples were highly expressed in children with HSCR compared with non-HSCR patients (Fig. 6d). These data indicated that these three plasma miRNAs could function as promising HSCR biomarkers.
a RT-qPCR analysis of these three miRNA transcripts was performed in 40 paired aganglionic and ganglionic bowel tissues; *P < 0.05 vs. ganglionic bowel tissues. b The correlation between these three miRNAs and their target genes ZEB2 and FNDC3B. c Western blot indicated protein levels of ZEB2 and FNDC3B in HSCR aganglionic bowel (AB) samples and paired ganglionic bowel (GB) tissues. d RT-qPCR measurement of miRNA levels in HSCR and non-HSCR plasma samples; *P < 0.05 vs. non-HSCR plasma samples. (Internal control: U6.) Exact P values are shown in Supplementary Table S2.
Discussion
HSCR is a digestive tract condition in neonates that manifests as incomplete colonization by neural crest-derived cells,35 and its diagnosis is based on a combination of clinical manifestations, contrast enema followed by radiology, anorectal manometry, and rectal biopsy.36,37 Diagnosis by rectal biopsy is most sensitive and specific, but requires examination of ganglion cells in serial sections of formalin-fixed tissue and acetylcholinesterase histochemistry using frozen sections.38 However, identifying ganglion cells in neonates can be difficult.39
The use of plasma miRNA has specific advantages for diagnostic purposes. MiRNAs originating in the bowel can cross the endothelium to communicate with the gut and distant organs through the bloodstream.40 MiRNAs are associated with numerous human pathologies, and blood-based miRNAs have become a potential source of diagnostic biomarkers.41 Therefore, these studies indicated the possibility of profiling tissue-specific miRNAs found in plasma.
To elucidate the possible roles of miRNAs in HSCR and find biomarkers for the early diagnosis of HSCR that can be evaluated by noninvasive blood-based testing, we compared the profiles of dysregulated miRNAs between AB and GB samples in HSCR from the public GEO (GSE77296) database, finding that 106 miRNAs were highly expressed and 55 miRNAs were downregulated in the AB relative to the GB. Analysis of the TAM 2.0 database showed that hsa-miR-200a, hsa-miR-192, and hsa-miR-200b are colon tissue-specific genes. GO and KEGG enrichment analyses indicated that the target genes of these miRNAs were enriched in various processes that may be associated with the proliferation and migration of neural crest-derived cells. Previous findings have shown that dysregulation of the ErbB pathway leads to developmental disorders of human gastrointestinal motility.42 These three miRNAs decreased cell growth and migration and could inhibit the enteric nervous system. In addition, our study discovered that these three miRNAs were significantly upregulated in aganglionic segment tissues and in plasma from patients with HSCR. These findings show that these three miRNAs could be used as noninvasive biomarkers. Previous research has shown that hsa-miR-192-5p participates in the pathophysiology of Crohn’s disease and colorectal cancer.43 Moreover, among the three miRNAs, hsa-miR-192-5p has been used to predict survival in patients with stage IIIB colon cancer.44 HSCR is a severe neonatal defect in which the distal bowel lacks ganglion cells.45 These three miRNAs were upregulated in aganglionic tissues and in plasma from patients with HSCR, suggesting that they may be used as biomarkers for HSCR. Through luciferase and western blot assays, the mechanism at the base of these three miRNAs reduced viability and migration through interaction with ZEB2 and FNDC3B. Our research indicated that ZEB2 and FNDC3B were downregulated in HSCR children with AB, and the inhibition of ZEB2 expression could reduce neuronal differentiation in HSCR.46 We further explored the mechanism of the increase in these three miRNAs through the TransmiR v2.0 database,47 which is a transcription factor-miRNA regulation database. We explored the transcription factors that coregulate these three miRNAs through the TransmiR v2.0 database, and the results showed that there were a total of 25 transcription factors (Supplementary Fig. S4a). Next, we selected transcription factors that were statistically significant and tissue-specific in the digestive tract, and only the transcription factors ATF2 and TFAP4 were identified (Supplementary Fig. S4b). Therefore, we speculate that the transcription factors ATF2 and TFAP4 regulate and upregulate three miRNAs in HSCR.
Despite these findings, this study has some limitations. First, the number of miRNA profiles of HSCR samples that we obtained from GSE77296 was small, resulting in bias when analyzing the miRNAs. Second, there are currently 40 specimens for verification, but this is far from sufficient, and more tissues and blood samples are needed to verify the results. Third, although the pathogenesis underlying the overexpression of these three miRNAs in the aganglionic colon suggests that they are related to the transcription factors ATF2 and TFAP4, further verification is needed.
In conclusion, by mining the miRNA expression profiles of HSCR bowel segments coupled with analysis of the TAM 2.0 database, we identified three colon tissue-specific miRNAs and explored their signaling pathways. Our data indicated that hsa-miR-192-5p, hsa-miR-200a-3p, and hsa-miR-200b-3p are upregulated in AB segments and in the plasma of patients with HSCR. The portfolio of HSCR biomarkers,6,9,10 including plasma miRNA biomarkers, should be used complementarily to provide blood-based testing approaches to facilitate the early diagnosis of HSCR. As these three miRNAs are promising HSCR markers with great diagnostic value, their identification represents an important step toward developing a noninvasive and low-risk diagnostic test.
References
Burkardt, D. D., Graham, J. J., Short, S. S. & Frykman, P. K. Advances in Hirschsprung disease genetics and treatment strategies: an update for the primary care pediatrician. Clin. Pediatr. 53, 71–81 (2014).
Soret, R. et al. A collagen VI-dependent pathogenic mechanism for Hirschsprung’s disease. J. Clin. Invest. 125, 4483–4496 (2015).
Jaroy, E. G. et al. “Too much guts and not enough brains”: (epi)genetic mechanisms and future therapies of Hirschsprung disease—a review. Clin. Epigenet. 11, 135 (2019).
Lof, G. A. et al. Maternal risk factors and perinatal characteristics for Hirschsprung disease. Pediatrics 138, e20154608 (2016).
Bettolli, M. et al. Colonic dysmotility in postsurgical patients with Hirschsprung’s disease. Potential significance of abnormalities in the interstitial cells of Cajal and the enteric nervous system. J. Pediatr. Surg. 43, 1433–1438 (2008).
Jain, S. et al. Organotypic specificity of key RET adaptor-docking sites in the pathogenesis of neurocristopathies and renal malformations in mice. J. Clin. Invest. 120, 778–790 (2010).
Miyamoto, R. et al. Loss of Sprouty2 partially rescues renal hypoplasia and stomach hypoganglionosis but not intestinal aganglionosis in Ret Y1062F mutant mice. Dev. Biol. 349, 160–168 (2011).
Sanchez, M. P. et al. Renal agenesis and the absence of enteric neurons in mice lacking GDNF. Nature 382, 70–73 (1996).
Cantrell, V. A. et al. Interactions between Sox10 and EdnrB modulate penetrance and severity of aganglionosis in the Sox10Dom mouse model of Hirschsprung disease. Hum. Mol. Genet. 13, 2289–2301 (2004).
Stanchina, L., Van de Putte, T., Goossens, M., Huylebroeck, D. & Bondurand, N. Genetic interaction between Sox10 and Zfhx1b during enteric nervous system development. Dev. Biol. 341, 416–428 (2010).
Van de Putte, T. et al. Mice lacking ZFHX1B, the gene that codes for Smad-interacting protein-1, reveal a role for multiple neural crest cell defects in the etiology of Hirschsprung disease-mental retardation syndrome. Am. J. Hum. Genet. 72, 465–470 (2003).
Heuckeroth, R. O. Hirschsprung disease—integrating basic science and clinical medicine to improve outcomes. Nat. Rev. Gastroenterol. Hepatol. 15, 152–167 (2018).
Ambartsumyan, L., Smith, C. & Kapur, R. P. Diagnosis of Hirschsprung disease. Pediatr. Dev. Pathol. 23, 8–22 (2020).
Sergi, C. M., Caluseriu, O., McColl, H. & Eisenstat, D. D. Hirschsprung’s disease: clinical dysmorphology, genes, micro-RNAs, and future perspectives. Pediatr. Res. 81, 177–191 (2017).
Shen, Z. et al. Microarray expression profiling of dysregulated long non-coding RNAs in Hirschsprung’s disease reveals their potential role in molecular diagnosis. Neurogastroenterol. Motil. 28, 266–273 (2016).
Huntzinger, E. & Izaurralde, E. Gene silencing by microRNAs: contributions of translational repression and mRNA decay. Nat. Rev. Genet. 12, 99–110 (2011).
Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
Maffioletti, E., Tardito, D., Gennarelli, M. & Bocchio-Chiavetto, L. Micro spies from the brain to the periphery: new clues from studies on microRNAs in neuropsychiatric disorders. Front. Cell. Neurosci. 8, 75 (2014).
Vishnoi, A. & Rani, S. MiRNA biogenesis and regulation of diseases: an overview. Methods Mol. Biol. 1509, 1–10 (2017).
Saliminejad, K., Khorram, K. H., Soleymani, F. S. & Ghaffari, S. H. An overview of microRNAs: biology, functions, therapeutics, and analysis methods. J. Cell. Physiol. 234, 5451–5465 (2019).
Wen, Z. et al. Circular RNA CCDC66 targets DCX to regulate cell proliferation and migration by sponging miR-488-3p in Hirschsprung’s disease. J. Cell. Physiol. 234, 10576–10587 (2019).
Tang, W. et al. Aberrant reduction of MiR-141 increased CD47/CUL3 in Hirschsprung’s disease. Cell. Physiol. Biochem. 32, 1655–1667 (2013).
Li, Y. et al. Long non-coding RNA FAL1 functions as a ceRNA to antagonize the effect of miR-637 on the down-regulation of AKT1 in Hirschsprung’s disease. Cell Prolif. 51, e12489 (2018).
Edgar, R., Domrachev, M. & Lash, A. E. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 30, 207–210 (2002).
Li, J. et al. TAM 2.0: tool for microRNA set analysis. Nucleic Acids Res. 46, W180–W185 (2018).
Li, J. H., Liu, S., Zhou, H., Qu, L. H. & Yang, J. H. starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res. 42, D92–D97 (2014).
Liu, J. A. et al. Identification of GLI mutations in patients with Hirschsprung disease that disrupt enteric nervous system development in mice. Gastroenterology 149, 1837–1848 (2015).
Olsen, R. R. et al. MYCN induces neuroblastoma in primary neural crest cells. Oncogene. 36, 5075–5082 (2017).
Anitha, M. et al. Characterization of fetal and postnatal enteric neuronal cell lines with improvement in intestinal neural function. Gastroenterology 134, 1424–1435 (2008).
Tang, W. et al. Multiple ‘omics’-analysis reveals the role of prostaglandin E2 in Hirschsprung’s disease. Free Radic. Biol. Med. 164, 390–398 (2021).
Wang, G. et al. Downregulation of microRNA-483-5p promotes cell proliferation and invasion by targeting GFRA4 in Hirschsprung’s disease. DNA Cell Biol. 36, 930–937 (2017).
Wu, F. et al. MPGES-1 derived PGE2 inhibits cell migration by regulating ARP2/3 in the pathogenesis of Hirschsprung disease. J. Pediatr. Surg. 54, 2032–2037 (2019).
Hong, M. et al. Runt-related transcription factor 1 promotes apoptosis and inhibits neuroblastoma progression in vitro and in vivo. J. Exp. Clin. Cancer Res. 39, 52 (2020).
Collins, T. J. ImageJ for microscopy. Biotechniques 43, 25–30 (2007).
Fattahi, F. et al. Deriving human ENS lineages for cell therapy and drug discovery in Hirschsprung disease. Nature 531, 105–109 (2016).
Guinard-Samuel, V. et al. Calretinin immunohistochemistry: a simple and efficient tool to diagnose Hirschsprung disease. Mod. Pathol. 22, 1379–1384 (2009).
Nabi, Z., Shava, U., Sekharan, A. & Nageshwar, R. D. Diagnosis of Hirschsprung’s disease in children: preliminary evaluation of a novel endoscopic technique for rectal biopsy. JGH Open 2, 322–326 (2018).
Moore, S. W. & Johnson, G. Acetylcholinesterase in Hirschsprung’s disease. Pediatr. Surg. Int. 21, 255–263 (2005).
Lv, X. et al. Molecular function predictions and diagnostic value analysis of plasma exosomal miRNAs in Hirschsprung’s disease. Epigenomics 12, 409–422 (2020).
Scaldaferri, F. et al. Gelatin tannate ameliorates acute colitis in mice by reinforcing mucus layer and modulating gut microbiota composition: emerging role for ‘gut barrier protectors’ in IBD? United Eur. Gastroenterol. J. 2, 113–122 (2014).
Leidinger, P., Backes, C., Meder, B., Meese, E. & Keller, A. The human miRNA repertoire of different blood compounds. BMC Genomics 15, 474 (2014).
Le, T. L. et al. Dysregulation of the NRG1/ERBB pathway causes a developmental disorder with gastrointestinal dysmotility in humans. J. Clin. Invest. 131, e146389 (2021).
Zhao, H. et al. miR-192/215-5p act as tumor suppressors and link Crohn’s disease and colorectal cancer by targeting common metabolic pathways: An integrated informatics analysis and experimental study. J. Cell. Physiol. 234, 21060–21075 (2019).
Li, P., Ou, Q., Braciak, T. A., Chen, G. & Oduncu, F. S. MicroRNA-192-5p is a predictive biomarker of survival for Stage IIIB colon cancer patients. Jpn J. Clin. Oncol. 48, 619–624 (2018).
Soret, R. et al. Glial cell-derived neurotrophic factor induces enteric neurogenesis and improves colon structure and function in mouse models of Hirschsprung disease. Gastroenterology 159, 1824–1838 (2020).
Watanabe, Y. et al. Differentiation of mouse enteric nervous system progenitor cells is controlled by endothelin 3 and requires regulation of Ednrb by SOX10 and ZEB2. Gastroenterology 152, 1139–1150 (2017).
Tong, Z., Cui, Q., Wang, J. & Zhou, Y. TransmiR v2.0: an updated transcription factor-microRNA regulation database. Nucleic Acids Res. 47, D253–D258 (2019).
Funding
This study was supported by the National Natural Science Foundation of China (81670511 and 81873848).
Author information
Authors and Affiliations
Contributions
M.H., X.L., and Y.L. carried out the RT-qPCR, CCK-8, western blot, luciferase reporter, and migration assays. X.L., Y.Z., Y.L., S.C., G.C., and S.L. helped with statistical analysis of the data. M.H. and S.T. designed the research and wrote the paper. The authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Hong, M., Li, X., Li, Y. et al. Hirschsprung’s disease: key microRNAs and target genes. Pediatr Res 92, 737–747 (2022). https://doi.org/10.1038/s41390-021-01872-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41390-021-01872-1