Abstract
Background
The S-adenosyl-methionine (SAM) availability is crucial for DNA methylation, an epigenetic mechanism involved in nonsyndromic cleft lip with or without cleft palate (NSCL/P) expression. The aim of this study was to assess the association between single-nucleotide polymorphisms (SNPs) of genes involved in SAM synthesis and NSCL/P in a Chilean population.
Methods
In 234 cases and 309 controls, 18 SNPs in AHCY, MTR, MTRR, and MAT2A were genotyped, and the association between them and the phenotype was evaluated based on additive (allele), dominant, recessive and haplotype models, by odds ratio (OR) computing.
Results
Three deep intronic SNPs of MTR showed a protective effect on NSCL/P expression: rs10925239 (OR 0.68; p = 0.0032; q = 0.0192), rs10925254 (OR 0.66; p = 0.0018; q = 0.0162), and rs3768142 (OR 0.66; p = 0.0015; q = 0.0162). Annotations in expression database demonstrate that the protective allele of the three SNPs is associated with a reduction of MTR expression summed to the prediction by bioinformatic tools of its potentiality to modify splicing sites.
Conclusions
The protective effect against NSCL/P of these intronic MTR SNPs seems to be related to a decrease in MTR enzyme expression, modulating the SAM availability for proper substrate methylation. However, functional analyses are necessary to confirm our findings.
Impact
-
SAM synthesis pathway genetic variants are factors associated to NSCL/P.
-
This article adds new evidence for folate related genes in NSCL/P in Chile.
-
Its impact is to contribute with potential new markers for genetic counseling.
Similar content being viewed by others
Introduction
Epigenetic modifications include both DNA and histone methylation.1 These modifications lead to differential gene expression-producing cells with diverse phenotypes from the same genome.2 Epigenetics also reflects the effects of a changing environment on an individual during his/her life, a phenomenon that has generated great interest in complex diseases, in considering the possibility that complex diseases may have causes unrelated to the genetic sequence.3 Thus, epigenetic changes could explain part of the heritability of complex traits where genome-wide association studies have only found small contributions from genetic variants.4
Nonsyndromic orofacial clefts (OFCs) are the most prevalent craniofacial birth defects in Chile and worldwide.5 These malformations include nonsyndromic cleft lip with or without cleft palate (NSCL/P), and nonsyndromic cleft palate not associated with other major malformations.6 Affected subjects present a wide variety of complications related not only to orofacial issues but also experience delays in cognitive development and a shorter lifespan, which is associated with an increased risk of certain cancers.7,8 NSCL/P is a complex trait where its susceptibility seems to depend on the interaction of genetic and environmental factors.6,9 NSCL/P etiology also hints at a missing heritability phenomenon where genetic variants explain only ~20% of its heritability.10 In the past decade, several studies have been performed assessing the possible role of epigenetics in NSCL/P etiology in humans, focusing on both global and specific regions DNA methylation from blood or oral tissues.11,12,13,14 In addition, evidence from animal models has demonstrated the role of histone methylation in the expression of OFCs.15,16,17
The methyl group donor for DNA, histone methylation, and more than 50 other methylation reactions, is S-adenosyl-methionine (SAM), which is used by several methyl-transferases at the nuclear and cytosolic levels.18 After the production of the methylated substrate, SAM becomes S-adenosyl-l-homocysteine (SAH).19 SAH is a strong inhibitor of SAM-dependent reactions, and is hydrolyzed to adenine and homocysteine (Hcy) by a reaction catalyzed by S-adenosyl-l-homocysteine hydrolase (SAHH).20 Hcy is also a negative regulator of methyl-transferases, and is re-methylated to methionine by methionine synthase (MS or MTR), which transfers a methyl group from 5-methyl-tetrahydrofolate, a reaction that uses methylcobalamin as a cofactor, derived from vitamin B12.21 In this reaction, the oxidized cofactor inactivates MTR, which is later regenerated by MTRR enzyme.22 Finally, methionine is converted to SAM by methionine adenosyltransferase (MAT) in an ATP-dependent reaction.23 In addition to its synthesis pathway enzymes, SAM levels depend on the intake of methyl donors, such as folates and vitamin B12.18
Variants in genes in the aforementioned SAM synthesis pathway have been associated with the risk of NSCL/P in several populations.24,25,26,27,28,29,30,31 In addition, these phenotypes are related to the maternal consumption of folates and vitamin B12 during the periconceptional period.32 In Chile, our group has reported that a functional variant in the MTHFR gene (codes for the enzyme-producing 5-methyl-tetrahydrofolate) is a risk factor for NSCL/P.33 The purpose of this study was to assess the association between single-nucleotide polymorphism (SNP) variants of the genes involved in the SAM synthesis pathway and NSCL/P in a Chilean population, based on a case–control design. To achieve this purpose, our report includes the genes encoding the enzymes SAHH (AHCY, 20q11.22), MS (MTR, 1q43), MTRR (MTRR, 5p15.31), and MAT (MAT2A, 2q11.2).
Methods
Subjects
Our case sample was composed of 234 unrelated NSCL/P subjects. This group contained 38.0% females, and 70% had no family history of OFCs. These cases were recruited from 2017 to 2019 from the following centers in Santiago, Chile: The Craniofacial Malformations Unit, School of Dentistry, Universidad de Chile; The Cleft Lip/Palate Center, Hospital Exequiel Gonzalez Cortes; The Dental Service, Hospital Roberto del Rio, and The Maxillofacial Service, Hospital San Borja-Arriaran. The control group was composed of 309 unrelated individuals, and does not significantly differ in the proportion of sexes as compared to cases (37.2% females, p = 0.8645). These subjects were recruited from blood donors from the blood banks of San Jose and San Juan de Dios Hospitals and from un-clefted patients from the Dental Clinic, School of Dentistry, Universidad de Chile (Santiago, Chile). All control subjects had a negative family history of OFCs. Clinical and demographic data for cases and controls are detailed in Supplementary Table S1 (online). Our study was conducted following the guidelines of the Declaration of Helsinki and was approved by The Institutional Review Board of the Faculty of Dentistry, Universidad de Chile (Protocol #2017/07). All participants or their legal guardians provided their written informed consent. For this study, an alliance with the University of Chile BioBank (BTUCH) was established for all aspects related to the collection, processing, and secure storage of both clinical and epidemiological data and biological samples. BTUCH meets international standards for storage, tracking, processing, and distribution of samples and data policies.34,35 This platform applied a broad informed consent authorized by the Institutional Review Board of the Clinical Hospital, University of Chile.
SNP selection and genotype extraction
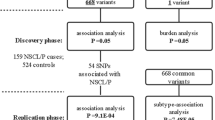
Genomic DNA samples from each participant were purified from venous peripheral blood or from buccal swabs using a standard method (QIAamp DNA Blood Mini Kit; Qiagen, Venlo, Netherlands). Genotypes for AHCY, MTR, MTRR, and MAT2A SNPs were obtained by performing an Infinium Global Screening Array-24 BeadChip (Illumina, San Diego, CA, USA) at the Human Genomics Facility (HuGe-F) in Erasmus MC, Netherlands according to the manufacturer’s instructions. Subjects and markers included here passed the quality control tests according to previously stated criteria.36 For cases and controls, a genotype call rate ≥95% was used. Then, SNPs were selected among those meeting all the following criteria: (a) located between 5 kb upstream from the transcription start site and 5 kb downstream from the stop triplet (according to human genome GRCh37 assembly); (b) a minor allele frequency (MAF) >0.10; (c) not in linkage disequilibrium (LD) with other SNPs assayed in the array (r2 > 0.8); and (d) no significant departure from the Hardy–Weinberg equilibrium (HWE) in the control population (p < 0.01). Supplementary Table S2 (online) describes the list of SNPs included in this study.
Statistical analyses
Genotype set manipulation and additive (allele), recessive, and dominant model associations were performed using PLINK 1.9 (http://www.cog-genomics.org/plink/1.9/). In order to detect HWE departures we applied an exact test. Association between SNPs and phenotype was assessed under the three above-mentioned models by performing a logistic regression analysis. Because the association of genetic markers and NSCL/P differs by sex,33,37 p values for additive model association were also adjusted for sex. To infer the effect of population stratification on association, we obtained a set of genotypes from Infinium Global Screening Array (Illumina, CA, USA) where SNPs from extended regions of high LD (r2 > 0.2) were excluded. The genotypes for a set of 284,256 autosome SNPs were employed for a principal component analysis (PCA). PC1 and PC2 were included as covariates in the logistic regression analyses in order to adjust p values for population stratification.38 False discovery rate (FDR) was applied for multiple comparisons correction according to the Benjamini–Hochberg method.39 For this purpose, a threshold for FDR-adjusted p values (q value) of 0.05 was considered. For haplotype-based association analysis, UNPHASED software was used, which performs a likelihood-ratio test for each haplotype.40 The statistical power (1 − β) for genetic association based on the additive allele model was computed using Quanto 1.2.4 (http://biostats.usc.edu/Quanto.html).
Functional prediction and annotation for significant SNPs
Association with gene expression was evaluated using the Genotype-Tissue Expression (GTEx) database (https://www.gtexportal.org/home/). For variants located within intron and/or exon–intron boundaries, its potential effects on splicing was evaluated using the Human Splice Finder online tool (http://www.umd.be/HSF/).41 This tool detects significant changes produced by sequence variants in sites, such as donor, acceptor, branch point, exonic enhancer, and silencer sites. In addition, associated intronic variants were assessed using the regSNP-intron (https://regsnps-intron.ccbb.iupui.edu/), which predicts the likelihood of an intronic SNP to cause a disease.42
Results
After filters were applied as described in the “Methods” section, 18 SNPs of AHCY, MTR, MTRR, and MAT2A were included in this association study. Their rsIDs, chromosome position, alleles, and MAFs in NSCL/P cases and controls are detailed in Supplementary Table S2 (online). After a multiple comparison correction, the additive (allele) model showed that three variants within MTR are associated with NSCL/P in our sample: rs10925239 (odds ratio (OR) 0.68; 95% confidence interval (CI) 0.53–0.89; p = 0.0032; q = 0.0192), rs10925254 (OR 0.66; 95% CI 0.50–0.86; p = 0.0018; q = 0.0162), and rs3768142 (OR 0.66; 95% CI 0.50–0.86; p = 0.0015; q = 0.0162) (Table 1). None of the other markers for MTR or the other three genes exhibited significant results. The association between these three MTR SNPs and NSCL/P remained significant when its p values were adjusted for sex: rs10925239 (p = 0.0023; q = 0.0138), rs10925254 (p = 0.0017; q = 0.0138) and rs3768142 (p = 0.0014; q = 0.0138) (Table 1). Regarding population stratification, after adjusting for PC1 and PC2, the significance of the association between these three MTR SNPs and NSCL/P was comparable with the unadjusted data: rs10925239 (p = 0.0016; q = 0.0096), rs10925254 (p = 0.0013; q = 0.0096) and rs3768142 (p = 0.0011; q = 0.0096) (Table 1). When the dominant model of association was evaluated, none of the 19 SNPs from the four genes considered showed significant results (Table 2). On the other hand, when a recessive model was assessed, association with NSCL/P was detected for the same three aforementioned SNPs of MTR: rs10925239 (OR 0.37; 95% CI 0.20–0.68; p = 0.0014; q = 0.0108), rs10925254 (OR 0.33; 95% CI 0.17–0.65; p = 0.0018; q = 0.0108) and rs3768142 (OR 0.33; 95% CI 0.17–0.65; p = 0.0018; q = 0.0108) (Table 2). Considering the additive model OR, MAF, and our sample size, we estimated the statistical power for the association of the three MTR SNPs to be 85% for rs10925239, 88% for rs10925254, and 88% for rs3768142. Haplotype-based association showed that the haplotype T–T–G, composed, respectively, by the minor alleles of rs10925239, rs10925254, and rs3768142, is more frequent in controls than in cases (OR 0.73; 95% CI 0.57–0.95; p = 0.0171) (Table 3). Haplotypes including all MTR SNPs did not show significant association with the trait, as well as the combination for MTRR and AHCY gene variants (Supplementary Table S3, online).
From the GTEx database, we found that the three MTR SNPs are associated with the expression levels of MTR in several human tissues and may be considered expression quantitative trait loci. The tissues where these variants significantly alter gene expression are detailed in the Supplementary Table S4 (online). In five human tissues, the major allele of these three SNPs is associated with increased expression of MTR; therefore, we can infer that the associated alleles correlate with a decrease in expression. According to Ensembl (GRCh38.p13; https://www.ensembl.org/index.html) the three associated SNPs are located within introns of the human MTR gene. Thus, rs10925239 is located 2817 bp upstream of exon 9, rs10925254 is 200 bp downstream from exon 18, and rs3768142 is 1710 bp downstream from exon 22. Based on the results of regSNP-intron, we found that none of the MTR-associated SNPs are probable causes of disease (Supplementary Table S4, online). Using the tool LDlink (https://ldlink.nci.nih.gov/), we also found that none of these SNPs are in LD (r2 > 0.8) with other potential functional variants found in American admixed population data (data not shown). Thus, applying the algorithms from the Human Splice Finder tool, we found that all associated SNPs may have an effect on MTR splicing. Considering its position relative to its closest exon, one can highlight that the minor allele of rs10925239 (T) breaks an exon splice silencer site, in comparison to the major allele (G). In the case of the T allele of rs10925254, this variant creates a new exon splice enhancer site. Finally, for rs3768142, we showed that its G allele, in comparison to a T allele, generates a new donor splice site and two new exon splice enhancer sites (Supplementary Table S4, online).
Discussion
Population-based association between a biallelic marker and a disease is based on the premise that the model of inheritance for the risk/protective allele is unknown.43 Using the additive (allele) model, we found that the minor allele of three intronic MTR SNPs (rs10925239, rs10925254, and rs3768142) is significantly less frequent in cases than in controls, demonstrating a protective effect against the expression of NSCL/P in a Chilean population (Table 1). Association for this model may be interpreted as the carriers of two copies of the minor allele have double the protective effect as compared to heterozygotes.43 In addition, this additive association is independent of sex for the three MTR intronic SNPs, evidenced after adjusting p values for this association by this co-variable (Table 1). Differences related with sex have been described in prevalence of OFCs, where NSCL/P male cases are around double those of female cases. In a Chilean population with a lower sample size than the current one, we have detected differences among sexes for the association of MTHFR and SHMT1 variants and NSCL/P.42,44 The recessive model of association was consistent with the additive model for the SNPs rs10925239, rs10925254, and rs3768142 (Table 2). Thus, there is a protective effect for carriers of two copies of the minor allele of these variants in comparison to carriers of one or zero copies.43 We also assessed if the protective association with the phenotype was observed when these markers form a haplotype. Our findings showed that, although it borders the significant (Table 3), to carry one copy of the minor allele for the three SNPs on the same chromosome decreases the risk of NSCL/P. Haplotype-based methods may capture the interactive effect between two or more risk/protective variants on the same chromosome. Therefore, this can be considered a more powerful approach than single-marker assessments of susceptible gene mapping for complex traits.45
The MTR gene is expressed during craniofacial development in mice, specifically in the first branchial arch, mandibular, and maxillary prominences, and secondary palate shelves (https://bgee.org). Its product catalyzes the re-methylation of Hcy into methionine, which in turn is necessary for subsequent SAM synthesis and protein synthesis.21 Thus, one can infer that MTR expression is necessary for substrate methylation during craniofacial morphogenesis. The minor alleles for the three associated SNPs are associated with a decrease in the expression of MTR in human tissues (Supplementary Table S4, online). If one considers the results of the recessive model and the haplotype association, the carriers of two copies of the minor allele rs10925239, rs10925254, and/or rs3768142 have a reduced risk of NSCL/P associated with a decreased expression of MTR. This may result in the accumulation of Hcy, and a reduction of both methionine and SAM. This hypothesis could sound paradoxical seeing as the nutrients that stimulate SAM synthesis, such as folates and vitamin B12 in the maternal periconceptional diet are protective against OFCs.32,44 Nevertheless, in support of our hypothesis, there is evidence of association of fetal and maternal genotypes of the MTR SNP rs1805087 (A > G) with NSCL/P in different populations.24,25,46,47,48 This SNP generates a missense variant that seems to increase enzyme activity based on findings that the risk allele (G) carriers have lower Hcy and higher folate circulating levels than common allele carriers.49 In addition, GG subjects have an increase in global DNA methylation in comparison to the AA genotype.50 Another piece of evidence supporting our hypothesis is the fact that when SAM concentration is higher than physiological levels, it inhibits MTHFR activity, the enzyme providing the methyl donor folate for methionine re-methylation.51 Thereby, one can infer that the increase of methionine and the subsequent increase of SAM may represent a risk for NSCL/P, whereas a protective effect may be expected with a reduction of MTR expression. We propose the existence of a fine regulation of the physiological levels of these molecules.
The three MTR SNPs associated with NSCL/P in our population may be considered deep intronic variants, defined as pathogenic or disease-associated variants located more than 100 bp away from the exon–intron limits.52 The predictions of regSNP-intron, based on data deposited in 1000 Genomes, HGMD, and ClinVar sources42 showed that none of these variants cause disease. In the past decade, several articles have reported the potential pathogenicity of deep intronic variants in both monogenic and complex traits.52,53,54,55 One of the mechanisms is the activation of intronic non-canonical splice sites, competing with natural sites and leading to the inclusion of pseudo-exons through the creation of new acceptor sites, donor sites or enhancers, or disruption of silencers.52 Pseudo-exon inclusion is related to the appearance of a premature stop codon, where this isoform of messenger RNA is degraded through a mechanism called nonsense-mediated decay.56 In this context, our prediction analysis based on Human Splice Finder tools (Supplementary Table S2) reveals that the protective alleles seem to eliminate an exon splice silencer site (rs10925239) to create a new exon splice enhancer site (rs10925254), and to generate a new donor splice (rs3768142), acting individually or forming a haplotype. Thus, one can infer that these deep intronic-associated variants potentially may decrease the expression of MTR through a mechanism involving splicing alterations and possibly nonsense-mediated decay.
Regarding the strengths of the current report, we can highlight the statistical power of the three associated SNPs, which range from 85% to 88%. Another strength is the fact that the p values for all associated SNPs remain significant after adjustment for PC1 and PC2, reflecting the absence of a population stratification effect on our findings. PCA considers a set of genotypes in order to construct a genetic population structure and is an adequate method for addressing the effect of population stratification on population-based association.38 It has been demonstrated that this design may generate spurious association if populations show stratification by ethnicity, a phenomenon reported in Chile.57,58,59 Nonetheless, the main weakness of our study is the absence of functional evidence for any of the three associated MTR SNPs, which creates an opportunity for future studies either in assessing their effects on in vitro splicing or in vivo in animal models for craniofacial development.
In summary, we conclude that three intronic SNPs of human MTR gene are protective markers against NSCL/P in our sample of the Chilean population, either as individual variants or conforming a haplotype. The bioinformatic prediction reveals that these deep intronic variants potentially alter the splicing process, which may explain their association with the decreased MTR expression, as shown by annotation evidence for these SNPs. We hypothesize that a decrease in MTR enzyme levels modulates methionine and SAM availability for proper substrate methylation. Our findings can be used to support further in vitro or in vivo analysis in order to confirm our hypothesis.
References
Xavier, M. J., Roman, S. D., Aitken, R. J. & Nixon, B. Transgenerational inheritance: how impacts to the epigenetic and genetic information of parents affect offspring health. Hum. Reprod. Update 25, 518–540 (2019).
Marcho, C., Oluwayiose, O. A. & Pilsner, J. R. The preconception environment and sperm epigenetics. Andrology (2020) (ahead of print).
Slatkin, M. Epigenetic inheritance and the missing heritability problem. Genetics 182, 845–850 (2009).
Trerotola, M., Relli, V., Simeone, P. & Alberti, S. Epigenetic inheritance and the missing heritability. Hum. Genomics 9, 17 (2015).
Nazer, H. J. & Cifuentes, O. L. Prevalence of congenital malformations at birth in Chilean maternity hospitals. Rev. Med. Chil. 142, 1150–1156 (2014).
Saleem, K., Zaib, T., Sun, W. & Fu, S. Assessment of candidate genes and genetic heterogeneity in human non syndromic orofacial clefts specifically non syndromic cleft lip with or without palate. Heliyon 5, e03019 (2019).
Vieira, A. R. Unraveling human cleft lip and palate research. J. Dent. Res. 87, 119–125 (2008).
Wehby, G. L., Pedersen, D. A., Murray, J. C. & Christensen, K. The effects of oral clefts on hospital use throughout the lifespan. BMC Health Serv. Res. 12, 58 (2012). (2012).
Beaty, T. H., Marazita, M. L. & Leslie, E. J. Genetic factors influencing risk to orofacial clefts: today’s challenges and tomorrow’s opportunities. F1000Res. 5, 2800 (2016).
Leslie, E. J. et al. Association studies of low-frequency coding variants in nonsyndromic cleft lip with or without cleft palate. Am. J. Med. Genet. A 173, 1531–1538 (2017).
Sharp, G. C. et al. Distinct DNA methylation profiles in subtypes of orofacial cleft. Clin. Epigenet. 9, 63 (2017).
Khan, M. et al. Evaluating LINE-1 methylation in cleft lip tissues and its association with early pregnancy exposures. Epigenomics 10, 105–113 (2018).
Khan, M. et al. LINE-1 methylation in cleft lip tissues: Influence of infant MTHFR c.677C>T genotype. Oral Dis. 25, 1668–1671 (2019).
Xu, Z., Lie, R. T., Wilcox, A. J., Saugstad, O. D. & Taylor, J. A. A comparison of DNA methylation in newborn blood samples from infants with and without orofacial clefts. Clin. Epigenet. 11, 40 (2019).
Fortschegger, K. et al. PHF8 targets histone methylation and RNA polymerase II to activate transcription. Mol. Cell Biol. 30, 3286–3298 (2010).
Liu, S., Higashihori, N., Yahiro, K. & Moriyama, K. Retinoic acid inhibits histone methyltransferase Whsc1 during palatogenesis. Biochem. Biophys. Res. Commun. 458, 525–530 (2015).
Gou, Y. et al. Protein arginine methyltransferase PRMT1 is essential for palatogenesis. J. Dent. Res. 97, 1510–1518 (2018).
Serefidou, M., Venkatasubramani, A. V. & Imhof, A. The impact of one carbon metabolism on histone methylation. Front. Genet. 10, 764 (2019).
Mato, J. M., Alvarez, L., Ortiz, P. & Pajares, M. A. S-adenosylmethionine synthesis: molecular mechanisms and clinical implications. Pharm. Ther. 73, 265–280 (1997).
Baric, I. et al. S-adenosylhomocysteine hydrolase deficiency in a human: a genetic disorder of methionine metabolism. Proc. Natl Acad. Sci. USA 101, 4234–4239 (2004).
Li, Y. N. et al. Cloning, mapping and RNA analysis of the human methionine synthase gene. Hum. Mol. Genet. 5, 1851–1858 (1996).
Leclerc, D. et al. Cloning and mapping of a cDNA for methionine synthase reductase, a flavoprotein defective in patients with homocystinuria. Proc. Natl Acad. Sci. USA 95, 3059–3064 (1998).
Mao, Z., Liu, S., Cai, J., Huang, Z. Z. & Lu, S. C. Cloning and functional characterization of the 5′-flanking region of human methionine adenosyltransferase 2A gene. Biochem. Biophys. Res. Commun. 248, 479–484 (1998).
Mostowska, A., Hozyasz, K. K. & Jagodzinski, P. P. Maternal MTR genotype contributes to the risk of non-syndromic cleft lip and palate in the Polish population. Clin. Genet. 69, 512–517 (2006).
Blanton, S. H. et al. Folate pathway and nonsyndromic cleft lip and palate. Birth Defects Res. A 91, 50–60 (2011).
Bhaskar, L. V., Murthy, J. & Venkatesh Babu, G. Polymorphisms in genes involved in folate metabolism and orofacial clefts. Arch. Oral Biol. 56, 723–737 (2011).
Murthy, J., Gurramkondab, V. B. & Lakkakula, B. V. Genetic variant in MTRR A66G, but not MTR A2756G, is associated with risk of non-syndromic cleft lip and palate in Indian population. J. Oral Maxillofac. Surg. 27, 782–785 (2015).
Marini, N. J. et al. Sequence variation in folate pathway genes and risks of human cleft lip with or without cleft palate. Am. J. Med. Genet. A 170, 2777–2787 (2016).
Funato, N. & Nakamura, M. Identification of shared and unique gene families associated with oral clefts. Int. J. Oral Sci. 9, 104–109 (2017).
Wang, P. et al. Evidence of interaction between genes in the folate/homocysteine metabolic pathway in controlling risk of non-syndromic oral cleft. Oral Dis. 24, 820–828 (2018).
Marini, N. J., Asrani, K., Yang, W., Rine, J. & Shaw, G. M. Accumulation of rare coding variants in genes implicated in risk of human cleft lip with or without cleft palate. Am. J. Med. Genet. A 179, 1260–1269 (2019).
Jahanbin, A., Shadkam, E., Miri, H. H., Shirazi, A. S. & Abtahi, M. Maternal folic acid supplementation and the risk of oral clefts in offspring. J. Craniofac. Surg. 29, e534–e541 (2018).
Ramírez-Chau, C., Blanco, R., Colombo, A., Pardo, R. & Suazo, J. MTHFR c.677C>T is a risk factor for non-syndromic cleft lip with or without cleft palate in Chile. Oral Dis. 22, 703–708 (2016).
IARC. Common Minimum Technical Standards and Protocols for Biobanks Dedicated to Cancer Research. IARC Technical Publication No. 44 (IARC, 2017).
OECD. OECD Guidelines on Human Biobanks and Genetic Research Databases (Organization for Economic Cooperation and Development (OECD), Paris, 2009).
Anderson, C. A. et al. Data quality control in genetic case–control association studies. Nat. Protoc. 5, 1564–1573 (2010).
Suazo, J. et al. Risk variants in BMP4 promoters for nonsyndromic cleft lip/palate in a Chilean population. BMC Med. Genet. 12, 163 (2011).
Price, A. L., Zaitlen, N. A., Reich, D. & Patterson, N. New approaches to population stratification in genome-wide association studies. Nat. Rev. Genet. 11, 459–463 (2010).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Statistic. Soc. Ser. B 57, 289–300 (1995).
Dudbridge, F. Likelihood-based association analysis for nuclear families and unrelated subjects with missing genotype data. Hum. Hered. 66, 87–98 (2008).
Desmet, F. O. et al. Human Splicing Finder: an online bioinformatics tool to predict splicing signals. Nucleic Acids Res. 37, e67 (2009).
Lin, H. et al. RegSNPs-intron: a computational framework for predicting pathogenic impact of intronic single nucleotide variants. Genome Biol. 20, 254 (2019).
Salanti, G. et al. Underlying genetic models of inheritance in established type 2 diabetes associations. Am. J. Epidemiol. 170, 537–545 (2009).
Wallenstein, M. B., Shaw, G. M., Yang, W. & Carmichael, S. L. Periconceptional nutrient intakes and risks of orofacial clefts in California. Pediatr. Res. 74, 457–465 (2013).
Salamanca, C. et al. A SHMT1 variant decreases the risk of nonsyndromic cleft lip with or without cleft palate in Chile. Oral Dis. 26, 159–165 (2020).
Liu, N., Zhang, K. & Zhao, H. Haplotype-association analysis. Adv. Genet. 60, 335–405 (2008).
Chorna, L. B., Akopian, H. R., Makukh, H. V. & Fedoryk, I. M. Allelic polymorphism of MTHFR, MTR and MTRR genes in patients with cleft lip and/or palate and their mothers. Tsitol. Genet. 45, 51–56 (2011).
Jin, L. L., Chen, E. J., Hou, W., Liu, X. H. & Hu, Y. The Association between folate pathway genes and cleft lip with or without cleft palate in a Chinese population. Biomed. Environ. Sci. 28, 136–139 (2015).
Wang, W. et al. MTRR, and MTHFR gene polymorphisms and susceptibility to nonsyndromic cleft lip with or without cleft palate. Genet. Test. Mol. Biomark. 20, 297–303 (2016).
Goode, E. L., Potter, J. D., Bigler, J. & Ulrich, C. M. Methionine synthase D919G polymorphism, folate metabolism, and colorectal adenoma risk. Cancer Epidemiol. Biomark. Prev. 13, 157–162 (2004).
Weiner, A. S., Boyarskikh, U. A., Voronina, E. N., Mishukova, O. V. & Filipenko, M. L. Methylenetetrahydrofolate reductase C677T and methionine synthase A2756G polymorphisms influence on leukocyte genomic DNA methylation level. Gene 533, 168–172 (2014).
Finkelstein, J. D. The metabolism of homocysteine: pathways and regulation. Eur. J. Pediatr. 157(Suppl. 2), S40–S44 (1998).
Vaz-Drago, R., Custódio, N. & Carmo-Fonseca, M. Deep intronic mutations and human disease. Hum. Genet. 136, 1093–1111 (2017).
Venables, J. P. Downstream intronic splicing enhancers. FEBS Lett. 581, 4127–4131 (2007).
Cooper, D. N. Functional intronic polymorphisms: buried treasure awaiting discovery within our genes. Hum. Genomics 4, 284–288 (2010).
Abramowicz, A. & Gos, M. Splicing mutations in human genetic disorders: examples, detection, and confirmation. J. Appl. Genet. 59, 253–268 (2018).
Grellscheid, S. N. & Smith, C. W. An apparent pseudo-exon acts both as an alternative exon that leads to nonsense-mediated decay and as a zero-length exon. Mol. Cell. Biol. 26, 2237–2246 (2006).
Santos, J. L. et al. Applicability of the case-parent design in the etiological research of Type 1 diabetes in Chile and other genetically mixed populations. Diabetes Res. Clin. Pract. 43, 143–146 (1999).
Santos, J. L., Pérez, F., Carrasco, E. & Albala, C. Use of case–parents trio for epidemiological studies of association between genetic polymorphisms and complex diseases. Rev. Med. Chil. 130, 1307–1315 (2002).
Acknowledgements
We thank the patients, their families, and healthy individuals who voluntarily cooperated with us, and the staff members of all cleft children rehabilitation centers and blood banks. We also thank the University of Chile BioBank (BTUCH) staff members for their fundamental work. This study was supported by the Fondo Nacional de Desarrollo Científico y Tecnológico (FONDECYT, Chile) Grant #1170805.
Author information
Authors and Affiliations
Contributions
C.S. made substantial contributions to conception and design, analysis and interpretation of data, drafting, revising, and approved the final version of the manuscript. P.G-H. made substantial contributions to conception, analysis and interpretation of data, drafting, revising, and approved the final version of the manuscript. A.S.R. made substantial contributions to acquisition of data, drafting, revising, and approved the final version of the manuscript. P.A.R. made substantial contributions to acquisition of data, drafting, revising, and approved the final version of the manuscript. R.P. made substantial contributions to the acquisition of data, drafting, revising, and approved the final version of the manuscript. N.L. made substantial contributions to acquisition of data, drafting, revising, and approved the final version of the manuscript. R.P. made substantial contributions to acquisition of data, drafting, revising, and approved the final version of the manuscript. J.S. made substantial contributions to conception and design, acquisition, analysis and interpretation of data, drafting, revising, and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Informed consent
Informed consent for all participants was required, which was approved by the Institutional Review Board of the Faculty of Dentistry, Universidad de Chile (Protocol #2017/07).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Salamanca, C., González-Hormazábal, P., Recabarren, A.S. et al. Genetic variants in S-adenosyl-methionine synthesis pathway and nonsyndromic cleft lip with or without cleft palate in Chile. Pediatr Res 89, 1020–1025 (2021). https://doi.org/10.1038/s41390-020-0994-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41390-020-0994-3
This article is cited by
-
Vitamin B12 Transport Genes and Nonsyndromic Cleft Lip With or Without Cleft Palate in Chile
Reproductive Sciences (2022)