Abstract
C-terminal mutation of Nucleophosmin 1 (NPM1C+) was thought to be a primary driving event in acute myeloid leukemia (AML) that reprograms leukemic-associated transcription programs to transform hematopoietic stem and progenitor cells (HSPCs). However, molecular mechanisms underlying NPM1C+-driven leukemogenesis remain elusive. Here, we report that NPM1C+ activates signature HOX genes and reprograms cell cycle regulators by altering CTCF-driven topologically associated domains (TADs). Hematopoietic-specific NPM1C+ knock-in alters TAD topology leading to disrupted regulation of the cell cycle as well as aberrant chromatin accessibility and homeotic gene expression, which results in myeloid differentiation block. Restoration of NPM1 within the nucleus re-establishes differentiation programs by reorganizing TADs critical for myeloid TFs and cell cycle regulators that switch the oncogenic MIZ1/MYC regulatory axis in favor of interacting with coactivator NPM1/p300, and prevents NPM1C+-driven leukemogenesis. In sum, our data reveal that NPM1C+ reshapes CTCF-defined TAD topology to reprogram signature leukemic transcription programs required for cell cycle progression and leukemic transformation.
Similar content being viewed by others
To the Editor
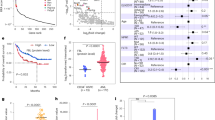
Mutation of NPM1 results in cytoplasmic sequestration of NPM1C+ protein that maintains a unique expression pattern of HOXA/B genes and MEIS1 oncogene by directly binding to chromatin target sites to facilitate AML leukemogenesis [1,2,3]. However, molecular mechanisms underlying NPM1C+-driven gene regulatory networks and leukemogenesis remain elusive. Since wild-type (WT) NPM1/NPM1C+-associates with CTCF and may influence CTCF-mediated loops by anchoring for CTCF cluster [4, 5], it has been long speculated that NPM1C+ mislocalization may alter CTCF-mediated gene expression. We test whether mislocalization of NPM1C+ to cytoplasm alters CTCF-mediated three-dimensional (3D) topologically associated domains (TADs) by Hi-C comparing mouse bone marrow (BM) Lin-cKit+ (LK) stem- and progenitor-enriched cells purified from WT and NPM1C+-KI animals induced for NPM1C+ expression for 4-months (Table S1 listed kits, constructs, cells, antibodies). Mice conditionally expressing NPM1C+ in the BM develop a myeloproliferative neoplasm/myelodysplasia-like condition with a long latency characterized by enlarged spleens, leukocytosis, and thrombocytopenia [6, 7]. We confirmed that NPM1C+ cKI aberrantly activated Hoxa/Hoxb and Meis1 genes that coincided with a gain of chromatin accessibility as seen by RNA-seq and ATAC-seq (Fig. S1A, B) [1, 8]. NPM1C+ expression led to an increase of 46 TADs and a decrease of 31 TADs (Fig. 1A). The increased TADs encompassed 434 genes of which approximately 18% (78 genes) overlapped with genes increased in expression upon NPM1C+-cKI measured by RNA-seq (Fig. 1B, top). Theses overlapping genes are mainly NPM1C+-signature genes involved in pathways regulating transcription, anterior/posterior patterning, homeobox genes, and cell cycle (Fig. 1C, top). Top TADs strengthened by the NPM1C+-cKI included Hoxa/b, and Pbx3 loci (Fig. 1D–F). Many of these genes are specifically involved in unique NPM1C+-driven leukemic transcription program [1, 8]. In contrast, the decreased TADs included cell cycle kinase inhibitors (CDKIs), such as Cdkn1a (p21) gene (Fig. 1G). The altered TADs or sub-TADs were defined by CTCF binding when we integrated with public available mouse CTCF ChIP-seq dataset (GSM918744), whereby CTCF bound to the altered TAD or sub-TAD boundaries (Fig. 1D-G, Red arrowheads). As expected, chromatin accessibility in Hoxa/b and Pbx3 loci were coordinately increased (Figs. S1B, 1H). In light of NPM1C+ directly interacting with CTCF and binding to its chromatin targets [2, 3, 5], NPM1C+ and CTCF may coordinate to remodels NPM1C+-signature gene TADs resulting in altered chromatin accessibility and oncogenic gene expression signatures.
A Overlapping of mouse BM HS/PC TADs and NPM1C+ KI HS/PC TADs identified by Hi-C assay using WT and NPM1C+ cKI mouse BM LK cells of which NPM1C+ was induced at 8 weeks old animals for 4 months. B Overlapping of enhanced or reduced TAD encompassed gene and upregulated or downregulated genes by RNA-seq upon NPM1C+ KI in BM HS/PCs, respectively. C GO term analysis of overlapped enhanced or reduced TAD encompassed gene and upregulated or downregulated genes by RNA-seq upon NPM1C+ KI in BM HS/PCs, respectively. NPM1C+ was induced at 8 weeks old animals for 4 months. D Hi-C interacting maps in part of mouse chromosome 6 regions containing Hoxa locus comparing WT and NPM1C+ KI mouse BM HS/PCs. Publicly available CTCF ChIP-seq data (GSM918744) in MEL cells was integrated on top with Hi-C data. Note: CTCF binding at the boundaries of altered TADs or sub-TADs (red arrow heads). E Hi-C interacting maps in part of mouse chromosome 11 regions containing Hoxb locus comparing WT and NPM1C+ KI mouse BM HS/PCs. Publicly available CTCF ChIP-seq data (GSM918744) in MEL cells was integrated on top with Hi-C data. Note: CTCF binding at the boundaries of altered TADs or sub-TADs (red arrow heads). F Hi-C interaction map at the Pbx3 locus comparing WT and NPM1C+ KI in BM HS/PCs. Publicly available CTCF ChIP-seq data (GSM918744) in MEL cells was integrated on top with Hi-C data. Note: CTCF binding at the boundaries of altered TADs or sub-TADs (red arrow heads). G Hi-C interaction map at the Cdkn1a locus comparing WT and NPM1C+ KI in BM HS/PCs. Publicly available CTCF ChIP-seq data (GSM918744) in MEL cells was integrated on top with Hi-C data. Note: CTCF binding at the boundaries of altered TADs or sub-TADs (red arrow heads). H ATAC-seq chromatin accessibility of Pbx3 gene locus compared WT and NPM1C+ KI in BM HS/PCs.
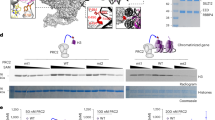
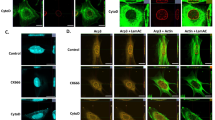
We next transfected OCI-AML3 cells with vector caring WT NPM1 cDNA to restore nuclear NPM1 (NPM1OE), or treated OCI-AML3 cells with 50 nM XPO1 inhibitor (XPO1i), selinexor, for 3 days to block the NPM1C+ cytoplasmic export, and test if restoration of nuclear NPM1 reversed NPM1C+-associated HOX expression signature and TAD formation. Immunofluorescent staining confirmed that XPO1i or NPM1OE resulted in nuclear localization of NPM1 and enhanced its colocalization with CTCF (Fig. 2A). Nuclear retention of NPM1 reduced MYC, MEIS1, and HOX gene expression, while increased CDKIs, CDKN1A and CDKN2A genes (Fig. 2B), consistent with the transcriptomic GO analysis (Fig. S2A, B).
A Immunofluorescence staining of NPM1 and CTCF localization and association upon XPO1i treatment or WT NPM1 expression. Note: The NPM1 antibody used recognized both WT and NPM1C+ mutant. B Heat map of RNA-seq analysis shows upregulated and downregulated genes upon WT NPM1 overexpression or XPO1i treatment in OCI-AML3 cells. C Overlapping of TADs identified from WT OCI-AML3 cells and NPM1OE OCI-AML3 cells by Hi-C assay. D Overlapping of enhanced or reduced TAD encompassed gene and upregulated or downregulated genes by RNA-seq upon NPM1OE in OCI-AML3 cells. E Hi-C interacting maps in part of human chromosome 6 regions containing CDKN1A gene comparing WT, XPO1i treated, or NPM1OE OCI-AML3 cells. Publicly available CTCF ChIP-seq data in human K562 cells was integrated on top with Hi-C data (GSM1010820). Note: CTCF binding at the boundaries of altered TADs or sub-TADs (red arrow heads). F Hi-C interacting maps in part of human chromosome 9 regions containing CDKN2A/CDNK2B genes comparing WT, XPO1i treated, or NPM1OE OCI-AML3 cells. G Hi-C interaction map at the HOXB loci comparing WT, XPO1i treated, or NPM1OE OCI-AML3 cells. Publicly available CTCF ChIP-seq data in human K562 cells was integrated on top with Hi-C data (GSM1010820). Note: CTCF binding at the boundaries of altered TADs or sub-TADs (red arrow heads). H Survival of NSG mice (n = 5) transplanted with WT or NPM1OE OCI-AML3 cells. Data are presented as mean ± SD. *p ≤ 0.05; **p ≤ 0.01; ***p ≤ 0.001.
Next, we carried out Hi-C and RNA-seq experiments in WT, XPO1i-treated or NPM1OE OCI-AML3 cells, and integrated our Hi-C data with available CTCF ChIP-seq profiles generated in K562 cells (GSM1010820). NPM1OE altered 96 TADs defined by CTCF boundaries. Among those, 55 TADs were decreased, while 41 TADs were increased by NPM1OE (Fig. 2C). Similarly, XPO1i treatment altered 97 CTCF-defined TADs (59 decreased TADs and 38 increased TADs, data not shown). We further integrated the transcriptomic data with Hi-C datasets from XPO1i-treated or NPM1OE OCI-AML3 cells. Upon NPM1OE, 24% of genes residing within up-regulated TADs were transcriptionally up-regulated determined by RNA-seq (Fig. 2D, top), and they were predominantly involved in cell cycle, cell differentiation, apoptosis, DNA damage and repair, and transcription regulation (Fig. S2C, left). Only 2.4% of genes residing within up-regulated TADs were transcriptionally down-regulated (Fig. S2D, Left). On the other hand, 16% of genes encompassed within reduced TADs were significantly down-regulated according to their expression levels (Fig. 2D, bottom), and were enriched in pathways involved in HOX, pathways in cancer, PI3K-AKT signaling, Wnt signaling, positive regulation of ERK1/ERK2, transcriptional regulation, cell cycle, and positive regulation of cell proliferation (Fig. S2C, right). Only 4.5% of genes residing within down-regulated TADs were transcriptionally up-regulated (Fig. S2D, Right). XPO1i treatment reveal very similar TADs and transcriptomic changes (Fig. S2E, F). Thus, NPM1C+-driven TAD alteration may be critical for the NPM1C+ signature gene expression.
In particular, Hi-C data revealed that top ten upregulated TADs encompassed genes encoding CDKIs, p21, p15, p16, and p27, while top ten reduced TADs included HOXB in both XPO1i or NPM1OE (Fig. 2E–G). These altered TADs or sub-TADs are defined by CTCF binding when we integrated with CTCF ChIP-seq data from K562 cells (GSM1010820), whereby CTCF bound to boundaries of the altered TADs or sub-TADs (Fig. 2E–G; Red arrowheads). Interestingly, CDKI genes are regulated by MYC-MIZ-1 transcription axis that is involved in control of LSC proliferation and cell cycle progression [9].
To further support that NPM1 nuclear relocalization is in part inducing genome TAD reorganization, we performed time course experiments by treated OCI-AML3 cells with XPO1i for 0, 1, 3, 6, 24, and 48 h. We observed a significantly increase in nuclear localization of NPM1C+ after 24 h treatment by WB (Fig. S3A) and immunofluorescent-staining (Fig. S3B, C). Time course 3 C assay revealed that CTCF-loops encompassed anterior HOXB genes (HOXB1-HOXB5) were significantly impaired as early as 3 hr upon blocking NPM1C+ cytoplasmic localization, while the CTCF mediated looped interactions encompassing CEBPA/G genes were increased immediately upon blockage of NPM1C+ cytoplasmic translocation (Fig. S3D, Table S2 listed all primers/probes). Coordinately, expression levels of CDKIs and CEBPA genes were only upregulated upon 24 hr and 3 days XPO1i treatment, respectively (Fig. S3E), suggesting that TAD reorganization precede gene expression changes.
It has been proposed that MIZ-1 activates its target genes including CDKN1A and CDKN2B by associating with NPM1 and p300, while in MYC transformed cancer cells, association of MYC and MIZ-1 repels NPM1 and p300 leading to aberrant proliferation [10, 11]. We further tested whether restoration of nuclear NPM1 switches the MIZ-1 transcription circuit from repression to activation of CDKIs by disassociating from MYC in favor of interacting with NPM1. Although MIZ-1 level remains relatively consistent upon XPO1i treatment or NPM1OE, these interventions promoted MIZ-1 interacting with NPM1 in nucleus compared to untreated OCI-AML3 cells (Fig. S4A, bottom). As a consequence, the amount of MIZ-1-associated p300 was increased, while MIZ-1 binding to MYC and G9a, a MYC-associated H3K9-metyltransferase required for repression, was markedly decreased upon nuclear retention of NPM1 (Fig. S4A). RT-qPCR and RNA-seq revealed that expression of several MIZ-1 targets, CDKI and CEBP genes, was significantly increased (Fig. 2B, S3E, S4B). ChIP-qPCR further revealed that MYC and G9a recruitment to these loci was significantly reduced comparing to the constant MIZ-1 chromatin binding (Fig. S4C–E), while recruitment of NPM1 and p300 was significantly increased upon XPO1i treatment or NPM1OE (Fig. S4F, G). As a consequence, active H3K27ac chromatin marks were increased (Fig. S4H) while H3K9me2 repressive chromatin marks were significantly reduced at these MIZ-1 target genes (Fig. S4I) comparing to control IgG (Fig. S4J), However, we cannot distinguish whether WT-NPM1 or NPM1C+ regulates CDKI and CEBP genes upon nuclear retention of NPM1 because the antibody used recognized both forms.
Although NPM1 antibody that we used cannot distinguish WT from mutant protein and their role in CDKI and CEBP gene regulation, analyzing Bru-seq data of WT and XPO1i treated OCI-AML3 cells from recent published data [2] also revealed that the nascent transcripts of CDKN1A and CEBPA genes are consistently enhanced by XPO1i treatment (Fig. S4K), supporting that nuclear restoration of NPM1 reverses aberrant TAD topology and switches the MYC-MIZ-1 axis to restore cell cycle control and myeloid differentiation. However, it remains to be determined how NPM1c+/NPM1 were targeted to distinct chromatin sites to differentially activate oncogenic vs myeloid differentiation pathway. Given that NPM1 interacts with MIZ-1 and chromatin bound CRM1/XPO1 recruits NPM1C+ onto HOX cluster [2, 3, 10, 12]. It is reasonable to suspect that MIZ-1 or CRM1/XPO1 directs NPM1 or NPM1C+ to the intended chromatin targets, respectively. In addition, HOX associated lncRNAs, HOTTIP and HOXBLINC, are aberrantly activated in NPM1C+-AML cells that may also play a role in NPM1C+-driven oncogenic HOX gene expression [8, 13, 14].
Consistent with upregulation of CDKIs along with myeloid TFs, CEBPA and CEBPD, nuclear retention of NPM1 promoted OCI-AML3 cell differentiation seen by increased expression of myeloid-specific surface markers CD11b and CD14 (Fig. S5A–C). Furthermore, nuclear retention of NPM1 consistently increased the proportion of cells in G1 phase at the expense of cell in the S phase and G2/M phase transition (Fig. S5D), which coincided with enhanced apoptosis (Fig. S5E) and decreased cellular proliferation (Fig. S5F). Of note, XPO1i impairs OCI-AML3 cell growth and differentiation in a p53 independent manner [15]. We further test whether restored NPM1 in nuclei of NPM1C+ AML cells affects leukemogenesis and survival of AML mouse models in vivo, we transplanted 1 × 106 WT, NPM1OE OCI-AML3 cells into NSG mice. All animals transplanted with WT OCI-AML3 cells died around 38 days after transplantation, while mice that received OCI-AML3 cells with NPM1OE survived between 54–58 days (Fig. 2H). Thus, nuclear NPM1 localization mitigates leukemogenesis in a xenograft mouse model of AML with NPM1C+, perhaps blocking NPM1C+-driven TAD topology and signature gene expression and restoring cell cycle control and myeloid differentiation.
Data availability
Sequence reads have been deposited in the National Center for Biotechnology Information GENE expression Omnibus (NCBI GEO) under accession number GSE208022.
References
Brunetti L, Gundry MC, Sorcini D, Guzman AG, Huang YH, Ramabadran R, et al. Mutant NPM1 maintains the leukemic state through HOX expression. Cancer Cell. 2018;34:499–512.e499. https://doi.org/10.1016/j.ccell.2018.08.005
Wang XQD, Fan D, Han Q, Liu Y, Miao H, Wang X, et al. Mutant NPM1 hijacks transcriptional hubs to maintain pathogenic gene programs in acute myeloid leukemia. Cancer Discov. 2023;13:724–745. https://doi.org/10.1158/2159-8290.Cd-22-0424
Uckelmann HJ, Haarer EL, Takeda R, Wong EM, Hatton C, Marinaccio C, et al. Mutant NPM1 directly regulates oncogenic transcription in acute myeloid leukemia. Cancer Discov. 2023;13:746–765. https://doi.org/10.1158/2159-8290.Cd-22-0366
Yusufzai TM, Tagami H, Nakatani Y, Felsenfeld G. CTCF tethers an insulator to subnuclear sites, suggesting shared insulator mechanisms across species. Mol Cell. 2004;13:291–298. https://doi.org/10.1016/s1097-2765(04)00029-2
Wang AJ, Han Y, Jia N, Chen P, Minden MD. NPM1c impedes CTCF functions through cytoplasmic mislocalization in acute myeloid leukemia. Leukemia. 2020;34:1278–1290. https://doi.org/10.1038/s41375-019-0681-8
Uckelmann HJ, Kim SM, Wong EM, Hatton C, Giovinazzo H, Gadrey JY, et al. Therapeutic targeting of preleukemia cells in a mouse model of NPM1 mutant acute myeloid leukemia. Science. 2020;367:586–590. https://doi.org/10.1126/science.aax5863
Mallardo M, Caronno A, Pruneri G, Raviele PR, Viale A, Pelicci PG, et al. NPMc+ and FLT3_ITD mutations cooperate in inducing acute leukaemia in a novel mouse model. Leukemia. 2013;27:2248–2251. https://doi.org/10.1038/leu.2013.114
Zhu G, Luo H, Feng Y, Guryanova OA, Xu J, Chen S, et al. HOXBLINC long non-coding RNA activation promotes leukemogenesis in NPM1-mutant acute myeloid leukemia. Nat Commun. 2021;12:1956 https://doi.org/10.1038/s41467-021-22095-2
Zhang L, Li J, Xu H, Shao X, Fu L, Hou Y, et al. Myc-Miz1 signaling promotes self-renewal of leukemia stem cells by repressing Cebpα and Cebpδ. Blood. 2020;135:1133–1145. https://doi.org/10.1182/blood.2019001863
Wiese KE, Walz S, Eyss BV, Wolf E, Athineos D, Sansom O, et al. The role of MIZ-1 in MYC-dependent tumorigenesis. Cold Spring Harb Perspect Med. 2013;3:a014290 https://doi.org/10.1101/cshperspect.a014290
Shostak A, Ruppert B, Ha N, Bruns P, Toprak UH, Eils R, et al. MYC/MIZ1-dependent gene repression inversely coordinates the circadian clock with cell cycle and proliferation. Nat Commun. 2016;7:11807 https://doi.org/10.1038/ncomms11807
Oka M, Mura S, Otani M, Miyamoto Y, Nogami J, Maehara K, et al. Chromatin-bound CRM1 recruits SET-Nup214 and NPM1c onto HOX clusters causing aberrant HOX expression in leukemia cells. eLife. 2019;8:e46667 https://doi.org/10.7554/eLife.46667
Luo H, Zhu G, Xu J, Lai Q, Yan B, Guo Y, et al. HOTTIP lncRNA promotes hematopoietic stem cell self-renewal leading to AML-like disease in mice. Cancer Cell. 2019;36:645–659.e648. https://doi.org/10.1016/j.ccell.2019.10.011
Luo H, Zhu G, Eshelman MA, Fung TK, Lai Q, Wang F, et al. HOTTIP-dependent R-loop formation regulates CTCF boundary activity and TAD integrity in leukemia. Mol Cell. 2022;82:833–851.e811. https://doi.org/10.1016/j.molcel.2022.01.014
Pianigiani G, Gagliardi A, Mezzasoma F, Rocchio F, Tini V, Bigerna B, et al. Prolonged XPO1 inhibition is essential for optimal antileukemic activity in NPM1-mutated AML. Blood Adv. 2022;6:5938–5949. https://doi.org/10.1182/bloodadvances.2022007563
Acknowledgements
We are grateful for the helpful discussions from members of the Huang and Qiu laboratories.
Funding
This work is supported by Four Diamond research fund (SH), National Natural Science Foundation of China (BX, 82170180, 81770126; JZ, 81800163), National Institute of Health (YQ, R01HL144712; OAG, R01DK121831; SH, R01CA260729, R01CA 264932).
Author information
Authors and Affiliations
Contributions
SH, YQ, OAG, BX conceived and designed experiments. QL, KH, JL, ZZ, CZ, JZ performed experiments. KH and HL performed bioinformatics/statistical analysis. SH and YQ wrote/edited the manuscript. SH and BX supervised the studies.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lai, Q., Hamamoto, K., Luo, H. et al. NPM1 mutation reprograms leukemic transcription network via reshaping TAD topology. Leukemia 37, 1732–1736 (2023). https://doi.org/10.1038/s41375-023-01942-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-023-01942-9