Abstract
Cardiac hypertrophy is a common cardiovascular disease that is found worldwide and is characterized by heart enlargement, eventually resulting in heart failure. Exploring the regulatory mechanism of cardiac hypertrophy is beneficial for understanding its pathogenesis and treatment. In our study, we have showed TINCR was downregulated and miR-211–3p was upregulated in TAC- or Ang II-induced models of cardiac hypertrophy. Dual luciferase and RIP assays revealed that TINCR served as a competitive endogenous RNA (ceRNA) for miR-211–3p. Then, we observed that knockdown of miR-211–3p alleviated TAC- or Ang II-induced cardiac hypertrophy both in vivo and in vitro. Mechanistically, we demonstrated that miR-211–3p directly targeted VEGFB and thus regulated the expression of SDF-1α and CXCR4. Rescue assays further confirmed that TINCR suppressed the progression of cardiac hypertrophy by competitively binding to miR-211–3p, thereby enhancing the expression of VEGFB and activating the VEGFB-SDF-1α- CXCR4 signal. Furthermore, overexpression of TINCR suppressed TAC-induced cardiac hypertrophy in vivo by targeting miR-211–3p-VEGFB-SDF-1α- CXCR4 signalling. In conclusion, our research suggests that LncRNA TINCR improves cardiac hypertrophy by targeting miR-211–3p, thus relieving its suppressive effects on the VEGFB-SDF-1α-CXCR4 signalling axis. TINCR and miR-211–3p might act as therapeutic targets for the treatment of cardiac hypertrophy.
Similar content being viewed by others
Introduction
Cardiac hypertrophy arises from increased cardiomyocyte size and generally causes thickened cardiac muscle and heart enlargement, which has been well recognized as a crucial mechanism to adapt to stressful conditions, such as hypertension and valvular disorder1,2. Cardiac hypertrophy consists of two types: physiological and pathological hypertrophy. Physiological hypertrophy is generally a reversible and mild response to physiological stimuli, such as endurance exercise and pregnancy, and maintains heart functions. However, pathological cardiac hypertrophy is accompanied by myofibroblast activation, myocardial fibrosis, and cell death, and eventually causes systolic dysfunction and heart failure3. With the development of diagnostic methods, drugs including calcium channel blockers and thiazide diuretics, and surgical procedures, great progress has been made in the diagnosis and treatment of cardiac hypertrophy in recent years4. Even so, cardiac hypertrophy is still a serious cardiovascular issue found worldwide5. It is very important to explore the molecular mechanisms by which cardiac hypertrophy is regulated to identify novel diagnostic biomarkers and therapeutic targets.
Emerging evidence has shown that long noncoding RNAs (lncRNAs) play vital roles in cardiac hypertrophy6,7. Decreased expression of lncRNA TINCR was observed in hypertrophic mice, and silencing of TINCR caused cardiomyocyte enlargement8,9. However, the mechanisms of TINCR in cardiac hypertrophy are still unclear. LncRNAs can act as competitive endogenous RNAs (ceRNAs), relieving their inhibitory effects on downstream targets. MiR-214–5p10, miR-31–5p9, miR-76111, and miR-544a12 have been reported to be competitive combined by lncRNA TINCR in various cells. Importantly, an increasing number of studies have revealed that several miRNAs are dysregulated in cardiomyocytes and facilitate cardiac hypertrophy13,14. MiR-155 was reported to promote cardiac hypertrophy15. Qiu et al. found that miR-20b facilitated cardiac hypertrophy by suppressing Mfn-2-mediated calcium overload16. Overexpression of miR-217 accelerated cardiac hypertrophy by reducing the expression of PTEN17. However, it is not clear whether lncRNA TINCR plays a role in cardiac hypertrophy by competitive combining miRNA.
The vascular endothelial growth factor (VEGF) family consists of placental growth factor, VEGFA, VEGFB, VEGFC, and VEGFD, and plays key roles in regulating angiogenesis, which has been recognized as a key regulator in the cardiovascular system and disorders, including cardiac hypertrophy18,19. As a member of the VEGF family, VEGFB is downregulated in several heart failure models and heart disorders20. In addition, VEGFB improves heart function and protects against heart failure by inducing vascular growth and cardiac hypertrophy21. VEGFB exerts its functions primarily by binding to VEGF receptor 1 (VEGFR1)22. Activation of the VEGFB/VEGFR1 pathway results in increased expression of stromal cell-derived factor-1α (SDF-1α), which contributes to myocardial repair in myocardial infarction23. The chemokine SDF-1α is a ligand for C-X-C motif chemokine receptor 4 (CXCR4), a G protein-coupled receptor24. The SDF-1α-CXCR4 axis is implicated in cell survival and angiogenesis in cardiovascular disorders, and has been regarded as a therapeutic target for cardiovascular diseases25. Intriguingly, Wang et al. demonstrated that blockade of the SDF-1α-CXCR4 signalling axis by deleting CXCR4 exacerbates isoproterenol-induced pathological cardiac hypertrophy in a mouse model. Moreover, overexpression of CXCR4 improves heart function and cardiac hypertrophy in isoproterenol-treated CXCR4 knockout mice26. However, the regulation of the VEGFB/SDF-1α/CXCR4 axis in cardiac hypertrophy is still not fully understood.
In summary, we assume that lncRNA TINCR might regulate cardiac hypertrophy by targeting the miR-211-3p-VEGFB-SDF-1α-CXCR4 pathway. Indeed, we demonstrate for the first time that TINCR and a miR-211-3p inhibitor alleviate cardiac hypertrophy. Mechanistically, TINCR targets miR-211-3p and relieves miR-211-3p-mediated suppression of the VEGFB-SDF-1α-CXCR4 signalling axis. Our study not only elucidates a novel regulatory mechanism of cardiac hypertrophy but also identifies potential diagnostic biomarkers and therapeutic targets for cardiac hypertrophy.
Methods
Cell culture and treatment
The rat cardiomyocyte cell line H9C2 and human embryonic kidney 293T cells were obtained from the American Type Culture Collection (ATCC, Manassas, VA, USA). Cells were grown in Dulbecco’s modified Eagle’s medium with 10% foetal bovine serum at 37 °C in a cell incubator with 5% CO2. Cell culture reagents were purchased from Thermo (Waltham, MA, USA). For the cell model of cardiac hypertrophy, H9C2 cells were treated with Ang II (Sigma, St. Louis, MO, USA) at 1 µM for 24 h and harvested for subsequent assays.
Cell transfection
The miR-211-3p inhibitor, inhibitor NC, shRNAs against VEGFR1 (sh-VEGFR1) and CXCR4 (sh-CXCR4), and shRNA NC were all obtained from RiboBio (Guangzhou, China). For TINCR overexpression, TINCR was inserted into the pcDNA 3.1 vector (OE-TINCR), and empty vector (OE-NC) was used as a control. H9C2 cells were grown to ~80% confluency and transfected with OE-NC, OE-TINCR, inhibitor NC, miR-211-3p inhibitor, shRNA NC, sh-VEGFR1, miR-211-3p inhibitor+sh-VEGFR1, sh-CXCR4, or OE-TINCR + sh-CXCR4 with a Neon Transfection System (Thermo). After 48 h, cells were treated with Ang II or untreated for 24 h and used for subsequent assays.
A transverse aortic constriction (TAC)-induced rat model of cardiac hypertrophy
The TAC-induced rat model of cardiac hypertrophy was established as described previously27. Male 8–10-week Sprague-Dawley rats were divided into two groups: TAC surgery and sham procedure. For TAC surgery, rats were anaesthetized and placed on a heating pad. Rat chests were opened, and a 6G blunt needle was placed parallel to the ascending aorta. Then, a silk suture was tied around the aorta and needle prior to removing the needle. Sham rats received the same procedure except for the tying of knots. Rat chests were closed, and rats were maintained on a heating pad until waking. For gene modification in vivo, adenoviruses for stably knocking down miR-211-3p and overexpressing miR-211-3p or TINCR were prepared, and rats were injected intravenously with adenovirus. Adenoviruses with miR-NC, inhibitor NC, or OE-NC were used as negative controls. Animal experiments were in accordance with National Institutes of Health guidelines and approved by the Animal Care and Use Committee of the Third Xiangya Hospital of Central South University.
Dual-luciferase reporter assay
Potential binding sites for miR-211-3p in TINCR or the 3′UTR of VEGFB were predicted with RNAhybrid (https://bibiserv.cebitec.uni-bielefeld.de/rnahybrid) or TargetScanHuman (http://www.targetscan.org/vert_72/), respectively. Wild-type (VEGFB-WT, TINCR-WT) and mutated (VEGFB-MUT, TINCR-MUT) binding sites for miR-211-3p were cloned into the psiCheck2 vector from Promega (Madison, WI, USA). 293T cells were cotransfected with miR-211-3p mimics and VEGFB-WT, VEGFB-MUT, TINCR-WT, or TINCR-MUT reporters. MiR-211-3p NC (miR-NC) was used as a negative control. Forty-eight hours post transfection, cells were harvested and lysed. The Dual-Glo luciferase system (Promega) was used to examine luciferase activity. Firefly luciferase activity was normalized to Renilla luciferase activity.
RNA immunoprecipitation (RIP)
Cardiac tissues from TAC-induced cardiac hypertrophy rats were lysed in lysis buffer containing protease/ribonuclease inhibitors on ice for 30 min. Supernatants were collected, and 10 µL of each sample was aliquoted as the input. Protein A magnetic beads were precoated with an Ago-2 antibody and added to the supernatants, and samples were incubated at 4 °C overnight. RNA was recovered on the next day. TINCR and miR-211 were amplified from recovered RNA and input by PCR followed by electrophoresis.
Haematoxylin and eosin (H&E) and immunohistochemistry (IHC) staining
Hearts were excised gently from rats and fixed in 4% paraformaldehyde at 4 °C for 16 h. Then, the hearts were dehydrated and embedded in paraffin prior to being cut into 5-µm sections. For H&E staining, after rehydration, sections were stained with H&E and cleared in xylene. Mounted slides were imaged under an Olympus BX51 microscope (Tokyo, Japan). For IHC staining, after rehydration, sections were placed in boiling pH 6.0 sodium citrate buffer for 10 min for antigen retrieval. Sections were then incubated in H2O2 solution at room temperature to inactivate endogenous peroxidase. After washing, the sections were incubated with primary antibodies against ANP (1:200, Abcam, Cambridge, UK) and β-MHC (1:100, Novus) overnight at 4 °C. Sections were rinsed and probed with an HRP-conjugated anti-rabbit secondary antibody (1:1000, Abcam). DAB substrate was used for visualizing the signal. Finally, sections were stained with haematoxylin and imaged under an Olympus BX51 microscope.
Real-time quantitative reverse transcription-PCR (RT-qPCR)
Total RNA was extracted from cardiac tissues from rats and H9C2 cells with the indicated treatments using TRIzol reagent (Thermo) according to the product manual. RNA was quantified with a Qubit 4 fluorometer (Thermo) and then reverse-transcribed into cDNA. The relative expression of TINCR, ANP, BNP, Col1, β-MHC, VEGFB, SDF-1α, and CXCR4 was assessed by quantitative PCR with SYBR green qPCR mix (Toyobo, Osaka, Japan). All gene expression was normalized to GAPDH and calculated using the 2−∆∆Ct method. The primers used for RT-qPCR in this study are listed in Table 1.
Western blot
Homogenized rat cardiac tissues and H9C2 cells with the indicated treatments were lysed in RIPA lysis buffer (Santa Cruz, Dallas, TX, USA) for 1 h (cardiac tissues) or 30 min (H9C2 cells) on ice. After centrifugation, supernatants were harvested and quantified using a BCA kit (Abcam). Protein (30 μg) was loaded in each lane on a 10% SDS-PAGE gel and electrophoresed and then transferred to PVDF membranes (Bio-Rad, Hercules, CA, USA). After blocking in 8% BSA/TBST solution for 1 h, membranes were incubated with primary antibodies against ANP (1:500, Thermo), BNP (1:1000, Abcam), Col1 (1:500, Abcam), β-MHC (1:500, Novus), VEGFB (1:500, Thermo), SDF-1α (1:1000, Thermo), and CXCR4 (1:1000, Novus) for 4 h at room temperature. After rinsing in TBST solution, membranes were incubated with HRP-conjugated secondary antibodies for 1 h at room temperature prior to visualizing the band with ECL substrates (Thermo). The intensity of bands was quantified using ImageJ software.
Statistical analysis
The results from three independent experiments are presented as the mean ± standard deviation (SD). SPSS 17.0 was used in this study for statistical analysis of the data. The variance of two groups was analyzed using Student’s t test. Fisher’s least significant difference was used for multiple comparisons. P < 0.05 was considered as statistically significant.
Results
TINCR targeted miR-211-3p, and silencing miR-211-3p inhibited TAC-induced cardiac hypertrophy in vivo
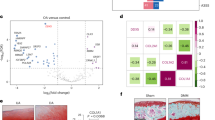
Since lncRNAs exert functions through various mechanisms, including acting as ceRNAs for miRNAs, we assumed that TINCR might target miRNAs to regulate cardiac hypertrophy. A dual-luciferase assay was carried out to validate the interaction between TINCR and miR-211-3p. As shown in Fig. 1A, overexpression of miR-211-3p significantly reduced the luciferase activity of wild-type TINCR but not that of mutated TINCR. Moreover, the RIP assay showed that both TINCR and miR-211-3p were specifically enriched in the anti-Ago-2-immunoprecipitated fractions from cardiac tissues of TAC-induced rats, implying that TINCR might target miR-211-3p in cardiac hypertrophy (Fig. 1B). We also found that TINCR was downregulated and miR-211-3p was upregulated in TAC-treated cardiac tissues (Fig. 1C). To investigate the role of miR-211-3p in cardiac hypertrophy, miR-211-3p was knocked down in rat cardiac tissues (Fig. 1D). We also found that knockdown of miR-211-3p led to a decreased ratio of heart weight to tibial length and reduced cross-sectional area of cardiomyocytes, revealing that knockdown of miR-211-3p had a protective role in TAC-induced cardiomyocyte enlargement (Fig. 1E, F). In addition, we observed that TAC induced the expression of hypertrophic markers, including ANP, BNP, and β-MHC, which were suppressed by the miR-211-3p inhibitor (Fig. 1G-I). Collectively, these results demonstrated that TINCR might target miR-211-3p and that knockdown of miR-211-3p inhibited TAC-induced cardiac hypertrophy in vivo.
A The predicted binding site for miR-211-3p and luciferase activity of TINCR wild-type or mutated reporters (n = 3). B The interaction between miR-211-3p and TINCR. C The relative expression of TINCR and miR-211-3p in rats treated with TAC or sham (n = 3). D The relative expression of miR-211-3p in rats with the indicated treatment (n = 3). E Heart weight to tibial length and cross-sectional areas of cardiomyocytes (n = 3). F Heart photo and H&E staining of heart sections. G The relative mRNA expression of ANP, BNP, β-MHC, and Col1 in rats (n = 3). H Western blot analysis of ANP and β-MHC in rats with the indicated treatment (n = 3). GAPDH was used as a loading and normalization control. I IHC staining of ANP and β-MHC. Data were from at least three independent assays. *P < 0.05, **P < 0.01, and ***P < 0.001.
Knockdown of miR-211-3p suppressed Ang II-induced cardiomyocyte hypertrophy in H9C2 cells
To investigate the function of miR-211-3p in vitro, we established a cell model of cardiac hypertrophy by treating H9C2 cells with Ang II. We found that the cardiomyocyte size of Ang II-treated cells was increased compared to that of control cells (Fig. 2A). In addition, the expression of ANP, BNP, β-MHC, and Col1 was increased in the Ang II treatment group, suggesting that Ang II treatment successfully induced cardiomyocyte hypertrophy (Fig. 2B, C). As expected, the expression of TINCR was decreased and miR-211-3p expression was enhanced in Ang II-treated H9C2 cells (Fig. 2D). Then, miR-211-3p was knocked down through inhibitor transfection (Fig. 2E). Consistent with the in vivo observations, silencing miR-211-3p reduced the size of Ang II-induced H9C2 cells and the expression of ANP, BNP, β-MHC, and Col1 (Fig. 2F-H). These observations showed that knockdown of miR-211-3p improved Ang II-induced cardiomyocyte hypertrophy in vitro.
A Cell size analysis in untreated and Ang II-treated cells (n = 3). B The relative mRNA expression of ANP, BNP, β-MHC, and Col1 (n = 3). C Western blot analysis of ANP, BNP, β-MHC, and Col1 (n = 3). GAPDH was used as a loading and normalization control. D The relative expression of TINCR and miR-211-3p in H9C2 cells treated with Ang II and untreated cells (n = 3). E The relative expression of miR-211-3p in H9C2 cells transfected with miR-211-3p inhibitor or inhibitor NC (n = 3). F Cell size analysis in Ang-induced H9C2 cells transfected with miR-211-3p inhibitor or inhibitor NC. G The relative mRNA expression of ANP, BNP, β-MHC, and Col1 in H9C2 cells with the indicated treatment by RT-qPCR (n = 3). H Western blot analysis of ANP, BNP, β-MHC, and Col1 in H9C2 cells with the indicated treatment (n = 3). GAPDH was used as a loading and normalization control. Data were from at least three independent assays. *P < 0.05, **P < 0.01, and ***P < 0.001.
MiR-211-3p targeted VEGFB and regulated the VEGFB-SDF-1α-CXCR4 axis
To explore downstream targets of miR-211-3p, through bioinformatic analysis, we predicted a potential binding site for miR-211-3p in the 3′UTR of VEGFB, a key regulator in the cardiovascular system (Fig. 3A). The luciferase assay showed that miR-211-3p overexpression significantly suppressed the luciferase activity of wild-type VEGFB but did not affect that of mutated VEGFB, demonstrating that miR-211-3p targeted VEGFB (Fig. 3A). Since the VEGFB-SDF-1α-CXCR4 signalling axis plays important roles in cardiovascular functions and disorders25, we examined whether miR-211-3p could affect the VEGFB-SDF-1α-CXCR4 axis. The results showed that overexpression of miR-211-3p decreased the expression of VEGFB, SDF-1α, and CXCR4 (Fig. 3B, C), but silencing of miR-211-3p increased their expression in H9C2 cells (Fig. 3D, E). In conclusion, miR-211-3p targeted VEGFB and thus suppressed the VEGFB-SDF-1α-CXCR4 signalling axis in H9C2 cells.
A The predicted binding site for miR-211-3p in the 3′UTR of VEGFB and luciferase activity of VEGFB wild-type or mutated reporters (n = 3). B The relative mRNA expression of VEGFB, SDF-1α, and CXCR4 in H9C2 cells transfected with miR-211-3p mimics or mimics NC (n = 3). C Western blot analysis of VEGFB, SDF-1α, and CXCR4 in H9C2 cells (n = 3). GAPDH was used as a loading and normalization control. D The relative mRNA expression of VEGFB, SDF-1α, and CXCR4 in H9C2 cells transfected with miR-211-3p inhibitor or inhibitor NC (n = 3). E Western blot analysis of VEGFB, SDF-1α, and CXCR4 in H9C2 cells (n = 3). GAPDH was used as a loading and normalization control. Data were from at least three independent assays. *P < 0.05, **P < 0.01, and ***P < 0.001.
MiR-211-3p-mediated regulation of cardiac hypertrophy was partially dependent on the VEGFB-VEGFR1-SDF-1α axis
H9C2 cells were treated with Ang II and transfected with miR-211-3p inhibitor, sh-VEGFR1, or both. The results showed that knockdown of miR-211-3p reduced H9C2 cell size, which was partially reversed by silencing VEGFR1 (Fig. 4A). Moreover, the miR-211-3p inhibitor abrogated Ang II-mediated suppression of the expression of VEGFB and SDF-1α (Fig. 4B, C). Silencing VEGFR1 further reduced SDF-1α expression and partially reversed the miR-211-3p inhibitor-induced expression of SDF-1α in Ang II-treated H9C2 cells (Fig. 4B, C). In addition, the miR-211-3p inhibitor suppressed Ang II-induced expression of ANP, BNP, β-MHC, and Col1, which was partially reversed by knockdown of VEGFR1 (Fig. 4D, E). Taken together, these observations suggested that miR-211-3p-mediated regulation of cardiac hypertrophy was partially dependent on the VEGFB-VEGFR1-SDF-1α signalling axis.
H9C2 cells were treated with Ang II, Ang II + shRNA NC, Ang II + inhibitor NC, Ang II + miR-211 inhibitor, Ang II + sh-VEGFR1, or Ang II + miR-211 inhibitor+sh-VEGFR1. Untreated cells were used as the control. A Cell size analysis. B The relative mRNA expression of VEGFB and SDF-1α (n = 3). C Western blot analysis of VEGFB and SDF-1α (n = 3). GAPDH was used as a loading and normalization control. (n = 3). D The relative mRNA expression of ANP, BNP, β-MHC, and Col1 (n = 3). E Western blot analysis of ANP, BNP, β-MHC, and Col1 (n = 3). GAPDH was used as a loading and normalization control. Data were from at least three independent assays. *P < 0.05, **P < 0.01, and ***P < 0.001.
TINCR regulated cardiac hypertrophy by targeting miR-211-3p-VEGFB-SDF-1α signalling and increasing CXCR4 expression
Since CXCR4 is a receptor for SDF-1α and has been regarded as a therapeutic target for several cardiovascular disorders25, we explored whether CXCR4 was a downstream target of the TINCR-miR-211-VEGFB-SDF-1α axis in cardiac hypertrophy. TINCR decreased H9C2 cell size, which was abrogated by knockdown of CXCR4 (Fig. 5A). Moreover, overexpression of TINCR significantly enhanced the expression of VEGFB and SDF-1α in Ang II-treated cells (Fig. 5B, C). However, silencing CXCR4 did not affect the expression of VEGFB and SDF-1α in Ang II-treated cells (Fig. 5B, C). In addition, TINCR overexpression suppressed the expression of ANP, BNP, β-MHC, and Col1 in Ang II-treated H9C2 cells, which was reversed by silencing CXCR4 (Fig. 5D, E). Furthermore, the expression of VEGFB, SDF-1α, and CXCR4 was suppressed in cardiac tissues from TAC-induced cardiac hypertrophy rats and restored by miR-211-3p knockdown (Fig. 5F, G). These data indicated that TINCR regulated cardiac hypertrophy by targeting the miR-211-VEGFB-SDF-1α signalling axis and thus upregulating CXCR4 expression.
H9C2 cells were treated with Ang II, Ang II + vector, Ang II + shRNA NC, Ang II + TINCR, Ang II + sh-CXCR4, or Ang II + TINCR + sh-CXCR4. Untreated cells were used as the control. A Cell size analysis. B The relative mRNA expression of VEGFB and SDF-1α in H9C2 cells (n = 3). C Western blot analysis of VEGFB and SDF-1α (n = 3). GAPDH was used as a loading and normalization control. D The relative mRNA expression of ANP, BNP, β-MHC, and Col1 in H9C2 cells (n = 3). E Western blot analysis of ANP, BNP, β-MHC, and Col1 (n = 3). GAPDH was used as a loading and normalization control. F The relative mRNA expression of VEGFB, SDF-1α, and CXCR4 in cardiac tissues from rats with the indicated treatment (n = 3). G Western blot analysis of VEGFB, SDF-1α, and CXCR4 in cardiac tissues from rats with the indicated treatment (n = 3). GAPDH was used as a loading and normalization control. Data were from at least three independent assays. *P < 0.05, **P < 0.01, and ***P < 0.001.
TINCR inhibited the activation of miR-211-3p-VEGFB-SDF-1α-CXCR4 signalling to suppress TAC-induced cardiac hypertrophy in vivo
Then, we examined whether TINCR suppressed TAC-induced cardiac hypertrophy by regulating miR-211-3p-VEGFB-SDF-1α-CXCR4 signalling in vivo. We found that TAC treatment increased heart weight and cardiomyocyte size, but they were reduced by overexpression of TINCR (Fig. 6A, B). Simultaneous overexpression of miR-211-3p reversed TINCR-mediated suppression of heart weight to tibial length and cross-sectional area of cardiomyocytes (Fig. 6A, B). Furthermore, the expression of VEGFB, SDF-1α, and CXCR4 was reduced by TAC treatment, which was reversed by overexpression of TINCR (Fig. 6C, D). Overexpression of miR-211-3p abrogated TINCR-mediated inhibition of the expression of VEGFB, SDF-1α, and CXCR4 (Fig. 6C, D). In addition, TAC-induced expression of ANP and β-MHC was suppressed by overexpression of TINCR, but these effects were all reversed by simultaneous overexpression of miR-211-3p (Fig. 6E–G). These data demonstrated that TINCR improved TAC-induced cardiac hypertrophy in vivo by targeting the miR-211-3p-VEGFB-SDF-1α-CXCR4 axis.
A Heart photos and heart weight to tibial length from rats with the indicated treatment. B H&E staining of heart sections and cross-sectional areas of cardiomyocytes. C The relative mRNA expression of VEGFB, SDF-1α, and CXCR4 in cardiac tissues (n = 3). D Western blot analysis of VEGFB, SDF-1α, and CXCR4 in cardiac tissues (n = 3). GAPDH was used as a loading and normalization control. E The relative mRNA expression of ANP and β-MHC in cardiac tissues (n = 3). F Western blot analysis of ANP and β-MHC in cardiac tissues (n = 3). GAPDH was used as a loading and normalization control. G IHC staining of ANP and β-MHC. *P < 0.05, **P < 0.01, and ***P < 0.001.
Discussion
Pathological cardiac hypertrophy is a very common cardiovascular disorder and eventually causes heart failure28, which is a serious health issue found worldwide29. Therefore, understanding the regulatory mechanism of cardiac hypertrophy is truly important for its treatment. Recently, growing evidence has demonstrated that lncRNAs exert vital roles in the regulation of cardiac hypertrophy30. In this study, we demonstrate that TINCR alleviates cardiac hypertrophy by acting as a ceRNA for miR-211 to reduce its abundance in cardiac hypertrophy, thereby relieving its suppression of the VEGFB-SDF-1α-CXCR4 signalling axis.
MiRNAs are one of the key regulators in cardiac hypertrophy. It was reported that miR-211-3p was downregulated and inhibited cell proliferation and migration in breast cancers31. Similarly, a miR-211-3p inhibitor can act as an inducer of oral squamous cell carcinoma by targeting MAFG32. Moreover, miR-211-3p has been reported to be involved in regulating endoplasmic reticulum stress and chondrocyte apoptosis33. To the best of our knowledge, our study is the first to reveal that miR-211-3p is dramatically upregulated in cardiac hypertrophy and that knockdown of miR-211-3p alleviates TAC- or Ang II-induced cardiac hypertrophy. This finding revealed a novel function of miRNA-211-3p in TAC- or Ang II-induced hypertrophy, thus highlighting its potential use as a valuable biomarker for cardiac hypertrophy.
To further explore the underlying mechanism by which miR-211-3p regulates cardiac hypertrophy, we identified VEGFB as a novel target of miR-211-3p in cardiac hypertrophy. Blockade of VEGFB-VEGFR1 signalling enhanced the expression of hypertrophic markers, which reversed the miR-211 inhibitor-mediated downregulation of hypertrophic markers, suggesting that miR-211-3p regulated cardiac hypertrophy by targeting the VEGFB-VEGFR1 signalling pathway. These findings are consistent with recent observations that the activation of VEGFB-VEGFR1 alleviated cardiomyocyte hypertrophy induced by Ang II34, as well as with the report that VEGFB activates VEGFR1 to elicit a particular hypertrophic response in cultured cardiomyocytes and in infarcted hearts35. Moreover, SDF-1α-CXCR4 signalling is a crucial signalling cascade in cardiovascular disorders, including cardiac hypertrophy36, and CXCR4 deletion exacerbates isoproterenol-induced cardiac dysfunction and hypertrophy26. In addition, evidence suggests that VEGFR1/CXCR4 may mediate local inflammation and augment tissue perfusion by initiating a cascade of events, including the local release of SDF-137. VEGF is known to recruit bone marrow-derived inflammatory cells to ischaemic vessels with SDF-1, facilitating their retention via the chemokine receptor CXCR438. Here, we observed that knockdown of miR-211-3p enhanced the expression of VEGFB and SDF-1α. In summary, miR-211-3p-mediated regulation of cardiac hypertrophy was dependent on regulating the VEGFB-SDF-1α-CXCR4 signalling axis.
Salmena et al. proposed a ceRNA hypothesis that ceRNAs, such as pseudogenes, lncRNAs, and circular RNAs, contain microRNA response elements and sequester miRNAs to control mRNA transcription39. LncRNAs can act as ceRNAs to compete with miRNAs to suppress miRNA activity, thus modifying the expression of downstream genes in various physiological and pathological conditions40,41. Previous research showed that TINCR was significantly downregulated in hypertrophic mice9, which is consistent with the results we obtained in the TAC mouse model and Ang II-induced H9C2 model of cardiomyocyte hypertrophy. In contrast, they revealed that the lncRNA TINCR regulates PKCɛ expression in a miR-31-5p-dependent manner in cardiomyocyte hypertrophy. In our study, we provide evidence to demonstrate that TINCR is involved in cardiomyocyte hypertrophy by regulating VEGFB-SDF-1α-CXCR4 expression, which is regulated in a miR-211-3p-dependent manner.
In summary, TINCR functions as a ceRNA for miR-211-3p and improves cardiac hypertrophy by competitive combining miR-211-3p, thus relieving its suppressive effects on the VEGFB-SDF-1α-CXCR4 signalling axis. TINCR/miR-211-3p may act as a novel therapeutic target for the treatment of cardiac hypertrophy. However, this study presented some limitations. Although H9C2 is a widely used cell model for exploring the pathological process of cardiac hypertrophy, primary cardiomyocytes might be more representative of the adult heart and will be involved in our future investigations. In addition, since lncRNAs generally target several miRNAs and miRNAs generally have various downstream targets, other signalling cascades may be targeted by TINCR and miR-211-3p to regulate cardiac hypertrophy, which will be further explored in our future studies.
Data availability
The datasets used or analyzed during the current study are available from the corresponding author on reasonable request.
References
Samak, M. et al. Cardiac hypertrophy: an introduction to molecular and cellular basis. Med. Sci. Monit. Basic Res. 22, 75–79 (2016).
Frey, N. & Olson, E. N. Cardiac hypertrophy: the good, the bad, and the ugly. Annu. Rev. Physiol. 65, 45–79 (2003).
Nakamura, M. & Sadoshima, J. Mechanisms of physiological and pathological cardiac hypertrophy. Nat. Rev. Cardiol. 15, 387–407 (2018).
Bisping, E., Wakula, P., Poteser, M. & Heinzel, F. R. Targeting cardiac hypertrophy: toward a causal heart failure therapy. J. Cardiovasc. Pharmacol. 64, 293–305 (2014).
Braunwald, E. The war against heart failure: the Lancet lecture. Lancet 385, 812–824 (2015).
Liu, L., Zhang, D. & Li, Y. LncRNAs in cardiac hypertrophy: from basic science to clinical application. J. Cell. Mol. Med. 24, 11638–11645 (2020).
Hobuss, L., Bar, C. & Thum, T. Long non-coding RNAs: at the heart of cardiac dysfunction? Front. Physiol. 10, 30 (2019).
Shao, M. et al. LncRNA TINCR attenuates cardiac hypertrophy by epigenetically silencing CaMKII. Oncotarget 8, 47565–47573 (2017).
Li, H. et al. LncRNA Tincr regulates PKCvarepsilon expression in a miR-31-5p-dependent manner in cardiomyocyte hypertrophy. Naunyn Schmiedebergs Arch. Pharmacol. https://doi.org/10.1007/s00210-020-01847-9 (2020).
Hu, M., Han, Y., Zhang, Y., Zhou, Y. & Ye, L. lncRNA TINCR sponges miR-214-5p to upregulate ROCK1 in hepatocellular carcinoma. BMC Med. Genet. 21, 2 (2020).
Zheng, J. et al. lncRNA-TINCR functions as a competitive endogenous RNA to regulate the migration of mesenchymal stem cells by sponging miR-761. Biomed. Res. Int. 2020, 9578730 (2020).
Liu, X., Ma, J., Xu, F. & Li, L. TINCR suppresses proliferation and invasion through regulating miR-544a/FBXW7 axis in lung cancer. Biomed. Pharmacother. 99, 9–17 (2018).
Da Costa Martins, P. A. & De Windt, L. J. MicroRNAs in control of cardiac hypertrophy. Cardiovasc. Res. 93, 563–572 (2012).
Wang, J. & Yang, X. The function of miRNA in cardiac hypertrophy. Cell. Mol. Life Sci. 69, 3561–3570 (2012).
Heymans, S. et al. Macrophage microRNA-155 promotes cardiac hypertrophy and failure. Circulation 128, 1420–1432 (2013).
Qiu, Y. et al. MicroRNA-20b promotes cardiac hypertrophy by the inhibition of mitofusin 2-mediated Inter-organelle Ca(2+) cross-talk. Mol. Ther. Nucleic Acids 19, 1343–1356 (2020).
Nie, X. et al. miR-217 promotes cardiac hypertrophy and dysfunction by targeting PTEN. Mol. Ther. Nucleic Acids 12, 254–266 (2018).
Braile, M. et al. VEGF-A in cardiomyocytes and heart diseases. Int. J. Mol. Sci. 21, 5294 (2020).
Xu, X. H. et al. VEGF attenuates development from cardiac hypertrophy to heart failure after aortic stenosis through mitochondrial mediated apoptosis and cardiomyocyte proliferation. J. Cardiothorac. Surg. 6, 54 (2011).
Bry, M., Kivela, R., Leppanen, V. M. & Alitalo, K. Vascular endothelial growth factor-B in physiology and disease. Physiol. Rev. 94, 779–794 (2014).
Kivela, R. et al. VEGF-B-induced vascular growth leads to metabolic reprogramming and ischemia resistance in the heart. EMBO Mol. Med. 6, 307–321 (2014).
Claesson-Welsh, L. VEGF-B taken to our hearts: specific effect of VEGF-B in myocardial ischemia. Arterioscler. Thromb. Vasc. Biol. 28, 1575–1576 (2008).
Tang, J. M. et al. VEGF/SDF-1 promotes cardiac stem cell mobilization and myocardial repair in the infarcted heart. Cardiovas. Res. 91, 402–411 (2011).
Kajiyama, H. et al. Involvement of SDF-1alpha/CXCR4 axis in the enhanced peritoneal metastasis of epithelial ovarian carcinoma. Int. J. Cancer 122, 91–99 (2008).
Wen, J., Zhang, J. Q., Huang, W. & Wang, Y. SDF-1alpha and CXCR4 as therapeutic targets in cardiovascular disease. Am. J. Cardiovasc. Dis. 2, 20–28 (2012).
Wang, E. R. et al. Deletion of CXCR4 in cardiomyocytes exacerbates cardiac dysfunction following isoproterenol administration. Gene Ther. 21, 496–506 (2014).
Chen, Y. et al. Effects of wenxin keli on the action potential and L-type calcium current in rats with transverse aortic constriction-induced heart failure. Evid. Based Complement. Alternat. Med. 2013, 572078 (2013).
Shimizu, I. & Minamino, T. Physiological and pathological cardiac hypertrophy. J. Mol. Cell. Cardiol. 97, 245–262 (2016).
Shimizu, T. & Liao, J. K. Rho kinases and cardiac remodeling. Circ. J. 80, 1491–1498 (2016).
Wehbe, N. et al. MicroRNAs in cardiac hypertrophy. Int. J. Mol. Sci. 20, 4714 (2019).
Kong, Q. & Qiu, M. Long noncoding RNA SNHG15 promotes human breast cancer proliferation, migration and invasion by sponging miR-211-3p. Biochem. Biophys. Res. Commun. 495, 1594–1600 (2018).
Yan, D., Wu, F., Peng, C. & Wang, M. Silencing of LINC00284 inhibits cell proliferation and migration in oral squamous cell carcinoma by the miR-211-3p/MAFG axis and FUS/KAZN axis. Cancer Biol Ther. 22, 149–163 (2021).
Xu, J., Zhang, R. & Zhao, J. The novel long noncoding RNA TUSC7 Inhibits Proliferation by Sponging MiR-211 in colorectal cancer. Cell. Physiol. Biochem. 41, 635–644 (2017).
Shen, Z., Zhang, Z., Wang, X. & Yang, K. VEGFB-VEGFR1 ameliorates Ang II-induced cardiomyocyte hypertrophy through Ca(2+)-mediated PKG I pathway. J. Cell Biochem. 119, 1511–1520 (2018).
Zentilin, L. et al. Cardiomyocyte VEGFR-1 activation by VEGF-B induces compensatory hypertrophy and preserves cardiac function after myocardial infarction. FASEB J 24, 1467–1478 (2010).
Jackson, E. K. et al. SDF-1alpha (Stromal Cell-Derived Factor 1alpha) induces cardiac fibroblasts, renal microvascular smooth muscle cells, and glomerular mesangial cells to proliferate, cause hypertrophy, and produce collagen. J. Am. Heart Assoc. 6, e007253 (2017).
Wragg, A. et al. VEGFR1/CXCR4-positive progenitor cells modulate local inflammation and augment tissue perfusion by a SDF-1-dependent mechanism. J. Mol. Med. 86, 1221–1232 (2008).
Jin, D. K. et al. Cytokine-mediated deployment of SDF-1 induces revascularization through recruitment of CXCR4+ hemangiocytes. Nat. Med. 12, 557–567 (2006).
Salmena, L., Poliseno, L., Tay, Y., Kats, L. & Pandolfi, P. P. A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell 146, 353–358 (2011).
Zhao, D. et al. Identification of a six-lncRNA signature based on a competing endogenous RNA network for predicting the risk of tumour recurrence in bladder cancer patients. J. Cancer 11, 108–120 (2020).
Wu, T. et al. Long noncoding RNA (lncRNA) CTTN-IT1 elevates skeletal muscle satellite cell proliferation and differentiation by acting as ceRNA for YAP1 Through Absorbing miR-29a in Hu sheep. Front. Genet. 11, 843 (2020).
Author contributions
S.T.: concepts, design, experimental studies, data acquisition, data analysis, editing;. X.-Y.W.: data analysis;. L.-X.Z.: experimental studies, data analysis. Z.-J.S. data acquisition, editing;. Z.-H.Z.: supervision, concepts, review. All the authors approved the final version.
Funding
This work was supported by the General Projects of the National Natural Science Foundation of China (81870303).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
Animal experiments were in accordance with National Institutes of Health guidelines and approved by the Animal Care and Use Committee of the Third Xiangya Hospital of Central South University.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Tu, S., Wang, XY., Zeng, LX. et al. LncRNA TINCR improves cardiac hypertrophy by regulating the miR-211-3p-VEGFB-SDF-1α-CXCR4 pathway. Lab Invest 102, 253–262 (2022). https://doi.org/10.1038/s41374-021-00678-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41374-021-00678-3