Abstract
Genetic disorders are a leading cause of morbidity and mortality in infants admitted to neonatal intensive care units. This population has immense potential to benefit from genomic medicine, as early precision diagnosis is critical to early personalized management. However, the implementation of genomic medicine in neonatology thus far has arguably worsened health inequities, and strategies are urgently needed to achieve equitable access to genomics in neonatal care. In this perspective, we demonstrate the utility of genomic sequencing in critically ill infants and highlight three key recommendations to advance equitable access: recruitment of underrepresented populations, education of non-genetics providers to empower practice of genomic medicine, and development of innovative infrastructure to implement genomic medicine across diverse settings.
Similar content being viewed by others
Introduction
Genomic medicine (GM) has rapidly advanced since exome sequencing was first utilized for patient diagnosis in 2010 [1]. Critically ill infants admitted to neonatal intensive care units (NICUs) represent a population with high rates of genetic disorders and associated morbidity and mortality, and arguably have the greatest potential to benefit from GM, as early diagnosis is critical to optimizing the benefits of early tailored management, including a growing number of pioneering personalized therapies [2]. Genomic (exome or genome) sequencing (GS) has the potential to transform neonatal care, with multiple studies demonstrating diagnostic, clinical, personal, and informational utility for critically ill infants with underlying genetic disorders and their families [2, 3]. However, the implementation of GM in clinical care thus far has arguably worsened health inequities, and strategies are urgently needed to achieve equitable access in this critical population [4, 5].
Potential of genomic medicine
Genetic disorders contribute to significant morbidity—with often lifelong consequences—and mortality during infancy. The advances in obstetric care and neonatology over the past several decades have reduced morbidity and mortality (M&M) from other perinatal conditions (e.g., surfactant administration for respiratory distress syndrome), and the leading causes of infant mortality in the United States are now reported to be genetic [6, 7]. Reducing M&M due to genetic disorders will require precision medicine approaches, for which the first step is identifying the underlying genetic diagnoses. Up to 25% of critically ill infants in NICUs may have an undiagnosed genetic condition; the majority undiagnosed due to limited access to GS [8,9,10,11].
Broadly, any critically ill infant with a disease of unknown etiology may be suspected to have an underlying genetic disorder and considered for GS. Based on previous studies and our own experience, specific phenotypic criteria in the neonatal period to prioritize for GS may include multiple congenital anomalies (or a single major anomaly with accompanying syndromic features) as well as neurologic, metabolic, or other severe organ system abnormalities of unknown etiology [12,13,14]. A recent evidence-based guideline from the American College of Medical Genetics and Genomics recommends GS for infants (<1 year old) with one or more congenital anomalies [15]. However, it is important to remember that genetic disorders may present with nonspecific clinical findings during the neonatal period and certain clinical features associated with genetic disorders may not present until later in life. Ideally, GS should be considered as a comprehensive diagnostic test for any critically ill infant who lacks a clear non-genetic explanation for his or her presentation.
Rapid GS has multiple potential benefits for critically ill infants with suspected genetic disorders and their families. First, GS may lead to a genetic diagnosis and end the diagnostic odyssey (diagnostic utility). A recent review of 31 studies of rapid GS in neonatal and pediatric patients in intensive care settings reported a weighted average diagnostic rate of 36% [2]. Second, GS may lead to change(s) in clinical management (clinical utility); the same review reported a weighted average change in management rate of 27% [2]. A genetic diagnosis may influence treatment (lead to starting, changing, or stopping treatment), workup (lead to new workup or avoidance of unnecessary workup), and goals of care (e.g., transition to comfort care). Further, a genetic diagnosis may enable access to etiology-specific research like natural history studies and clinical trials of emerging precision therapies. We also note that nondiagnostic GS has been reported to lead to changes in management in some cases [16]. Third, a genetic diagnosis may influence prognostic and reproductive counseling (informational utility); for example, the likelihood of developmental delay/intellectual disability for the former and recurrence risk for the latter. Fourth, a genetic diagnosis may provide additional patient-reported benefits (personal utility) for the infant and family [17]. Finally, we note that recent studies have demonstrated the cost-effectiveness of GS in the NICU setting [16, 18].
Thus, while we recognize that inequitable access to genomic medicine is present across our health care system, we believe that GS currently has the greatest potential to impact medical care of the NICU population. In this perspective, we use critically ill infants in NICUs as a paradigm population to describe barriers to equitable access to genomic medicine and recommendations to overcome those barriers. When referring to GS below, we are specifically referring to “rapid” GS that can provide clinically accredited results in ≤1–2 weeks.
Barriers to implementation of genomic medicine
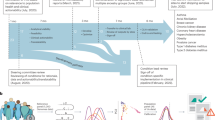
Barriers to equitable access exist at multiple steps of the GM implementation process, and it is important to acknowledge that racism on the internalized, interpersonal, institutional, and structural levels may be a barrier at every step [19, 20] (Fig. 1). Race and ethnicity are social, and not biological, constructs and are different from genetic ancestry; however, there are inequities in health care related to race and ethnicity [21].
Barriers to equitable access exist at multiple steps of the genomic medicine process, from identification of patients for genomic sequencing to return of genomic sequencing results and follow up. It is important to acknowledge that racism on the internalized, interpersonal, institutional, and structural levels may be a barrier at every step.
First, neonatal providers need to suspect that an admitted infant has an underlying genetic disorder and decide that GS is an appropriate test. At community and/or rural NICUs that often provide care for infants from (1) racial and ethnic minority populations, (2) lower income households, and/or (3) underserved areas, providers may not have access to clinical geneticists or genetic counselors (GCs) to assist with this process [22, 23]. Providers without formal training in clinical genetics may lack knowledge about genetics and genomics, have difficulty appropriately identifying infants with suspected genetic disorders, and/or have mixed attitudes toward genetic testing [24, 25]. Moreover, the “classic” dysmorphic features that may raise suspicion for certain genetic disorders are based on Northern European populations, and infants of non-European ancestry may not present with similar features [26]. Thus, infants from underrepresented and underserved populations may be less likely to be identified for GS.
Next, GS needs to be clinically available and approved. While neonatal intensive care occurs across a range of settings (e.g., rural and urban, community and academic), GS is mainly available at large academic referral centers that have the resources and expertise to sustainably carry out this process. Infants at community and/or rural NICUs where GS is unavailable may receive limited or no genetic workup or be transferred to referral centers for comprehensive genetic workup, which may impose additional burdens on families and costs to the healthcare system. At NICUs where GS is available, approval for testing is often required from an institutional committee and/or insurer. The presentation of a genetic disorder in a neonate may be nonspecific and difficult to discriminate from other causes of critical illness, and as noted above, may be more difficult to recognize in infants underrepresented in dysmorphology atlases. Thus, GS requests for infants from underrepresented populations may be less likely to be submitted to or approved by institutional committees. In addition, insurer approval remains a barrier, which may be exacerbated for infants with lower household income and public (Medicaid) insurance [27, 28].
Before ordering GS, a provider needs to consent and provide pre-test counseling to the infant’s family. This is often done by clinical geneticists or GCs, where available, and non-genetics providers (NGPs) may be uncomfortable with this process. Furthermore, racial and ethnic minority families may be more likely to have mistrust in genetic testing and/or the healthcare system due to systemic racism and historical injustices, as well as have language, literacy, cultural, and additional barriers to consent [29]. If the family consents, samples need to be collected and shipped to the sequencing laboratory. In the NICU, the infant sample is relatively easy to collect, but parental samples, which improve results interpretation, may be harder to collect for families who have transportation, childcare, work and/or other barriers that may limit visitation.
In the US, a Clinical Laboratory Improvement Amendments (CLIA) certified laboratory performs sequencing and analysis based on standardized diagnostic criteria [30], and rapid GS results are usually reported in ≤1–2 weeks. Although race and ethnicity are socially, not genetically, defined constructs, studies have reported that racial and ethnic minority infants may be more likely to have GS results that require additional interpretation or to receive nondiagnostic GS results [31, 32]. This may be in part due to the underrepresentation of non-European individuals in genetic variant databases leading to greater difficulty interpreting results for the subset of racial and ethnic minority infants of non-European ancestry [29, 31]. NGPs may also be uncomfortable with results return and post-test counseling, including making management changes based on the results, again limiting GS utilization outside of centers where clinical geneticists and/or GCs are available. Finally, identified genetic diagnoses often require complex follow up after NICU discharge by clinical geneticists and additional subspecialists, which may be difficult for families from underserved populations to access [33].
Recommendations for equitable access in NICUs
Defining and demonstrating the utility of GS in NICUs is an important ongoing research effort and provides evidence for institutional and insurer policies, which is necessary for achieving equitable access to these tests. Project Baby Bear provides an informative example: the study enrolled critically ill infants who were Medi-Cal beneficiaries (California’s Medicaid program), performed payer-funded rapid GS, and reported that GS led to changes in management and reduced healthcare costs [16]. Subsequently, the California government passed a bill to provide money to reimburse rapid GS in Medicaid-covered infants in 2022 [2].
In addition, we need to consciously build up research efforts focused on overcoming the many other barriers to equitable access to genomic medicine in NICUs. In the short-term, such efforts will provide urgently needed access to GS for critically ill infants who may otherwise not have access, and in the long term, will identify strategies for sustainable implementation of genomic medicine across diverse NICU settings. We recommend the following (Fig. 2).
Recruitment of underrepresented populations
Racial and ethnic minority participants are historically underrepresented in genomics research and populations of non-European ancestry are underrepresented in genetic variant databases [34, 35]. GS studies of critically ill infants have traditionally recruited from large academic referral centers and some initial studies did not report the racial and ethnic distribution of participants [36,37,38]. Recent studies have reported enrolling mostly infants from underserved populations (Project Baby Bear [16]) and infants with racial and ethnic distribution reflective of the US population (NICUSeq [39]). Future studies need to set recruitment diversity targets—the Clinical Sequencing Evidence-Generating Research Consortium (CSER) studies committed to at least 60% of participants being of non-European ancestry or from underserved areas—which will require recruitment from community and/or rural NICUs historically underrepresented in genomics research (ongoing CSER SouthSeq [31] and VIGOR [Virtual Genome Center for Infant Health] studies, clinicaltrials.gov IDs NCT03842995 and NCT05205356). Studies need to evaluate education and communication strategies for families with literacy, language, cultural, and other barriers that disproportionately impact racial and ethnic minority and underserved populations and engage with these families to understand their attitudes toward GS and build trust in genomic medicine during early interactions with the pediatric healthcare system. One-on-one interviews and focus groups, using certified interpreters for families with limited English proficiency and using plain language especially for families with lower health literacy, can be used to explore potential recruitment barriers, including the ethical, legal, and social implications of GS in these populations. It is important to understand how racial and ethnic minority populations who have been subjected to systemic racism and historical medical exploitation may feel about participating in GS studies, including providing samples for GS and sharing sequencing data with researchers and/or clinicians. Findings can be used to optimize study design and materials; for example, recruitment materials can be refined to better address recurrently brought up concerns and counseling materials can be adapted to use examples that resonate with the participant population. The ongoing SouthSeq study used diversity studios and literacy experts to optimize study materials for participants; 73% were non-white or from underserved areas [29].
Education of NGPs
Given limited availability of clinical geneticists and GCs and reported discomfort with genomic medicine among NGPs, studies need to provide education on genomic medicine to NGPs and evaluate subsequent comfort with the genomic medicine process, including identifying appropriate infants for GS, consenting families, and returning results to families (ongoing SouthSeq and VIGOR studies). Given the many demands on neonatal providers, education can be provided via virtual platforms that allow for asynchronous learning. To lower cognitive burden, GS decision support tools that NGPs can use in real time in the NICU can be developed and adapted to provide NICU-specific GS workflows. These can range from “big picture” criteria for when to consider GS in the NICU population to “little details” steps on consenting parents, ordering and collecting GS samples in a specific NICU system, and returning the different types of GS results (e.g., positive result, negative result, secondary finding, variant of uncertain significance). Preliminary results from SouthSeq, which randomized return of GS results to families by GCs or NGPs, demonstrated that NGPs reported increased confidence in interpreting results and managing care based on results after receiving education and training [40].
Development of infrastructure
Studies need to implement innovative infrastructure like virtual genomic medicine platforms and NICU “GS champions”, which have the potential to provide genomics education and expertise to NGPs and patients outside of large academic referral centers. As suggested by early implementation studies [41, 42], a NICU “GS champion”—for example, a NGP who has received basic education in genomic medicine and is passionate about GS—can assist in identifying appropriate infants for GS and consenting parents. For most centers, “in center” GS may not be feasible; rather, providers can order GS and send out the patient, and where available, parent samples to a CLIA-certified vendor who can rapidly perform, analyze, and provide initial interpretation of GS. Subsequently, providers receive a GS report and can return results to families. Depending on provider comfort and availability of center clinical geneticists or GCs, the NICU “GS champion” and/or virtual consulting genetics professionals can assist with consent and/or return of results as needed. Ideally, NGPs will gain comfort with the GS process and need to utilize virtual genetics professionals less over time for consent and initial return of results. The COVID-19 pandemic expanded the use of virtual medicine, which provides an opportunity to keep infants in the NICUs where they routinely receive care, removing cost and time barriers for families when infants are transferred, and ideally empowering NGPs who have developed trusting relationships with families to provide genomic care (ongoing VIGOR study). It will be critical to rigorously evaluate implementing these infrastructures using equity-focused methodology to identify strategies for sustainable genomic care after research studies end.
Conclusions
Genomic medicine is a powerful diagnostic tool in NICUs, but a large gap exists between the research setting, where a critically ill infant can be diagnosed in less than a day using the newest technologies [43], and “real world” clinical care, where many critically ill infants, and disproportionally racial and ethnic minority infants, do not have access to GS. There is an urgent need to implement equitable access to genomic medicine for critically ill infants, and we highlight three key recommendations for research efforts to overcome barriers: recruitment of underrepresented populations, education of NGPs to empower practice of genomic medicine, and development of innovative infrastructure to implement genomic medicine across diverse settings. These efforts have the potential to address inequities at the earliest stages of life and impact health outcomes. However, this potential can only be realized with sustained access to GS after research studies end and to follow up care, which will require institutional and insurer recognition of the utility of genomic medicine and continued efforts to overcome barriers to inequitable access to pediatric healthcare more broadly.
References
Ng SB, Buckingham KJ, Lee C, Bigham AW, Tabor HK, Dent KM, et al. Exome sequencing identifies the cause of a mendelian disorder. Nat Genet. 2010;42:30–5.
Kingsmore SF, Cole FS. The role of genome sequencing in neonatal intensive care units. Annu Rev Genomics Hum Genet. 2022;23:427–48.
Callahan KP, Mueller R, Flibotte J, Largent EA, Feudtner C. Measures of utility among studies of genomic medicine for critically ill infants: a systematic review. JAMA Netw Open. 2022;5:e2225980.
Amendola LM, Berg JS, Horowitz CR, Angelo F, Bensen JT, Biesecker BB, et al. The Clinical Sequencing Evidence-Generating Research Consortium: integrating genomic sequencing in diverse and medically underserved populations. Am J Hum Genet. 2018;103:319–27.
Jooma S, Hahn MJ, Hindorff LA, Bonham VL. Defining and achieving health equity in genomic medicine. Ethn Dis. 2019;29(Suppl 1):173–8.
Almli LM, Ely DM, Ailes EC, Abouk R, Grosse SD, Isenburg JL, et al. Infant mortality attributable to birth defects—United States, 2003-2017. Morb Mortal Wkly Rep. 2020;69:25–9.
Heron M. Deaths: leading causes for 2019. Natl Vital Stat Rep. 2021;70:1–114.
Hays T, Wapner RJ. Genetic testing for unexplained perinatal disorders. Curr Opin Pediatr. 2021;33:195–202.
Maron JL, Kingsmore SF, Wigby K, Chowdhury S, Dimmock D, Poindexter B, et al. Novel variant findings and challenges associated with the clinical integration of genomic testing: an interim report of the genomic medicine for ill neonates and infants (GEMINI) study. JAMA Pediatr. 2021;175:e205906.
Wojcik MH, Schwartz TS, Thiele KE, Paterson H, Stadelmaier R, Mullen TE, et al. Infant mortality: the contribution of genetic disorders. J Perinatol. 2019;39:1611–9.
Wojcik MH, Schwartz TS, Yamin I, Edward HL, Genetti CA, Towne MC, et al. Genetic disorders and mortality in infancy and early childhood: delayed diagnoses and missed opportunities. Genet Med. 2018;20:1396–404.
D’Gama AM, Del Rosario MC, Bresnahan MA, Yu TW, Wojcik MH, Agrawal PB. Integrating rapid exome sequencing into NICU clinical care after a pilot research study. NPJ Genom Med. 2022;7:51.
Gubbels CS, VanNoy GE, Madden JA, Copenheaver D, Yang S, Wojcik MH, et al. Prospective, phenotype-driven selection of critically ill neonates for rapid exome sequencing is associated with high diagnostic yield. Genet Med. 2020;22:736–44.
Wojcik MH, D’Gama AM, Agrawal PB. A model to implement genomic medicine in the neonatal intensive care unit. J Perinatol. 2023;43:248–52.
Manickam K, McClain MR, Demmer LA, Biswas S, Kearney HM, Malinowski J, et al. Exome and genome sequencing for pediatric patients with congenital anomalies or intellectual disability: an evidence-based clinical guideline of the American College of Medical Genetics and Genomics (ACMG). Genet Med. 2021;23:2029–37.
Dimmock D, Caylor S, Waldman B, Benson W, Ashburner C, Carmichael JL, et al. Project Baby Bear: rapid precision care incorporating rWGS in 5 California children’s hospitals demonstrates improved clinical outcomes and reduced costs of care. Am J Hum Genet. 2021;108:1231–8.
Hayeems RZ, Luca S, Assamad D, Bhatt A, Ungar WJ. Utility of genetic testing from the perspective of parents/caregivers: a scoping review. Children. 2021;8:259.
Diaby V, Babcock A, Huang Y, Moussa RK, Espinal PS, Janvier M, et al. Real-world economic evaluation of prospective rapid whole-genome sequencing compared to a matched retrospective cohort of critically ill pediatric patients in the United States. Pharmacogenomics J. 2022;22:223–9.
Fraiman YS, Wojcik MH. The influence of social determinants of health on the genetic diagnostic odyssey: who remains undiagnosed, why, and to what effect? Pediatr Res. 2021;89:295–300.
Montoya-Williams D, Fraiman YS, Pena MM, Burris HH, Pursley DM. Antiracism in the field of neonatology: a foundation and concrete approaches. Neoreviews. 2022;23:e1–12.
Flanagin A, Frey T, Christiansen SL, Committee AMAMoS. Updated guidance on the reporting of race and ethnicity in medical and science journals. JAMA. 2021;326:621–7.
Hawkins AK, Hayden MR. A grand challenge: providing benefits of clinical genetics to those in need. Genet Med. 2011;13:197–200.
Jenkins BD, Fischer CG, Polito CA, Maiese DR, Keehn AS, Lyon M, et al. The 2019 US medical genetics workforce: a focus on clinical genetics. Genet Med. 2021;23:1458–64.
Diamonstein C, Stevens B, Shahrukh Hashmi S, Refuerzo J, Sullivan C, Hoskovec J. Physicians’ awareness and utilization of genetic services in Texas. J Genet Couns. 2018;27:968–77.
Haga SB, Kim E, Myers RA, Ginsburg GS. Primary care physicians’ knowledge, attitudes, and experience with personal genetic testing. J Pers Med. 2019;9:29.
Koretzky M, Bonham VL, Berkman BE, Kruszka P, Adeyemo A, Muenke M, et al. Towards a more representative morphology: clinical and ethical considerations for including diverse populations in diagnostic genetic atlases. Genet Med. 2016;18:1069–74.
Lee G, Yu L, Suarez CJ, Stevenson DA, Ling A, Killer L. Factors associated with the time to complete clinical exome sequencing in a pediatric patient population. Genet Med. 2022;24:2028–33.
Suther S, Kiros GE. Barriers to the use of genetic testing: a study of racial and ethnic disparities. Genet Med. 2009;11:655–62.
Gutierrez AM, Robinson JO, Outram SM, Smith HS, Kraft SA, Donohue KE, et al. Examining access to care in clinical genomic research and medicine: experiences from the CSER Consortium. J Clin Transl Sci. 2021;5:e193.
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–24.
Bowling KM, Thompson ML, Finnila CR, Hiatt SM, Latner DR, Amaral MD, et al. Genome sequencing as a first-line diagnostic test for hospitalized infants. Genet Med. 2022;24:851–61.
Smith HS, Swint JM, Lalani SR, de Oliveira Otto MC, Yamal JM, Russell HV, et al. Exome sequencing compared with standard genetic tests for critically ill infants with suspected genetic conditions. Genet Med. 2020;22:1303–10.
Bohnhoff JC, Taormina JM, Ferrante L, Wolfson D, Ray KN. Unscheduled referrals and unattended appointments after pediatric subspecialty referral. Pediatrics. 2019;144:e20190545.
Landry LG, Ali N, Williams DR, Rehm HL, Bonham VL. Lack of diversity in genomic databases is a barrier to translating precision medicine research into practice. Health Aff. 2018;37:780–5.
Popejoy AB, Fullerton SM. Genomics is failing on diversity. Nature. 2016;538:161–4.
Meng L, Pammi M, Saronwala A, Magoulas P, Ghazi AR, Vetrini F, et al. Use of exome sequencing for infants in intensive care units: ascertainment of severe single-gene disorders and effect on medical management. JAMA Pediatr. 2017;171:e173438.
Willig LK, Petrikin JE, Smith LD, Saunders CJ, Thiffault I, Miller NA, et al. Whole-genome sequencing for identification of Mendelian disorders in critically ill infants: a retrospective analysis of diagnostic and clinical findings. Lancet Respir Med. 2015;3:377–87.
French CE, Delon I, Dolling H, Sanchis-Juan A, Shamardina O, Megy K, et al. Whole genome sequencing reveals that genetic conditions are frequent in intensively ill children. Intensive Care Med. 2019;45:627–36.
Group NIS, Krantz ID, Medne L, Weatherly JM, Wild KT, Biswas S, Devkota B, et al. Effect of whole-genome sequencing on the clinical management of acutely ill infants with suspected genetic disease: a randomized clinical trial. JAMA Pediatr. 2021;175:1218–26.
East KM, Cochran ME, Kelley WV, Greve V, Finnila CR, Coleman T, et al. Education and training of non-genetics providers on the return of genome sequencing results in a NICU setting. J Pers Med. 2022;12:405.
Best S, Brown H, Lunke S, Patel C, Pinner J, Barnett CP, et al. Learning from scaling up ultra-rapid genomic testing for critically ill children to a national level. NPJ Genom Med. 2021;6:5.
Franck LS, Kriz RM, Rego S, Garman K, Hobbs C, Dimmock D. Implementing rapid whole-genome sequencing in critical care: a qualitative study of facilitators and barriers to new technology adoption. J Pediatr. 2021;237:237–43. e2.
Owen MJ, Niemi AK, Dimmock DP, Speziale M, Nespeca M, Chau KK, et al. Rapid sequencing-based diagnosis of thiamine metabolism dysfunction syndrome. N Engl J Med. 2021;384:2159–61.
Funding
AMD is supported by NICHD T32 HD098061. PBA is supported by NHGRI R01 HG011798.
Author information
Authors and Affiliations
Contributions
AMD and PBA conceptualized the manuscript, AMD drafted the manuscript, and AMD and PBA critically reviewed and revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
D’Gama, A.M., Agrawal, P.B. Role of genomic medicine and implementing equitable access for critically ill infants in neonatal intensive care units. J Perinatol 43, 963–967 (2023). https://doi.org/10.1038/s41372-023-01630-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41372-023-01630-7
This article is cited by
-
Genomic medicine in neonatal care: progress and challenges
European Journal of Human Genetics (2023)