Abstract
The AML1-ETO fusion protein (A/E), which results from the t(8;21) translocation, is considered to be a leukemia-initiating event. Identifying the mechanisms underlying the oncogenic activity of A/E remains a major challenge. In this study, we identified a specific down-regulation of brain acid-soluble protein 1 (BASP1) in t(8;21) acute myeloid leukemia (AML). A/E recognized AML1-binding sites and recruited DNA methyltransferase 3a (DNMT3a) to the BASP1 promoter sequence, which triggered DNA methylation-mediated silencing of BASP1. Ectopic expression of BASP1 inhibited proliferation and the colony-forming ability of A/E-positive AML cell lines and led to apoptosis and cell cycle arrest. The DNMT inhibitor decitabine up-regulated the expression of BASP1 in A/E-positive AML cell lines. In conclusion, our data suggest that BASP1 silencing via promoter methylation may be involved in A/E-mediated leukemogenesis and that BASP1 targeting may be an actionable therapeutic strategy in t(8;21) AML.
Similar content being viewed by others
Introduction
Brain acid-soluble protein 1 (BASP1) (CAP-23, NAP-22) was initially identified as an abundant membrane-bound protein in the brain and was thought to play a role in axon regeneration and neuronal plasticity1,2,3. BASP1 has also been found in other cell types and at other subcellular locations, and its functions in transcriptional regulation, apoptosis, and differentiation have also been recently reported4,5,6.
In recent years, studies have shown that abnormal expression and methylation status of the BASP1 gene are closely related to the occurrence and prognosis of certain tumors and that BASP1 may also play a role as a tumor suppressor gene. For instance, in hepatocellular carcinoma (HCC), BASP1 expression has been shown to be down-regulated via promoter methylation; the methylation status of BASP1 could be used in the early diagnosis of HCC7. BASP1 gene methylation and its expression level are also associated with melanoma progression8.
The t(8;21)(q22;q22) chromosome translocation is one of the most common cytogenetic abnormalities in acute myeloid leukemia (AML), accounting for approximately 7–12% of newly diagnosed AML cases9,10,11. The new fusion protein AML1-ETO (A/E) is considered to be the leukemia-initiating event and also gives this subtype of AML unique molecular characteristics. The A/E fusion protein has been shown to bind specific DNA sequences and recruit DNA methyltransferases (DNMTs) to target genes, resulting in A/E-associated DNA methylation12, 13. Recent studies have shown that t(8;21) AML has a unique methylation pattern and that this abnormal methylation and silencing of tumor suppressor genes may be an important factor leading to the occurrence and development of this disease14. Here, we studied the expression of BASP1 in various AML cell lines and identified a specific down-regulation and promoter methylation of the BASP1 gene in t(8;21) AML. Our results suggest that methylation of the BASP1 gene may be regulated by the A/E fusion protein and play a role in A/E-driven leukemogenesis.
In this study, we showed that the A/E fusion protein directly bound to the promoter of the BASP1 gene. The A/E fusion protein recruited DNMT3a to the promoter region of BASP1, leading to methylation of the CpG island. As a result, the BASP1 gene was silenced. Upregulation of the BASP1 gene by overexpression or treatment with the DNMT inhibitor decitabine (DAC) was able to induce cell death in t(8;21) AML cell lines. Our findings suggest that in t(8;21) AML, BASP1 plays a tumor suppressive role in vitro and that targeting BASP1 may be an actionable treatment option in treating t(8;21) AML.
Materials and methods
Cell lines and clinical samples
Kasumi-1, HL-60, NB4, U937, THP-1, KG-1, MV-4-11, and HEL cells were cultured in RPMI 1640 (HyClone, GE Healthcare, Logan, UT, USA) medium containing 10% fetal calf serum (HyClone, GE Healthcare, Logan, UT, USA). SKNO-1 cells were cultured in RPMI 1640 (Gibco, Fisher Scientific, Waltham, MA, USA) containing 10% fetal calf serum (Gibco, Fisher Scientific, Waltham, MA, USA) and 10 ng/mL granulocyte–macrophage colony-stimulating factor. The 293T cells were cultured in Dulbecco's modified Eagle's medium (HyClone, GE Healthcare, Logan, UT, USA) containing 10% fetal calf serum (HyClone, GE Healthcare, Logan, UT, USA). SKNO-1-siA/E (A/E silenced by the lentiviral vector pRRL.cPPT.hPGK in SKNO-1 cells) and U937-A/E (produced by U937 electroporated with a hemagglutinin (HA)-tagged A/E complementary DNA (cDNA) and subcloned into a vector carrying the Zn2+-inducible mouse MT-1 promoter) cells were cultured in RPMI 1640 (HyClone, GE Healthcare, Logan, UT, USA) containing 10% fetal calf serum (HyClone, GE Healthcare, Logan, UT, USA) as described in a previous study13. U937-A/E cells were treated with 100 µM ZnSO42+ (Invitrogen, Carlsbad, CA, USA) for 16 h to induce A/E expression. Bone marrow samples from patients and healthy donors were obtained with informed consent in accordance with the principles of the Declaration of Helsinki. This study was approved by the Human Subject Ethics Committee of the Chinese PLA General Hospital.
RNA extraction and quantitative real-time polymerase chain reaction
Total RNA was extracted from cells using Trizol reagent (Invitrogen, Carlsbad, CA, USA), and cDNA was obtained after reverse transcription. Expression of mRNA was quantified by SYBR Green real-time quantitative polymerase chain reaction (qRT-PCR) analysis (Takara, Tokyo, Japan) on an Mx3000P Lightcycler (Stratagene, LaJolla, CA, USA). Gene expression was expressed as 2−ΔCT relative to GAPDH. The primers used are listed in Supplementary Table 1.
DNA extraction, BSP, and MSP
Genomic DNA from patients, healthy donors, and cell lines was isolated with the Wizard® Genomic DNA Purification Kit (Promega, Madison, WI, USA). Genomic DNA was treated with sodium bisulfate according to the manufacturer’s instructions supplied with the EpiTect Bisulfite Kit (Qiagen, Hilden, Germany). The bisulfite sequencing PCR (BSP) and methylation-specific PCR (MSP) primer sequences were designed by Methprimer and are shown in Supplementary Table 1. For BSP, amplified PCR products were purified and cloned into pGEM-T vectors (Promega, Madison, WI, USA), and five positive clones were sequenced for the analysis of DNA methylation. MSP was conducted in a 25 μL reaction solution with GoTaq Green Master Mix (Promega, Madison, WI, USA), and PCR products were analyzed on 2% agarose gels.
Luciferase reporter assays
DNA fragments were amplified by PCR of human genomic DNA. The primer sequences are shown in Supplementary Table 1. All DNA fragments were inserted into the pGL3-LUC reporter vector (Promega, Madison, WI, USA). Mutants were generated using overlap-extension PCR. The SuperFect reagent (Qiagen, Hilden, Germany) was used to co-transfect 293T cells at a density of 2 × 105 with the pcDNA3 vectors (0, 10, 50, or 100 ng) with or without A/E cDNA and plated in 24-well plates. The pRL-TK renilla luciferase reporter vector (Promega, Madison, WI, USA) was used as a control. According to the manufacturer’s instructions, cells harvested 48 h after transfection were analyzed using the Dual-Luciferase® Reporter assay (Promega, Madison, WI, USA).
ChIP assays
Chromatin immunoprecipitation (ChIP) assays were performed as described previously using the EZ-ChIP Assay Kit (Millipore, Billerica, MA, USA) according to the manufacturer’s instructions15. Briefly, SKNO-1 and SKNO-1-siA/E cells were seeded at a density of 2 × 106 and incubated for 10 min at 37 °C to crosslink the proteins to DNA after treatment with formaldehyde (1% final concentration). After sonication, chromatin was immunoprecipitated overnight with 5 µg of antibodies against AML1 (Santa Cruz, Dallas, TX, USA), ETO (Santa Cruz, Dallas, TX, USA), DNMT1 (Abcam, Cambridge, UK), DNMT3a (Abcam, Cambridge, UK), and DNMT3b (Abcam, Cambridge, UK). A negative control experiment was performed with a normal mouse immunoglobulin G (IgG) (Abcam, Cambridge, UK) antibody. DNA fragments obtained without antibody were used as the input controls. The genomic BASP1 upstream region close to the active AML1-binding site and unrelated negative control region were amplified by SYBR Green qRT-PCR analysis (Takara, Tokyo, Japan) on an Mx3000P Lightcycler (Stratagene, LaJolla, CA, USA). Their expression was quantified as 2−ΔCT relative to input. GAPDH served as an internal control to account for non specific precipitated sequences. The primer sequences are shown in Supplementary Table 1.
Western blot
Total protein extracts and western blot analysis were performed as previously described16. Rabbit polyclonal anti-BASP1 was purchased from Abcam (Cambridge, MA, USA, ab103315). Rabbit anti-human β-actin and horse radish peroxidase-linked anti-rabbit IgG were purchased from Cell Signaling Technology (Boston, MA, USA, 4970S). Densitometric analysis was performed using UVITEC-1D for quantification.
Cell proliferation, apoptosis, differentiation, and colony-forming unit analysis
Cell proliferation was evaluated by using the Cell-Counting Kit (CCK-8) (Dojindo, Japan), the concise method for which was that a minimum of 5000 cells per well cultured in 96-well plates were added to 10 μL of CCK-8 solution and incubated for another 3 h at 37 °C in a 5% CO2 atmosphere. Then, the absorbance was detected at a wavelength of 450 nm. The cell apoptosis assay was analyzed by a FACSCalibur flow cytometer (Becton Dickinson, Franklin Lakes, NJ, USA). For every sample, cells were collected in a BD Falcon tube and stained with 5 μL of Annexin V and 5 μL of 7AAD (Becton Dickinson, Franklin Lakes, NJ, USA). Cell differentiation was also assessed by a FACSCalibur flow cytometer (Becton Dickinson, Franklin Lakes, NJ, USA). Samples stained with anti-CD11b (Becton Dickinson, Franklin Lakes, NJ, USA) were analyzed using CellQuest Pro software (Becton Dickinson, Franklin Lakes, NJ, USA). For colony-forming unit assays, cells were plated in methylcellulose H4230 (Stemcell Technologies, Vancouver, BC, Canada) at a concentration of 1 × 103 cells per mL. The number of colonies with localized clusters of >50 cells were counted after incubation for 7 days. All experiments were performed in triplicate.
Statistical analysis
SPSS 13.0 software (SPSS Inc., Chicago, IL, USA) was used to process the data. All data are expressed as the mean ± standard deviation (SD) from at least three separate experiments. Student’s ttest was used to determine the statistical significance of the experimental results, and a p value ≤0.05 was considered to be significant.
Results
BASP1 is selectively silenced in A/E-positive AML
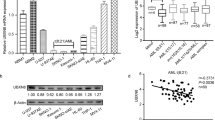
In an initial screen of BASP1 expression in AML, we examined the mRNA expression of BASP1 in nine AML cell lines (Kasumi-1, SKNO-1, NB4, U937, THP-1, KG-1, MV-4-11, HEL, and HL-60) and three sets of normal bone marrow (NBM) from healthy donors. Expression of BASP1 showed an interesting pattern: it was robustly down-regulated in A/E-positive AML cell lines (Kasumi-1 and SKNO-1) in contrast to the other A/E-negative AML cell lines and NBMs (Fig. 1a). The results were further validated in a published GEO microarray dataset (GSE13159) that included 60 t(8;21) AML, 54 t(15;17) AML, 47 inv16 or t(16;16) AML, 43 t(11q23)/MLL-rearrangement AML, 48 complex aberrant karyotype AML, and 5 hematopoietic stem cells (HSCs) (Fig. 1b). This trend of BASP1 was also confirmed by western blotting of the AML cell lines (Fig. 1c). Surprisingly, the expression of BASP1 was adversely correlated with the expression of AML1-ETO in t(8;21) AML (Fig. 1d). To determine the role of A/E in the regulation of BASP1 expression, A/E was knocked out in SKNO-1 (SKNO-1-siA/E) by using a lentiviral vector, and HA-tagged A/E cDNA was electroporated into U937 cells to produce the U937-A/E-HA clone in a zinc-inducible manner, as previously described13. The efficiency of AML1-ETO knockdown and overexpression was evaluated by western blotting, as described in our previous study17. BASP1 mRNA expression and protein levels were up-regulated in SKNO-1-siA/E compared to SKNO-1 and were down-regulated in U937-A/E compared to U937 (Fig. 1e), suggesting that BASP1 expression may be selectively silenced by A/E in A/E-positive AML.
a Expression of BASP1 is down-regulated in A/E-positive cell lines (Kasumi-1 and SKNO-1) compared to the other A/E-negative AML cell lines (NB4, U937, THP-1, KG-1, MV-4-11, HEL, and HL-60) and three sets of normal bone marrow (NBM). b Expression of BASP1 in different subtypes of AML and hematopoietic stem cells (HSC) analyzed from the GEO dataset (GSE13159). BASP1 expression was down-regulated in t(8;21) AML in contrast to the other subtypes of AML and HSC (***p < 0.001). c The protein level of BASP1 was detected by western blotting in AML cell lines. β-actin was used as a control. d mRNA expression of BASP1 was plotted against that of AML1-ETO in A/E-positive AML (n = 60) (GSE13159). The plot of BASP1 versus AML1-ETO expression showed an inverse correlation (r = -0.38, p = 0.0027). e Upper panel: mRNA expression of BASP1 was measured in two pairs of cell lines: SKNO-1 and SKNO-1-siA/E, and U937 and U937-A/E. GAPDH was used as a control (**p < 0.01, ***p < 0.001). Lower panel: The protein levels of BASP1 were detected by western blotting. β-Actin was used as the control
BASP1 silencing in A/E-positive AML by promoter methylation
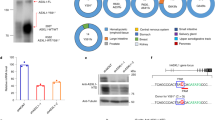
To further clarify the mechanism of BASP1 down-regulation in A/E-positive AML, we analyzed the genomic sequence of the BASP1 promoter region (−2000 to −1 bp) by performing a bioinformatics search (http://www.urogene.org/methprimer/) and identified one CpG island with CpGs that were densely clustered close to the transcription start site (Fig. 2a). Bisulfite sequencing of the genome region from −179 to −400 bp of the BASP1 promoter showed that this CpG island was hypermethylated in A/E-positive AML cell lines (Kasumi-1 and SKNO-1), but not in A/E-negative AML cell lines (HL-60 and SKNO-1-siA/E) and NBM (Fig. 2b). MSP showed that the BASP1 promoter was methylated in almost all primary t(8;21) AML samples (n = 9), but not in NBM (n = 3) (Fig. 2c). To elucidate whether DNA methylation was able to account for the regulation of BASP1 expression, we treated Kasumi-1 and SKNO-1 cells with a DNMT inhibitor, DAC, for 72 h. The mRNA expression and protein levels of BASP1 were significantly increased after treatment with DAC in a dose-dependent manner (Fig. 2d). These results indicated that DNA methylation may contribute to the transcriptional silencing of BASP1 in A/E-positive AML.
a Schematic diagrams of the CpG island of BASP1 promoter sequences (−2000 to −1 bp). The horizontal line (F1–R1) indicates the location of the primer used in the BSP assays. b Genomic bisulfate sequencing was performed to detect the methylation status of the DNA sequences in the CpG island of the BASP1 promoter from the indicated leukemia cell lines and normal bone marrow. Black dots and white dots indicate methylated and unmethylated CpG cytosine, respectively. c DNA methylation of BASP1 in nine A/E-positive AML cases (#) and three normal bone marrow samples (N) was demonstrated by MSP. M methylated allele, U unmethylated allele. d Upper panel: mRNA expression of BASP1 was up-regulated after continuous treatment with 0.5 or 1.0 µmol/L decitabine (DAC) for 3 days compared to the vehicle in Kasumi-1 and SKNO-1. GAPDH was used as a control. Error bars show the SD from three different assays (*p < 0.05, **p < 0.01). Lower panel: The protein levels of BASP1 were detected by western blotting. β-Actin was used as a control
A/E triggers epigenetic silencing of BASP1 through direct promoter binding
To address the question of whether the epigenetic silencing of BASP1 in t(8;21) AML is directly mediated by A/E, we searched the JASPAR database (http://jaspar.genereg.net/cgi-bin/jaspar_db.pl?rm=browse&db=core&tax_group=vertebrates) and found four putative AML1-binding sites in the BASP1 promoter region (−1500 to −1 bp). We cloned various lengths of the BASP1 promoter sequence with different wild-type A/E-binding sites into a pGL3-basic vector and monitored luciferase activity with co-transfection of the A/E plasmid into 293T cells. As shown in Fig. 3a, the promoter activity of BASP1 was significantly decreased after co-transfection with the A/E plasmid (10 ng) when the luciferase reporter contained sequences of a functional binding site (B-3), but not when the luciferase reporter contained no functional binding sites or a mutated sequence (B-3M). Furthermore, increasing the amount of the A/E plasmid to 50 and 100 ng did not further inhibit the luciferase activity of the BASP1 promoter (data not shown), suggesting that the presence of a low dose of A/E was sufficient to maximize inhibition of BASP1 promoter activity. These results suggest that AML1-binding site B-3 (nucleotides at −546 to −556) in the promoter region of BASP1 may be related to the A/E-dependent silencing of BASP1.
a The left diagram shows a luciferase reporter containing various lengths of the BASP1 promoter region with wild-type A/E-binding sites1,2,3,4 or mutated sequence (mut). Luciferase reporter assays showed that A/E repressed the transcriptional activity of the BASP1 promoter. b ChIP was performed in the SKNO-1 and SKNO-1-siA/E cell lines using anti-AML1, anti-ETO, anti-DNMT1, anti-DNMT3a, and anti-DNMT3b antibodies or a non-specific antibody (IgG control). Quantitative RT-PCR was performed with primers specific for the BASP1 promoter region surrounding the predicted −546/−556 binding site and expression was quantified as 2−ΔCT relative to input. ChIP-PCR showed that A/E and DNMT3a bound the BASP1 promoter (*p < 0.05, **p < 0.01)
To study whether A/E binds directly to the promoter region of BASP1 and leads to the methylation of the BASP1 promoter region, we performed ChIP-PCR assays in the SKNO-1 and SKNO-1-siA/E cell lines using anti-AML1, anti-ETO, anti-DNMT1, anti-DNMT3a, and anti-DNMT3b antibodies or a non-specific antibody (normal IgG). Quantitative RT-PCR was performed with primers specific to the BASP1 promoter region surrounding the predicted −546/−556 site or with the unrelated negative control primers. The results showed that A/E and DNMT3a were present at the BASP1 promoter region surrounding the functional AML1-binding site, but not for DNMT1 and DNMT3b (Fig. 3b and Supplementary Figure 1). This suggests that A/E and DNMT3a may form a co-repressor complex to inhibit the expression of BASP1 by binding directly to its promoter region and then methylating the DNA.
BASP1 plays a tumor suppressor role in A/E-positive cell lines in vitro
To evaluate the effects of BASP1 in t(8; 21) AML, we transduced SKNO-1 cells with the GFP-tagged BASP1 or GFP control lentiviral plasmid. Transfection was confirmed by flow cytometry (Supplementary Figure 2). BASP1 restoration significantly reduced cellular proliferation, as monitored by the CCK-8 assay to quantify the colony-forming ability and colony-forming assay in SKNO-1 cells (Fig. 4a, b). BASP1 restoration also blocked cells in the G1 phase of the cell cycle and reduced the number of cells in the S phase (Fig. 4c). In addition, BASP1 restoration increased the number of apoptotic SKNO-1 cells by 1.6-fold compared to the negative control (Fig. 4d). We also examined the biological effects of BASP1 on the expression of CD11b by using flow cytometry and found no significant differences between BASP1-restored cells and the control group (data not shown). These data indicate that BASP1 may play a key role in the regulation of cell proliferation by inducing apoptosis and cell cycle arrest in A/E-positive AML.
a The CCK-8 assay was performed on SKNO-1 cells transduced with the GFP-tagged BASP1 lentiviral or GFP control plasmids at the indicated time points. b A colony formation assay was performed on SKNO-1 cells transduced with the GFP-tagged BASP1 lentiviral or GFP control plasmids and grown for 7 days. c Cell cycle analyses in SKNO-1 cells by flow cytometry showed that BASP1 overexpression induced the G1 phase arrest and blocked S-phase entry. d The induction of apoptosis in BASP1 overexpressing cells was measured using Annexin V-7AAD staining via flow cytometry. Error bars indicates the SD from three different assays (*p < 0.05)
Discussion
As a transcription factor, the A/E fusion protein plays a critical role in the pathogenesis of t(8;21) AML and regulates gene expression through various mechanisms, that is, by direct DNA binding and recruitment of DNMTs, HDACs, P300, and PRMT1;12, 18,19,20 by interacting with other transcription factors;15, 21, 22 or by forming stable multiprotein complexes, including CBFβ/HEB/E2A/LYL1/AML123. Our previous studies showed that A/E can bind to the promoters of miR-9-1 and SIRT1 to form a regulatory circuit, resulting in the down-regulation of miR-9 and upregulation of SIRT1 and contributing to leukemogenesis15, 17. In this study, we showed for the first time that hypermethylation of the BASP1 gene promoter leads to specific down-regulation of BASP1 in t(8;21) AML. Our results showed that the A/E fusion protein could bind to the AML1-binding site on the BASP1 promoter sequence together with DNMT3a and inactivate transcription of BASP1 by CpG island hypermethylation. BASP1 may play a role as a tumor suppressor gene in vitro in t(8;21) AML.
BASP1 contains a nuclear localization sequence domain that can localize to promyelocytic leukemia bodies within the nucleus. Release of BASP1 from the nucleus to the cytoplasm was observed in apoptotic cells24. We also observed enrichment of BASP1 in the nucleus after transfection of the BASP1 lentiviral vector in 293T cells. Further, ectopic expression of BASP1 promoted apoptosis of A/E-positive leukemia cells. As a transcriptional co-regulator of Wilms’ tumor-1 (WT1) protein, BASP1 is involved in the regulation of many genes related to apoptosis pathways, such as BCL-2 family, c-Myc, p21, and others4, 6. A study in thyroid cancer cells showed that BASP1 overexpression increases the expression of bax, promote cytochrome C release, and activate caspase-3, which ultimately results in apoptosis25. Again, further work is needed to determine the exact mechanism of BASP1-induced apoptosis in t(8;21) AML cells. In addition, BASP1 has been reported to be involved in cell differentiation. After ectopic expression of BASP1 in K562 cells, more than 90% of cells were differentiated by PMA induction26. We found that BASP1 expression was gradually increased during the differentiation of myeloid cells using publicly available expression data (Supplementary Figure 3). However, we were unable to find any difference in the CD11b expression level after overexpression of BASP1 compared to controls in A/E-positive cells, but cell cycle arrest occurred. Therefore, we hypothesize that the expression level of BASP1 may be related to the degree of cell differentiation, but is probably not directly involved in the induction of cell differentiation in t(8;21) AML. Inhibition of proliferation in A/E-positive AML cells by ectopic expression of BASP1 may occur mainly through cell cycle arrest and induction of apoptosis.
Expression of WT1 might play an important role in A/E-mediated leukemogenesis27. Many studies using murine models have demonstrated that A/E alone was not able to independently induce leukemia28, 29. However, all mice rapidly developed AML after transplantation of A/E-transduced bone marrow cells co-expressing WT127. BASP1 has been identified as a transcriptional cosuppressor of WT1 and exerts its transcriptional activity by binding to the promoters of target genes together with WT1. Down-regulation of BASP1 weakens the inhibitory effect of WT1 on target genes causing the transcriptional activation domain to be released4, 30. BASP1 has also been identified as a specific MYC target that can inhibit v-Myc-induced cell transformation31. MYC is frequently activated in AML, and WT1 can also induce upregulation of MYC, which is correlated with the induction of leukemia, notably for sustained cell proliferation32. It will be important in the future to study the effect of BASP1 expression on signaling pathways regulated by WT1 and MYC to better understand the possible mechanisms of BASP1 promoter hypermethylation in the leukemogenicity of t(8;21) AML.
In conclusion, our results provide new insights into the role of BASP1 in leukemogenesis. We demonstrate that BASP1 plays a role as a tumor suppressor gene in vitro and is silenced via promoter methylation in t(8;21) AML. We suggest that manipulation of BASP1 expression may be a new strategy for the treatment of t(8; 21) AML.
References
Maekawa, S., Maekawa, M., Hattori, S. & Nakamura, S. Purification and molecular cloning of a novel acidic calmodulin binding protein from rat brain. J. Biol. Chem. 268, 13703–13709 (1993).
Bomze, H. M., Bulsara, K. R., Iskandar, B. J., Caroni, P. & Skene, J. H. Spinal axon regeneration evoked by replacing two growth cone proteins in adult neurons. Nat. Neurosci. 4, 38–43 (2001).
Korshunova, I. et al. Characterization of BASP1-mediated neurite outgrowth. J. Neurosci. Res. 86, 2201–2213 (2008).
Carpenter, B. et al. BASP1 is a transcriptional cosuppressor for the Wilms’ tumor suppressor protein WT1. Mol. Cell. Biol. 24, 537–549 (2004).
Sanchez-Nino, M. D. et al. BASP1 promotes apoptosis in diabetic nephropathy. J. Am. Soc. Nephrol. 21, 610–621 (2010).
Green, L. M., Wagner, K. J., Campbell, H. A., Addison, K. & Roberts, S. G. Dynamic interaction between WT1 and BASP1 in transcriptional regulation during differentiation. Nucleic Acids Res. 37, 431–440 (2009).
Moribe, T. et al. Identification of novel aberrant methylation of BASP1 and SRD5A2 for early diagnosis of hepatocellular carcinoma by genome-wide search. Int. J. Oncol. 33, 949–958 (2008).
Ransohoff, K. J. et al. Two-stage genome-wide association study identifies a novel susceptibility locus associated with melanoma. Oncotarget 8, 17586–17592 (2017).
Grimwade, D., Ivey, A. & Huntly, B. J. Molecular landscape of acute myeloid leukemia in younger adults and its clinical relevance. Blood 127, 29–41 (2016).
Licht, J. D. AML1 and the AML1-ETO fusion protein in the pathogenesis of t(8;21) AML. Oncogene 20, 5660–5679 (2001).
Peterson, L. F. & Zhang, D. E. The 8;21 translocation in leukemogenesis. Oncogene 23, 4255–4262 (2004).
Liu, S. et al. Interplay of RUNX1/MTG8 and DNA methyltransferase 1 in acute myeloid leukemia. Cancer Res. 65, 1277–1284 (2005).
Li, Y. et al. Epigenetic silencing of microRNA-193a contributes to leukemogenesis in t(8;21) acute myeloid leukemia by activating the PTEN/PI3K signal pathway. Blood 121, 499–509 (2013).
Figueroa, M. E. et al. DNA methylation signatures identify biologically distinct subtypes in acute myeloid leukemia. Cancer Cell 17, 13–27 (2010).
Zhou, L. et al. AML1-ETO promotes SIRT1 expression to enhance leukemogenesis of t(8;21) acute myeloid leukemia. Exp. Hematol. 46, 62–69 (2017).
Dou, L. et al. Methylation-mediated repression of microRNA-143 enhances MLL-AF4 oncogene expression. Oncogene 31, 507–517 (2012).
Fu, L. et al. A minicircuitry of microRNA-9-1 and RUNX1-RUNX1T1 contributes to leukemogenesis in t(8;21) acute myeloid leukemia. Int. J. Cancer 140, 653–661 (2017).
Saeed, S. et al. Chromatin accessibility, p300, and histone acetylation define PML-RARalpha and AML1-ETO binding sites in acute myeloid leukemia. Blood 120, 3058–3068 (2012).
Shia, W. J. et al. PRMT1 interacts with AML1-ETO to promote its transcriptional activation and progenitor cell proliferative potential. Blood 119, 4953–4962 (2012).
Wang, L. et al. The leukemogenicity of AML1-ETO is dependent on site-specific lysine acetylation. Science 333, 765–769 (2011).
Martens, J. H. et al. ERG and FLI1 binding sites demarcate targets for aberrant epigenetic regulation by AML1-ETO in acute myeloid leukemia. Blood 120, 4038–4048 (2012).
Gao, X. N. et al. AML1/ETO cooperates with HIF1alpha to promote leukemogenesis through DNMT3a transactivation. Leukemia 29, 1730–1740 (2015).
Sun, X. J. et al. A stable transcription factor complex nucleated by oligomeric AML1-ETO controls leukaemogenesis. Nature 500, 93–97 (2013).
Ohsawa, S., Watanabe, T., Katada, T., Nishina, H. & Miura, M. Novel antibody to human BASP1 labels apoptotic cells post-caspase activation. Biochem. Biophys. Res. Commun. 371, 639–643 (2008).
Guo, R. S. et al. Restoration of brain acid soluble protein 1 inhibits proliferation and migration of thyroid cancer cells. Chin. Med. J. (Engl.) 129, 1439–1446 (2016).
Goodfellow, S. J. et al. WT1 and its transcriptional cofactor BASP1 redirect the differentiation pathway of an established blood cell line. Biochem J. 435, 113–125 (2011).
Nishida, S. et al. AML1-ETO rapidly induces acute myeloblastic leukemia in cooperation with the Wilms tumor gene, WT1. Blood 107, 3303–3312 (2006).
Wang, Y. Y. et al. C-KIT mutation cooperates with full-length AML1-ETO to induce acute myeloid leukemia in mice. Proc. Natl. Acad. Sci. USA 108, 2450–2455 (2011).
Yuan, Y. et al. AML1-ETO expression is directly involved in the development of acute myeloid leukemia in the presence of additional mutations. Proc. Natl. Acad. Sci. USA 98, 10398–10403 (2001).
Toska, E., Shandilya, J., Goodfellow, S. J., Medler, K. F. & Roberts, S. G. Prohibitin is required for transcriptional repression by the WT1-BASP1 complex. Oncogene 33, 5100–5108 (2014).
Hartl, M., Nist, A., Khan, M. I., Valovka, T. & Bister, K. Inhibition of Myc-induced cell transformation by brain acid-soluble protein 1 (BASP1). Proc. Natl. Acad. Sci. USA 106, 5604–5609 (2009).
Delgado, M. D. & Leon, J. Myc roles in hematopoiesis and leukemia. Genes Cancer 1, 605–616 (2010).
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (81700133, 81370635, 81570137, 81470010, 81400135, 81500118, and 81670162), the Capital Medical Development Scientific Research Fund (SF2001-5001-07), Beijing Natural Science Foundation (7151009), the National Public Health Grant Research Foundation (No. 201202017), The capital of the public health project (Z111107067311070) and The China Postdoctoral Science Foundation funded project (project no. 2016M600443). The English in this document was checked by at least two professional editors, both of whom are native English speakers. For a certificate, please see: http://www.textcheck.com/certificate/mW3grs.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. If you remix, transform, or build upon this article or a part thereof, you must distribute your contributions under the same license as the original. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/.
About this article
Cite this article
Zhou, L., Fu, L., Lv, N. et al. Methylation-associated silencing of BASP1 contributes to leukemogenesis in t(8;21) acute myeloid leukemia. Exp Mol Med 50, 1–8 (2018). https://doi.org/10.1038/s12276-018-0067-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s12276-018-0067-4