Abstract
Glutaryl-CoA dehydrogenase (GCDH) deficiency causes glutaric aciduria type I (GA I), an inborn error of metabolism that is characterized clinically by dystonia and dyskinesia and pathologically by neural degeneration of the caudate and putamen. Studies of metabolite excretion allowed us to categorize 43 GA I Spanish patients into two groups: group 1 (26 patients), those presenting with high excretion of both glutarate and 3-hydroxyglutarate, and group 2 (17 patients) , those who might not be detected by routine urine organic acid analysis because glutarate might be normal and 3-hydroxyglutarate only slightly higher than controls. Single-strand conformation polymorphism (SSCP) screening and sequence analysis of the 11 exons and the corresponding intron boundaries of the GCDH gene allowed us to identify 13 novel and 10 previously described mutations. The most frequent mutations in group 1 were A293T and R402W with an allele frequency of 30% and 28%, respectively. These two mutations were also found in group 2, but always in heterozygosity, in particular in combination with mutations V400M or R227P. Interestingly, mutations V400M and R227P were only found in group 2, and at least one of these mutations was found in 11 of 15 unrelated alleles, accounting together for 53% of the mutant alleles in group 2. Therefore, it seems clear that two genetically and biochemically distinct groups of patients exist. The severity of the clinical phenotype seems to be closely linked to the development of encephalopathic crises rather than to residual enzyme activity or genotype. Comparison of GCDH protein with other acyl-CoA dehydrogenases (whose x-ray crystal structure has been determined) reveals that most of the mutations identified in GCDH protein seem to affect folding and tetramerization, as has been described for a number of mutations affecting mitochondrial β-oxidation acyl-CoA dehydrogenases.
Similar content being viewed by others
Main
GA I (MIM 231670) is an autosomal recessive disorder caused by deficient or nonfunctional GCDH (EC 1.3.99.7). The metabolic block leads to the accumulation of glutaric and 3-hydroxyglutaric acids as well as glutarylcarnitine in body fluids. The clinical picture is characterized by the sudden onset of a severe dystonic-dyskinetic disorder, hypotonia, irritability, macrocephaly, and degeneration of the caudate and putamen, which generally appear between 5 and 14 mo of age, but mild symptoms such as motor delay and hypotonia can be observed at earlier ages (1–4). However, clinical variability is common and some patients remain asymptomatic (2, 4). The condition is not exceedingly rare, and a frequency of 1:40,000 in Caucasians has been proposed (2).
The GCDH enzyme functions in the mitochondrial matrix as a homotetramer. The monomer is made up of 438 amino acids (48.2 kD), of which the 44 N-terminal residues are cleaved after being imported into the mitochondria. It also contains one molecule of FAD, which is reduced by ETF (5).
The human GCDH gene has been mapped to chromosome 19p13.2 (6), spans about 7 Kb, and comprises 11 exons and 10 introns. The genomic structure and most of the intronic sequences have been published (5, 7, 8) and the amino acid sequences of mouse, human, and pig GCDH are highly conserved (9). Single prevalent mutations, IVS1+5G>T and A421V, have been found in the Ojibway Indians of Canada (10) and in the Amish of Pennsylvania (7), respectively. However, mutations of the GCDH gene in general population are heterogeneous. To date, 66 disease-causing mutations and 6 polymorphisms and neutral mutations have been identified in the GCDH gene (7, 8, 10–16), the most common mutation being R402W with an allele frequency of 12 to 16% (7, 8).
In this study, we have analyzed metabolite excretion, GCDH activity, and the mutations responsible for GCDH deficiency in 43 Spanish patients with GA I. Our studies showed that there were two genetically and biochemically distinct groups of patients. We have attempted to establish a correlation between the GA I genotype and the clinical or biochemical phenotype.
MATERIALS AND METHODS
Patients.
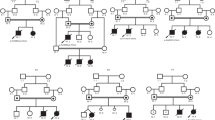
We studied 43 patients (35 unrelated families) with GA I from different areas of Spain. Diagnosis was established by clinical symptoms, characteristic findings on neuroimaging, and quantitative analysis of organic acids in urine. The disease was confirmed by analysis of GCDH activity in cultured skin fibroblasts, as previously described (17). Fibroblasts from nine patients were not available, but in six of them GCDH activity could be predicted on the basis of other patients belonging to the same GCDH genotype. Only in four families were the parents known to be consanguineous. Case histories of patients 4, 13, 19, 22, 23, 25, 26, 27, 28, 34, 38, 39, and 42 have been previously described (18–23).
In 24 patients the disease began abruptly, during infancy, with focal seizures or generalized convulsions, vomiting, and obtundation or lethargy, usually after an acute infectious illness. In 18 patients the onset was “insidious,” with slowly developing psychomotor delay, hypotonia as dystonic postures, lethargy, or macrocephaly. The phenotype was classified according to the following criteria: “severe,” when the patient showed dystonia (either static or progressive) and was unable to stand up, walk, or speak (25 patients); “mild,” when dystonia was static or improving and the child was able to walk without help (11 patients); and “slight,” when there were only minimal neurologic signs, and the child was able to lead a normal life (5 patients). Two patients were asymptomatic. The siblings of patients presented the same phenotype and clinical onset as their corresponding index case, except patient 11, who is an elder asymptomatic brother of patient 10, and patient 17, who is a younger brother of patient 16 with a milder clinical phenotype (Table 1).
The ethics committees of the participating hospitals and organizations approved the collection of samples on which these studies were performed.
Quantitative analysis of glutarate and 3-hydroxyglutarate.
Organic acids were isolated by strong anion exchange (SAX) column extraction as described (24) and were analyzed by GC-MS (Hewlett-Packard 5890, Palo Alto, CA, U.S.A.) as trimethylsilyl (TMS)-derivatives. For quantitation, the mass spectrometry was operated in the single ion monitoring mode, and the following ion currents were measured: glutarate m/z 261, 3-hydroxyglutarate m/z 259, and undecanedioate (internal standard) m/z 345. The sodium salts of glutaric and undecanedioc acids were obtained from Sigma Chemical Co. (St. Louis, MO, U.S.A.) and 3-hydroxyglutaric acid was from Dr. Brunet (Organic Chemistry Department, Universidad Autónoma de Madrid, Spain).
Mutation analysis.
Genomic DNA was prepared from fresh blood or from cultured skin fibroblasts by standard methods. Total RNA was prepared from cultured fibroblasts by the UltraspecTM RNA Isolation System (Biotecx, Laboratories, Houston, TX, U.S.A.).
Mutation screening of genomic DNA was performed by PCR amplification and SSCP analysis. PCR conditions and primers used were as described elsewhere (7, 14).
SSCP was performed at room temperature or at 4°C. DNA fragments were detected after silver staining. The bands with mobility shifts were excised from the gel, purified (QIAquick Gel Extraction Kit; QIAGEN, GmbH Germany, Hilden, Germany) and reamplified, followed by sequencing using a Sequenase Version 2.0 Kit (USB, Amersham Pharmacia Biotech, Buckinghamshire, U.K.) and 35S-dATP. In some cases, PCR products were purified (QIAquick PCR Purification Kit; QIAGEN) and sequenced with the DNA sequencing kit, dye terminator cycle sequencing ready reaction (Perkin Elmer Applied Biosystems Division, Foster City, CA, U.S.A.).
For cDNA analysis, about 5 μg of total RNA was transcribed into the first strand of cDNA in a 25-μL reaction volume containing avian myeloblastosis virus (AMV) 1× buffer, 400 μM dinucleotide triphosphate (dNTP), 10 U AMV reverse transcriptase (Promega, Madison, WI, U.S.A.), 33 U RNA in ribonuclease inhibitor (Promega), and 400 nM of primer for 45 min at 52°C. PCR amplification of cDNA was carried out using primers described elsewhere (10). PCR products were studied by SSCP analysis and sequenced as described above.
When possible, sequence changes were confirmed by restriction enzyme analysis as summarized in Table 2. To detect mutation A293T, a mismatch PCR was designed. We introduced an Hga I restriction site into the mutant PCR product. The two primers used were located on exon 8. The forward mismatch primer was F5′-CCTTCGGCTGCCTGAAC G AC-3′, where the mismatch nucleotide is underlined, and the reverse primer was as described (7).
Mutations were named according to suggested nomenclature (25). Throughout this article, all mutations except frameshift are referred to by their amino acid-based name.
RESULTS
Biochemical and clinical findings.
The use of quantitative, as opposed to qualitative, organic acid analysis allowed us to categorize the patients into two groups (Table 1): Group 1 (26 patients) comprised those who could be diagnosed simply by routine GC-MS organic acid analysis in urine. Patients presented with high excretion of both glutarate and 3-hydroxyglutarate, although the excretion of glutarate was always higher than that of 3-hydroxyglutarate. Group 2 (17 patients) comprised those who might go undetected by routine screening because glutarate excretion can be normal and that of 3-hydroxyglutarate might be only slightly increased. In contrast to group 1 patients, those in group 2 presented with a higher 3-hydroxyglutarate excretion than that of glutarate. The median and range (mmol/mol creatinine) of glutarate and 3-hydroxyglutarate excretion in 42 control urine samples were 3 (2–10) and 7 (2–14), respectively; values for group 1 patients were 2817 (204–8955) and 420 (75–3573), respectively. In contrast, group 2 showed a median value for glutarate, 10 (2–84), which is within the range of controls, whereas the median value for 3-hydroxyglutarate, 101 (16–571), fell within the range of group 1. When possible, only those values of glutarate and 3-hydroxyglutarate obtained at the time of diagnosis were considered to obviate the possible influence of any treatment.
Glutarylcarnitine in blood spots was tested in six patients from group 2: patient 38 (Dr. Millington, Duke University, Durham, NC, U.S.A.) and patients 30, 34, 36, 37, and 41 (Dr. Hoffmann, Philipps-Universitat, Marburg, Germany). All of them gave negative results (even under carnitine supplementation). This indicates that patients with this metabolite excretion pattern would not be detected in neonatal screening programs employing tandem mass spectrometry.
In group 1, all but two patients had undetectable GCDH activity in fibroblasts (<0.5% of controls), whereas in group 2, the residual activities were usually higher (Table 1).
Acute encephalopathy marked the clinical onset in 24 patients, of whom 21 presented the severe phenotype. Insidious onset was found in 18 patients, of whom only 4 presented the severe phenotype (Table 1). Interestingly, 71% of group 2 patients presented the severe phenotype, whereas it was only found in 50% of group 1 patients (Table 1). Therefore, the severity of the clinical phenotype does not seem to be directly related to the severity of the biochemical phenotype.
Mutation analysis.
SSCP screening of the 11 exons and intron boundaries of the GCDH gene allowed us to identify 13 novel (10 missense, 2 deletions, and 1 nonsense) and 10 already known mutations, which account for 99% (69/70) of the total number of mutant alleles (Table 2). Sequence analysis of GCDH cDNA and genomic DNA of patient 33 revealed that mutation V400M was the only consistent change in the entire protein-coding region, suggesting that the other mutations lie in the promoter region or elsewhere within the intronic sequences. Of the 23 mutations identified only three were common to both groups (A293T, R402W, and 1209delG). However, these mutations were found in group 2 always in heterozygosity, in particular in combination with mutations V400M or R227P. The latter mutations accounted for 53% of the group 2 alleles and were exclusive to this group, whereas R402W and A293T mutations, accounted for 58% of the group 1 alleles (Table 2). None of the missense mutations was found in 100 unrelated control chromosomes. Mendelian inheritance of the mutations was confirmed by use of a restriction-digestion test, where available, or by SSCP pattern, in the DNA of the parents of 25 families.
Flow chart for the diagnosis of GA I.
The flowchart depicted in Figure 1, based on the biochemical and mutational results reported here, is now being used in our laboratory. It is especially useful for the diagnosis of group 2 patients, and in most cases means that the measurement of GCDH activity is not necessary. Patients 29, 33, and 36 were, in fact, diagnosed using this flowchart.
DISCUSSION
The aim of this study was to identify the mutations responsible for GCDH deficiency in the Spanish patients to investigate a possible correlation between GA I genotypes and clinical and biochemical phenotypes.
Mutational spectrum.
We report 13 novel mutations in the GCDH gene. This means that, at present, a total of 79 potentially pathogenic mutations are known. Missense mutations are more frequently detected than nonsense mutations or deletions, and here 55% of nucleotide substitutions were found at hypermutable CpG sites in the GCDH gene (Table 2). Deletions and nonsense mutations all result in truncation of the mutant protein before the α-proton-abstracting base (E414) in enzyme catalysis (5); consequently, the encoded protein would not be expected to have any residual activity. Several lines of evidence suggest that missense mutations may be pathogenic: (1) after screening all of the coding region of the GCDH gene, no other mutations were found in the patients; (2) the nucleotide changes were not present in 100 normal chromosomes analyzed; and (3) all the amino acid residues, except R161 and A316, are conserved in pig and mouse GCDH homologs (5, 9).
According to the pattern of metabolite excretion, patients were categorized into two groups. In group 1, we have found 15 different mutations, of which 9 are novel. In contrast to other reports (7, 8, 15), we found a particular prevalence not only for mutation R402W (28%) but also for mutation A293T (30%). In addition, we have recently found linkage disequilibrium between mutation R402W and a specific haplotype, suggesting a single origin for this mutation (26). In group 2, we have found 11 mutations, of which 4 are novel. The most striking result from this group was the strong association with mutations V400M and R227P, which together account for more than 50% of mutant alleles. We have previously shown that the R227P allele was associated with minimal glutarate excretion when heterozygous (13) and it has recently been confirmed in a patient with genotype E365K/R227P, whereas patients homozygous for mutation E365K present with large amounts of glutarate and 3-hydroxyglutarate in urine (27). The same applies when mutation A293T was found in homozygosity, whereas in heterozygosity with R227P the biochemical phenotype changed to that of group 2 (Table 1). Consequently, R227P seems to be one of the causative mutations of this phenomena, but it is not clear how this mutation acts (7, 8). It might be the case that in these patients the decarboxylase activity was more severely affected than that of the dehydrogenase, but this was not sustained when mutation R227P was expressed in Escherichia coli, and partial reactions (dehydrogenation and decarboxylation) of GCDH activity were measured (28). A similar phenomena has been observed with mutation V400M. Genotypes V400M/R227P (patients 34, 35, and 36) and V400M/unknown (patient 33) presented the highest residual enzyme activities in fibroblasts, similar to those in the heterozygous range. If we take into account that patients homozygous for mutation V400M have lower enzyme activities than those compound heterozygous for V400M/R227P, a complementary effect between these two alleles might be speculated. Other private mutations (F71S, R161Q, and S255L) were also found to be associated with this group of patients.
Effects of GCDH mutations on protein structure.
Human GCDH displays a high degree of homology with other acyl-CoA dehydrogenases (Fig. 2), therefore, comparison with the x-ray structure of MCAD (29), IVD (30) and SCAD or butyryl-CoA dehydrogenase (BCAD) (31) may help to explain the molecular consequences of the identified mutations. As described for MCAD (29), the monomer structure of GCDH can be divided in three regions with a distinct secondary structure. Region A, from R-45 to E-172 (Fig. 2), consists of a bundle of six α-helices that does not appear to be directly involved in either substrate binding or oligomerization (except for the first five residues, which might participate in oligomerization). In this region we found six mutations (F71S, E90K, S119L, Y155H, R132Q, and R161Q), most of which are associated with residual activity. Two of these, F71S and S119L, may affect protein stability because of a change in polarity, which is, moreover, maintained among reference structures. In particular, the change in polarity brought about by F71S could prevent the salt bridge formation between residues R72 and E129, impairing GCDH folding, as has been described for mutations R28C and R22W in MCAD and SCAD proteins, respectively (32, 33). Residue Y-155 forms a hydrogen bond with G-284, which lies near the adenine ring of CoA. Therefore, mutation Y155H might compromise substrate binding. The consequences for mutations E90K, R132Q, and R161Q are less evident.
Alignment of the human GCDH sequence with the sequences of three acyl-CoA dehydrogenases (ACD) with known three-dimensional structure: pig MCAD (29), human IVD (30), and bacterial SCAD or BCAD (31). The GCDH sequence was aligned manually with the reference sequences taking into account conserved residues, residues in contact with substrates, and also the suitability of hydrophobicity and secondary structure profiles. All numbering is according to the precursor enzymes. The names of α-helices and β-sheets are given below their sequence. The position of the amino acid change due to each missense mutation identified is shown above its sequence with downward-pointing arrows. Residues shared by all ACD or residues that are identical in GCDH and MCAD are highlighted. Positions marked by a number are described in detail in the text: 1, E-129 in GCDH; 2, G-284 in GCDH; 3, K-164 in mature MCAD (K189 in precursor MCAD); 4, E-408 in GCDH.
Region B, from L-173 to P-278 (Fig. 2), is a β-sandwich in close proximity to FAD. In this region we found four mutations, two of them (E181K and A195T) were associated with group 1 patients, whereas the other two (R227P and S255L) were found in group 2 patients. The substitution E181K may alter the local fold around the riboflavine moiety of FAD. Moreover, residue E-181 is homologous to residue E-137 in MCAD, the carboxyl of which is crucial for β-domain folding, and which has been described to affect tetramerization (34). A homologue disease-causing mutation, E178K, in VLCAD has also been identified (35). A similar effect could be predicted for mutation S255L. The mutation A195T leads to an increase in the volume and polarity of a conserved residue buried inside the β-sandwich, and this might alter the stability of the whole region. Mutation R227P lies in a region that might be involved in the ETF interaction site, as suggested for MCAD protein (36).
Region C consists of a bundle of four antiparallel α-helices, the latter split in two halves. Loops connecting the helices are directly involved in the interaction with FAD molecule, the loop between α7 and α8 is in contact with the adenine ring, and the loop between α9 and α10 (including a patch of conserved glycine residues) is in contact with the pyrophospate group of the dinucleotide (Fig. 3). Moreover, FAD molecules interact at the same time with residues from the two subunits in such a way that FAD binding and subunit aggregation are closely linked. This region is structurally the most important, as regions C of the four subunits form the core of the tetramer. Mutations in this region are expected to compromise both FAD binding and intersubunit contacts. Figure 3 shows the position of the mutations found in this region (A293T, A304T, L309W, A316D, R372K, R383C, V400M, and R402W), together with a ribbon representation of regions C of two subunits. All of these mutations, except V400M, were associated with a lack of residual enzyme activity. The residual activity found in patients harboring mutation V400M can probably be accounted for by the fact that this does not constitute a drastic change. Three of these mutations (A293T, R402W, and V400M) were expressed in E. coli (15) and the results for R402W and V400M were inconsistent with the biochemical phenotype of our patients. The R402W mutant has been shown to have some residual activity (3%), whereas the residual activity for mutant V400M is almost undetectable (0.8%); the contrary has been observed when GCDH was assayed in fibroblasts. These results reflect the difference between in vivo and in vitro analyses, as also observed in other diseases (37).
Location of GCDH mutations found in region C. Ribbon representation of region C, comprising α7 to α10–11 (from 282 to 438 residues), of two subunits of the GCDH model. The view is oriented along the binary symmetry axis that relates the two subunits. Side chains of the mutated residues are shown as space-fill models and the two FAD molecules are indicated as ball and stick representations. Loops between α7 and α8 (adenine ring) and α9 and α10 (pyrophospate) interact directly with the coenzyme. All calculations were performed in a Silicon Graphics R8000 workstation using the molecular modeling programs Insight II, Homology, and Discover (Molecular Simulations, Inc., Princeton, NJ, U.S.A.).
Genotype-phenotype correlation.
Considerable clinical variability in patients with GA I has been reported (2–4). Patient 10 is an extreme example of this heterogeneity. This child presented a severe phenotype and died at 2 y of age, whereas her elder GA I brother is asymptomatic at 4 y of age (data not shown). Another apparent discrepancy of this disease is that a severe phenotype was recorded with greater frequency among group 2 patients (71%), the biochemical phenotype of whom is less severe than that of group 1. Therefore, it seems clear that high residual activity and low metabolite excretion do not prevent the development of a severe clinical phenotype (see patients 33 and 36). The severity of the phenotype seems to be closely linked to the development of encephalopathic crises, often in association with high fever. In our series, 91% of patients with encephalopathic onset presented a severe phenotype, and prevention of the encephalopathic crises is the only treatment available to avoid further deterioration. Thus, environmental factors, such us exposure to metabolic stress, are probably one of the main determinants of the clinical phenotype, irrespective of the GA I genotype. This is by no means a new finding in the history of acyl-CoA dehydrogenase deficiencies; in MCAD deficiency, patients with the most severe mutations may remain asymptomatic for years and patients with the mildest MCAD mutations may present with the most severe symptoms (38). Similar observations have been made in SCAD deficiency (39).
It is interesting to note that, in contrast to glutarate, the concentration of 3-hydroxyglutarate in urine is similar in both groups of patients. This compound, at concentrations much lower than glutarate, has been reported to induce massive nerve cell lesions in vitro (40, 41). Most probably, 3-hydroxyglutarate arises from β-oxidation of the intramitochondrial increased glutaryl-CoA through the β-oxidation enzyme, MCAD (1), but this enzyme is not expressed in brain (42) and, consequently, is unable to produce 3-hydroxyglutarate in this tissue. It is thus difficult to link brain damage to this metabolite, but it might be the case that 3-hydroxyglutarate crosses the blood-brain barrier due to increased permeability during viral or bacterial infections (43–45) and perhaps then provokes brain lesions and encephalopathic crisis. The fact that 3-hydroxyglutarate did not increase in cerebrospinal fluid during clinical stability (A.R., personal observation) and that other authors found it increased during an acute illness (46) favor this hypothesis.
From this study we can conclude the following: (1) For certain diseases it is necessary to perform mutation analysis in every population. In earlier reports (7, 8, 15) many private mutations were reported because, it would seem, the studies were conducted in heterogeneous populations. (2) Recently, it has been speculated that the number of group 2 patients was probably small (3), but this is not the case in our population, where 40% of patients were found to belong to this group. Whether this is a particular genetic characteristic of our population or reflects the results of ongoing attempts to improve diagnosis (18, 22, 23, 47) remains to be established. (3) The knowledge of GA I genotypes might be useful for prenatal diagnosis, particularly for families with high residual enzyme activity. (4) Comparison of GCDH protein with other crystal-solved acyl-CoA dehydrogenases reveals that most of the mutations identified might affect folding and tetramerization, as has been described for a number of mutations affecting mitochondrial β-oxidation acyl-CoA dehydrogenases. However, further experimental studies are needed to demonstrate this definitively. (5) With a few exceptions, a marked correlation between genotype and biochemical phenotype was found. However, no correlation was found between clinical symptoms, enzyme activities, and metabolite excretion in these Spanish GA I patients, which suggests that other factors contribute to the onset and severity of the disease.
Abbreviations
- GA I:
-
glutaric aciduria type I
- GCDH:
-
glutaryl-CoA dehydrogenase
- FAD:
-
flavin adenine dinucleotide
- ETF:
-
electron-transfer flavoprotein
- GC-MS:
-
gas chromatography-mass spectrometry
- MCAD:
-
medium-chain acyl-CoA dehydrogenase
- SCAD:
-
short-chain acyl-CoA dehydrogenase
- VLCAD:
-
very-long-chain acyl-CoA dehydrogenase
- IVD:
-
isovaleryl-CoA dehydrogenase
- SSCP:
-
single-strand conformation polymorphism
References
Goodman SI, Frerman FE 1995 Organic acidemias due to defects in lysine oxidation: 2-ketoadipic acidemia and glutaric acidemia. In: Scriver CR, Beaudet AL, Sly WS, Valle D (eds) The Metabolic and Molecular Bases of Inherited Disease, 7th Ed. McGraw-Hill, New York, 1451–1460.
Superti-Furga A, Hoffman G 1997 Glutaric aciduria type I (glutaryl-CoA dehydrogenase deficiency): advances and unanswered questions. Eur J Pediatr 156: 821–828.
Baric I, Zschocke J, Christensen E, Duran M, Goodman SI, Leonard JV, Müller E, Morton DH, Superti-Furga A, Hoffmann GF 1998 Diagnosis and management of glutaric aciduria type I. J Inherit Metab Dis 21: 326–340.
Hoffmann GF, Zschocke J 1999 Glutaric aciduria type I: from clinical, biochemical and molecular diversity to successful therapy. J Inherit Metab Dis 22: 381–391.
Goodman SI, Kratz LE, Di Giulio KA, Biery BJ, Goodman KE, Isaya G, Frerman FE 1995 Cloning of glutaryl-CoA dehydrogenase cDNA, and expression of wild type and mutant enzymes in Escherichia coli. Hum Mol Genet 4: 1493–1498.
Greenberg CR, Duncan AMV, Gregory CA, Singal R, Goodman SI 1994 Assignment of human glutaryl-CoA dehydrogenase gene (GCDH) to the short arm of chromosome 19 (19p13. Genomics 21: 289–290.
Biery BJ, Stein DE, Morton DH, Goodman SI 1996 Gene structure and mutations of glutaryl-CoA dehydrogenase: impaired association of enzyme subunits that is due to an A421V substitution causes glutaric acidemia type I in the Amish. Am J Hum Genet 59: 1006–1011.
Schwartz M, Christensen E, Superti-Furga A, Brandt NJ 1998 The human glutaryl-CoA dehydrogenase gene: report of intronic sequence and of 13 novel mutations causing glutaric aciduria type I. Hum Genet 102: 452–458.
Koeller DM, DiGiulio KA, Angeloni SV, Dowler LL, Frerman FE, White RA, Goodman SI 1995 Cloning, structure, and chromosome localization of the mouse glutaryl-CoA dehydrogenase gene. Genomics 28: 508–512.
Greenberg CR, Reimer D, Singal R, Triggs-Raine B, Chudley AE, Dilling LA, Philipps S, Haworth JC, Seargeant LE, Goodman SI 1995 A G-to-T transversion at the +5 position of intron 1 in the glutaryl-CoA dehydrogenase gene is associated with the Island Lake variant of glutaric acidemia type I. Hum Mol Genet 4: 493–495.
Haworth JC, Singal R, Goodman SI, Greenberg CR 1994 Taq I polymorphism in intron 2 of the GCDH gene. Hum Mol Genet 3: 678
Anikster Y, Shaag A, Joseph A, Mandel H, Ben-Zeev B, Christensen E, Elpeleg ON 1996 Glutaric aciduria type I in the Arab and Jewish communities in Israel. Am J Hum Genet 59: 1012–1018.
Christensen E, Ribes A, Busquets C, Pineda M, Duran M, Poll-The BT, Greenberg CR, Leffers H, Schwartz M 1997 Compound heterozygosity in the glutaryl-CoA dehydrogenase gene with R227P mutation in one allele is associated with no or very low free glutarate excretion. J Inherit Metab Dis 20: 383–386.
Busquets C, Coll MJ, Christensen E, Campistol J, Clusellas N, Vilaseca MA, Ribes A 1998 Feasibility of molecular prenatal diagnosis of glutaric aciduria type I in chorionic villi. J Inherit Metab Dis 21: 243–246.
Goodman SI, Stein DE, Schlesinger S, Christensen E, Schwartz M, Greenberg CR, Elpeleg ON 1998 Glutaryl-CoA dehydrogenase mutations in glutaric acidemia (type I): review and report of thirty novel mutations. Hum Mutat 12: 141–144.
Ikeda H, Kimura T, Ikegami T, Kato M, Matsunaga A, Yokoyama S, Yamaguchi S, Ohura T, Hayasaka K 1998 Novel mutations of the glutaryl-CoA dehydrogenase gene in two Japanese patients with glutaric aciduria type I. Am J Med Genet 80: 327–329.
Christensen E 1983 Improved assay of glutaryl-CoA dehydrogenase in cultured cells and liver: application to glutaric aciduria type I. Clin Chim Acta 129: 91–97.
Campistol J, Ribes A, Alvarez L, Christensen E, Millington DS 1992 Glutaric aciduria type I: unusual biochemical presentation. J Pediatr 121: 83–86.
Prats JM, Ribes A, Briones MP, Garaizar C, Sanjurjo P 1993 Aciduria glutarica tipo I. An Esp Pediatr 38: 343–347.
Martínez-Lage JF, Casas C, Fernandez MA, Puche A, Rodríguez-Costa T, Poza M 1994 Macrocephaly, dystonia, and bilateral temporal arachnoid cysts: glutaric aciduria type I. Childs Nerv Syst 10: 198–203.
Artigas J, Ribes A, Rovira A, Lorente I, Briones MP 1995 Aciduria glutárica tipo I con quistes aracnoideos. Rev Neurol 23: 153–156.
Merinero B, Pérez-Cerdá C, Font LM, Garcia MJ, Aparicio M, Lorenzo G, Martinez Pardo M, Garzo C, Martínez-Bermejo A, Pascual Castroviejo I, Christensen E, Ugarte M 1995 Variable clinical and biochemical presentation of seven Spanish cases with glutaryl-CoA dehydrogenase deficiency. Neuropediatrics 26: 238–242.
Pineda M, Ribes A, Busquets C, Vilaseca MA, Aracil A, Christensen E 1998 Glutaric aciduria type I with high residual glutaryl-CoA dehydrogenase activity. Dev Med Child Neurol 40: 840–842.
Verhaeghe BJ, Lefevere MF, De Leenheer AP 1988 Solid-phase extraction with strong anion-exchange columns for selective isolation and concentration of urinary organic acids. Clin Chem 3416: 1077–1083.
Antonorakis SE 1998 Recommendations for a nomenclature system for human gene mutations. Hum Mutat 11: 1–3.
Busquets C, Coll MJ, Ribes A 1999 Evidence of a single origin for the most frequent mutation R402W causing glutaryl-CoA dehydrogenase deficiency. Identification of 3 novel polymorphisms and haplotype definition. Hum Mutat, Mutation in Brief #291 on-line.
Nyhan WL, Zschocke J, Hoffmann G, Stein DE, Bao L, Goodman SI 1999 Glutaryl-CoA dehydrogenase deficiency presenting as 3-hydroxyglutaric aciduria. Mol Genet Metab 66: 199–204.
Liesert M, Zschocke J, Hoffmann GF, Mühlhäuser N, Buckel W 1999 Biochemistry of glutaric aciduria type I: Activities of in vitro expressed wild-type and mutant cDNA encoding human glutaryl-CoA dehydrogenase. J Inherit Metab Dis 22: 256–258.
Kim J-JP, Wang M, Paschke R 1993 Crystal structures of medium-chain acyl-CoA dehydrogenase from pig liver mitochondria with and without substrate. Proc Natl Acad Sci U S A 90: 7523–7527.
Tiffany KA, Roberts DL, Wang M, Paschke Mohsen A-WA, Vockley Kim J-JP 1997 Structure of human isovaleryl-CoA dehydrogenase at 2. Biochemistry 36: 8455–8464.
Djordjevic S, Pace CP, Stankovich MT, Kim J-JP 1995 Three-dimensional structure of butyryl-CoA dehydrogenase from Megasphaera elsdenii. Biochemistry 34: 2163–2171.
Bross P, Jespersen C, Jensen TG, Andresen BS, Kristensen MJ, Winter V, Nandy A, Kräutle F, Ghisla S, Bolund L, Kim J-JP, Gregersen N 1995 Effects of two mutations detected in medium chain acyl-CoA dehydrogenase (MCAD)-deficient patients on folding, oligomer assembly, and stability of MCAD enzyme. J Biol Chem 17: 10284–10290.
Naito E, Indo Y, Tanaka K 1990 Identification of two variant short chain acyl-CoA dehydrogenase alleles, each containing a different point mutation in a patient with short chain acyl-CoA dehydrogenase deficiency. J Clin Invest 85: 1575–1582.
Saijo T, Kim J-JP, Kuroda Y, Tanaka K 1998 The roles of threonine-136 and glutamate-137 of human medium chain acyl-CoA dehydrogenase in FAD binding and peptide folding using site-directed mutagenesis: creation of an FAD-dependent mutant, T136. Arch Biochem Biophys 358: 49–57.
Andresen BS, Olpin S, Poorthuis BJHM, Scholte JR, Vianey-Saban C, Wanders R, Ijlst L, Morris A, Pourfarzam M, Bartlett K, Baumgartner ER, deKlerk JBC, Schroeder LD, Corydon TJ, Lund H, Winter V, Bross P, Bolund L, Gregersen N 1999 Clear correlation of genotype with disease phenotype in very-long-chain acyl-CoA dehydrogenase deficiency. Am J Hum Genet 64: 479–494.
Roberts DL, Frerman FE, Kim J-JP 1996 Three-dimensional structure of human electron transfer flavoprotein to 2. Proc Natl Acad Sci U S A 93: 14355–14260.
Waters PJ, Parniak MA, Nowacki PN, Scriver CR 1998 In vitro expression analysis of mutations in phenylalanine hydroxylase: linking genotype to phenotype and structure to function. Hum Mutat 11: 4–7.
Andresen BS, Bross P, Udvari S, Kirk J, Gray G, Kmoch S, Chamoles N, Knudsen I, Winter V, Wilken B, Yokota I, Hart K, Packman S, Harpey JP, Saudubray JM, Hale DE, Bolund L, Kolvraa Gregersen N 1997 The molecular basis of medium-chain acyl-CoA dehydrogenase (MCAD) deficiency in compound heterozygous patients: is there correlation between genotype and phenotype. Hum Mol Genet 6: 695–707.
Gregersen N, Winter VS, Corydon MJ, Corydon TJ, Rinaldo P, Ribes A, Martinez G, Bennett MJ, Vianey-Saban C, Bhala A, Hale DE, Lehnert W, Kmoch S, Roig M, Riudor E, Eiberg H, Andresen BS, Bross P, Bolund LA, Kolvraa S 1998 Identification of four new mutations in the short-chain acyl-CoA dehydrogenase (SCAD) gene in two patients: one of the variant alleles, 511C∏ T, is present at an unexpectedly high frequency in the general population, as was the case for 625G∏ A, together conferring susceptibility to ethylmalonic aciduria. Hum Mol Genet 7: 619–627.
Flott-Rahmel B, Falter C, Schluff P, Fingerhut R, Christensen E, Jakobs C, Musshoff U, Fautek JD, Deufel T, Ludolph A, Ullrich K 1997 Nerve cell lesions caused by 3-hydroxyglutaric acid: a possible mechanism for neurodegeneration in glutaric acidaemia type I. J Inherit Metab Dis 20: 387–390.
Ullrich K, Flott-Rahmel B, Schluff P, Musshoff Das A, Lücke T, Steinfeld R, Christensen E, Jakobs C, Ludolph A, Neu A, Röper R 1999 Glutaric aciduria type I: pathomechanisms of neurodegeneration. J Inherit Metab Dis 22: 392–403.
Andresen BS, Bross P, Vianey-Saban C, Divry P, Zabot M, Roe CR, Nada MA, Byskov A, Kruse TA, Neve S, Kristiansen K, Knudsen I, Corydon MJ, Gregersen N 1996 Cloning and characterization of human very-long-chain acyl-CoA dehydrogenase cDNA, chromosomal assignment of the gene and identification in four patients of nine different mutations within the VLCAD gene. Hum Mol Genet 5: 461–472.
Shukla A, Dikshit M, Srimal RC 1995 Nitric oxide modulates blood-brain barrier permeability during infections with an inactivated bacterium. Neuroreport 6: 1629–1632.
Shinawi M, Gruener N, Lerner A 1998 CSF levels of carnitine in children with meningitis, neurologic disorders, acute gastroenteritis, and seizure. Neurology 50: 1869–1871.
Hathaway CA, Appleyard CB, Percy WH, Williams JL 1998 Collitis increases blood-brain barrier permeability in rabbits. FASEB J 12: A391
Land JM, Goulder P, Johnson A, Hockaday J 1992 Glutaric aciduria type I: an atypical presentation together with some observations upon treatment and the possible cause of cerebral damage. Neuropediatrics 23: 322–326.
Ribes A, Riudor E, Briones P, Christensen E, Campistol J, Millington DS 1992 Significance of bound glutarate in the diagnosis of glutaric aciduria type I. J Inherit Metab Dis 15: 367–370.
Acknowledgements
The authors thank Dr. S.I. Goodman (University of Colorado School of Medicine, Denver, CO, U.S.A.), who provided us with information on GCDH gene structure so that we might undertake our molecular studies. We also thank C. Ogg and C. Llordés for their excellent technical assistance, and Laura Gort for her help and advice concerning mutation studies. We also thank the GA I families and the following pediatricians and biochemists who contributed to this study by providing fibroblasts or blood samples: J. Artigas, G. Pintos, P. Villanueva, M.A. Ruiz, M. Tallada, M.A. Vilaseca, M.A. Fernández, E. Riudor, M. Aparicio, G. Lorenzo, I. Corral, C. Garzo, G. Bueno, C. Pedrón, and J.C. Cabrera.
Author information
Authors and Affiliations
Additional information
This work was supported by Fondo de Investigaciones Sanitarias (FIS) grant 97/0049–01.
Hospital Niño Jesús, Madrid, 28009, Spain
Rights and permissions
About this article
Cite this article
Busquets, C., Merinero, B., Christensen, E. et al. Glutaryl-CoA Dehydrogenase Deficiency in Spain: Evidence of Two Groups of Patients, Genetically, and Biochemically Distinct. Pediatr Res 48, 315–322 (2000). https://doi.org/10.1203/00006450-200009000-00009
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1203/00006450-200009000-00009
This article is cited by
-
Modeling Glutaric Aciduria Type I in human neuroblastoma cells recapitulates neuronal damage that can be rescued by gene replacement
Gene Therapy (2024)
-
Two novel compound heterozygous variants of the GCDH gene in two Chinese families with glutaric acidaemia type I identified by high-throughput sequencing and a literature review
Molecular Genetics and Genomics (2023)
-
Probability of high-risk genetic matching with oocyte and semen donors: complete gene analysis or genotyping test?
Journal of Assisted Reproduction and Genetics (2022)
-
Clinical, biochemical and molecular findings of 24 Brazilian patients with glutaric acidemia type 1: 4 novel mutations in the GCDH gene
Metabolic Brain Disease (2021)
-
Clinical, biochemical, neuroradiological and molecular characterization of Egyptian patients with glutaric acidemia type 1
Metabolic Brain Disease (2019)