Abstract
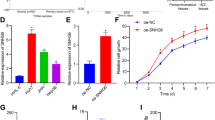
Hepatocellular carcinoma (HCC) is a frequent form of cancer with a poor prognosis and with limited possibilities for medical intervention. Recent evidence has accumulated that long noncoding RNAs (lncRNAs) are important regulators of disease processes including cancer. Chromatin remodeling in cancer cells may result in an unusual expression of lncRNAs and indeed it has been shown that more than 7000 unannotated lncRNAs are expressed in HCCs. We identified a novel long intergenic noncoding RNA, Linc00176, that plays a role in proliferation and survival of HCC. Linc00176 regulates expression of more than 200 genes by the sponge function for tumor suppressor miRNAs, miR-9 and miR-185. Linc00176 is expressed at a high level only in HCC, and is activated by Myc, Max and AP-4 transcription regulators. Myc also upregulates miR-9 and miR-185. In Linc00176-depleted HCC, these miRNAs were released from Linc00176 and downregulated their target mRNAs. Thus, depletion of Linc00176 disrupted the cell cycle and induced necroptosis in HCC via released tumor suppressor miRNAs. These data indicate that atypically expressed lncRNAs may be useful targets for cancer therapy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
Abbreviations
- LincRNA:
-
long intergenic noncoding RNA
- HCC:
-
hepatocellular carcinoma
- THOC5 :
-
suppressors of the transcriptional defects of hpr1 delta by overexpression
- GST:
-
glutathione S transferase
- DAPI:
-
4′,6-diamidin-2-phenylindole
- TUNEL:
-
terminal deoxynucleotidyl transferase dUTP nick end labeling.
References
Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D . Global cancer statistics. CA Cancer J Clin 2011; 61: 69–90.
Whittaker S, Marais R, Zhu AX . The role of signaling pathways in the development and treatment of hepatocellular carcinoma. Oncogene 2010; 29: 4989–5005.
Ferrin G, Aguilar-Melero P, Rodriguez-Peralvarez M, Montero-Alvarez JL, de la Mata M . Biomarkers for hepatocellular carcinoma: diagnostic and therapeutic utility. Hepat Med 2015; 7: 1–10.
Schulze K, Imbeaud S, Letouze E, Alexandrov LB, Calderaro J, Rebouissou S et al. Exome sequencing of hepatocellular carcinomas identifies new mutational signatures and potential therapeutic targets. Nat Genet 2015; 47: 505–511.
Consortium EP Consortium EP Birney E Consortium EP Stamatoyannopoulos JA Consortium EP Dutta A Consortium EP Guigo R Consortium EP Gingeras TR et al. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature 2007; 447: 799–816.
Evans JR, Feng FY, Chinnaiyan AM . The bright side of dark matter: lncRNAs in cancer. J Clin Invest 2016; 126: 2775–2782.
Iyer MK, Niknafs YS, Malik R, Singhal U, Sahu A, Hosono Y et al. The landscape of long noncoding RNAs in the human transcriptome. Nat Genet 2015; 47: 199–208.
Yang Y, Chen L, Gu J, Zhang H, Yuan J, Lian Q et al. Recurrently deregulated lncRNAs in hepatocellular carcinoma. Nat Commun 2017; 8: 14421.
Chen LL . Linking long noncoding RNA localization and function. Trends Biochem Sci 2016; 41: 761–772.
Saran S, Tran DD, Ewald F, Koch A, Hoffmann A, Koch M et al. Depletion of three combined THOC5 mRNA export protein target genes synergistically induces human hepatocellular carcinoma cell death. Oncogene 2015; 35: 3872–3879.
Padgett RA . New connections between splicing and human disease. Trends Genet 2012; 28: 147–154.
Martin G, Gruber AR, Keller W, Zavolan M . Genome-wide analysis of pre-mRNA 3' end processing reveals a decisive role of human cleavage factor I in the regulation of 3' UTR length. Cell Rep 2012; 1: 753–763.
Luscher B, Vervoorts J . Regulation of gene transcription by the oncoprotein MYC. Gene 2012; 494: 145–160.
Jung P, Hermeking H . The c-MYC-AP4-p21 cascade. Cell Cycle 2009; 8: 982–989.
Bossone SA, Asselin C, Patel AJ, Marcu KB . MAZ, a zinc finger protein, binds to c-MYC and C2 gene sequences regulating transcriptional initiation and termination. Proc Natl Acad Sci USA 1992; 89: 7452–7456.
Deng H, Jiang Q, Yang Y, Zhang S, Ma Y, Xie G et al. Intravenous liposomal delivery of the short hairpin RNAs against Plk1 controls the growth of established human hepatocellular carcinoma. Cancer Biol Ther 2011; 11: 401–409.
Remijsen Q, Goossens V, Grootjans S, Van den Haute C, Vanlangenakker N, Dondelinger Y et al. Depletion of RIPK3 or MLKL blocks TNF-driven necroptosis and switches towards a delayed RIPK1 kinase-dependent apoptosis. Cell Death Dis 2014; 5: e1004.
Steitz JA, Vasudevan S . miRNPs: versatile regulators of gene expression in vertebrate cells. Biochem Soc Trans 2009; 37 (Pt 5): 931–935.
Betel D, Koppal A, Agius P, Sander C, Leslie C . Comprehensive modeling of microRNA targets predicts functional non-conserved and non-canonical sites. Genome Biol 2010; 11: R90.
Zhang J, Cheng J, Zeng Z, Wang Y, Li X, Xie Q et al. Comprehensive profiling of novel microRNA-9 targets and a tumor suppressor role of microRNA-9 via targeting IGF2BP1 in hepatocellular carcinoma. Oncotarget 2015; 6: 42040–42052.
Wu WL, Wang WY, Yao WQ, Li GD . Suppressive effects of microRNA-16 on the proliferation, invasion and metastasis of hepatocellular carcinoma cells. Int J Mol Med 2015; 36: 1713–1719.
Zhu XC, Dong QZ, Zhang XF, Deng B, Jia HL, Ye QH et al. microRNA-29a suppresses cell proliferation by targeting SPARC in hepatocellular carcinoma. Int J Mol Med 2012; 30: 1321–1326.
Kim HS, Lee KS, Bae HJ, Eun JW, Shen Q, Park SJ et al. MicroRNA-31 functions as a tumor suppressor by regulating cell cycle and epithelial-mesenchymal transition regulatory proteins in liver cancer. Oncotarget 2015; 6: 8089–8102.
Dang YW, Zeng J, He RQ, Rong MH, Luo DZ, Chen G . Effects of miR-152 on cell growth inhibition, motility suppression and apoptosis induction in hepatocellular carcinoma cells. Asian Pac J Cancer Prev 2014; 15: 4969–4976.
Qadir XV, Han C, Lu D, Zhang J, Wu T . miR-185 inhibits hepatocellular carcinoma growth by targeting the DNMT1/PTEN/Akt pathway. Am J Pathol 2014; 184: 2355–2364.
Yoon JH, Srikantan S, Gorospe M . MS2-TRAP (MS2-tagged RNA affinity purification): tagging RNA to identify associated miRNAs. Methods 2012; 58: 81–87.
Derrien T, Johnson R, Bussotti G, Tanzer A, Djebali S, Tilgner H et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res 2012; 22: 1775–1789.
Higashi T, Hayashi H, Ishimoto T, Takeyama H, Kaida T, Arima K et al. miR-9-3p plays a tumour-suppressor role by targeting TAZ (WWTR1) in hepatocellular carcinoma cells. Br J Cancer 2015; 113: 252–258.
Cai L, Cai X . Up-regulation of miR-9 expression predicate advanced clinicopathological features and poor prognosis in patients with hepatocellular carcinoma. Diagn Pathol 2014; 9: 1000.
Drakaki A, Hatziapostolou M, Polytarchou C, Vorvis C, Poultsides GA, Souglakos J et al. Functional microRNA high throughput screening reveals miR-9 as a central regulator of liver oncogenesis by affecting the PPARA-CDH1 pathway. BMC Cancer 2015; 15: 542.
Sun J, Fang K, Shen H, Qian Y . MicroRNA-9 is a ponderable index for the prognosis of human hepatocellular carcinoma. Int J Clin Exp Med 2015; 8: 17748–17756.
Zhu S, Li W, Liu J, Chen CH, Liao Q, Xu P et al. Genome-scale deletion screening of human long non-coding RNAs using a paired-guide RNA CRISPR-Cas9 library. Nat Biotechnol 2016; 34: 1279–1286.
Levens D . How the c-myc promoter works and why it sometimes does not. J Natl Cancer Inst Monogr 2008; 39: 41–43.
Liu SJ, Horlbeck MA, Cho SW, Birk HS, Malatesta M, He D et al. CRISPRi-based genome-scale identification of functional long noncoding RNA loci in human cells. Science 2017; 355: pii.aah7111.
Tran DD, Koch A, Allister A, Saran S, Ewald F, Koch M et al. Treatment with MAPKAP2 (MK2) inhibitor and DNA methylation inhibitor, 5-aza dC, synergistically triggers apoptosis in hepatocellular carcinoma (HCC) via tristetraprolin (TTP). Cell Signal 2016; 28: 1872–1880.
Guria A, Tran DD, Ramachandran S, Koch A, El Bounkari O, Dutta P et al. Identification of mRNAs that are spliced but not exported to the cytoplasm in the absence of THOC5 in mouse embryo fibroblasts. RNA 2011; 17: 1048–1056.
Tamura T, Mancini A, Joos H, Koch A, Hakim C, Dumanski J et al. FMIP, a novel Fms-interacting protein, affects granulocyte/macrophage differentiation. Oncogene 1999; 18: 6488–6495.
Yang JS, Nam HJ, Seo M, Han SK, Choi Y, Nam HG et al. OASIS: online application for the survival analysis of lifespan assays performed in aging research. PLoS ONE 2011; 6: e23525.
Acknowledgements
We thank C Bruce Boschek for critically reading the manuscript. This work is a part of thesis of CK. This research was supported by Deutsche Krebshilfe (111153), DFG Ta-111/13-3, Niedersächsische Krebsgesellschaft to AK and DDHT, PhD program Molecular Medicine and Structure Medicine in HBRS and Leistungsorientierte Mittelvergabe with Frauenfaktor from MHH.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Rights and permissions
About this article
Cite this article
Tran, D., Kessler, C., Niehus, S. et al. Myc target gene, long intergenic noncoding RNA, Linc00176 in hepatocellular carcinoma regulates cell cycle and cell survival by titrating tumor suppressor microRNAs. Oncogene 37, 75–85 (2018). https://doi.org/10.1038/onc.2017.312
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2017.312
This article is cited by
-
Cadmium Exposure Induces Apoptosis and Necrosis of Thyroid Cells via the Regulation of miR-494-3p/PTEN Axis
Biological Trace Element Research (2024)
-
Risk coefficient model of necroptosis-related lncRNA in predicting the prognosis of patients with lung adenocarcinoma
Scientific Reports (2022)
-
C-Myc-activated long non-coding RNA LINC01050 promotes gastric cancer growth and metastasis by sponging miR-7161-3p to regulate SPZ1 expression
Journal of Experimental & Clinical Cancer Research (2021)
-
LncRNA LINC00337 sponges mir-1285-3p to promote proliferation and metastasis of lung adenocarcinoma cells by upregulating YTHDF1
Cancer Cell International (2021)
-
MYBL2-induced PITPNA-AS1 upregulates SIK2 to exert oncogenic function in triple-negative breast cancer through miR-520d-5p and DDX54
Journal of Translational Medicine (2021)