Abstract
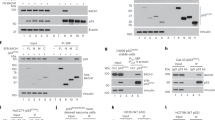
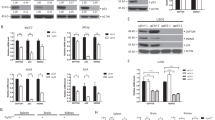
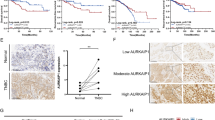
Tumor suppressor protein p19ARF (Arf; p14ARF in humans) functions in both p53-dependent and -independent modes to counteract hyper-proliferative signals caused by proto-oncogene activation, but its p53-independent activities remain poorly understood. Using the tandem affinity purification-tag technique, we purified Arf-containing protein complexes and identified p68 DEAD-box protein (DDX5) as a novel interacting protein of Arf. In this study, we found that DDX5 interacts with c-Myc, and harbors essential roles for c-Myc-mediated transcription and its transforming activity. Furthermore, when c-Myc was forcibly expressed, the expression level of DDX5 protein was drastically increased through the acceleration of protein synthesis of DDX5, suggesting the presence of an oncogenic positive feedback loop including c-Myc and DDX5. Strikingly, Arf blocked the physical interaction between DDX5 and c-Myc, and drove away DDX5 from the promoter of c-Myc target genes. These observations most likely indicate the mechanism by which Arf causes p53-independent tumor-suppressive activity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout










Similar content being viewed by others
References
Serrano M, Hannon GJ, Beach D . A new regulatory motif in cell-cycle control causing specific inhibition of cyclin D/CDK4. Nature 1993; 366: 704–707.
Quelle DE, Zindy F, Ashmun RA, Sherr CJ . Alternative reading frames of the INK4a tumor suppressor gene encode two unrelated proteins capable of inducing cell cycle arrest. Cell 1995; 83: 993–1000.
Kamijo T, Zindy F, Roussel MF, Quelle DE, Downing JR, Ashmun RA et al. Tumor suppression at the mouse INK4a locus mediated by the alternative reading frame product p19ARF. Cell 1997; 91: 649–659.
Zhang Y, Xiong Y, Yarbrough WG . ARF promotes MDM2 degradation and stabilizes p53: ARF-INK4a locus deletion impairs both the Rb and p53 tumor suppression pathways. Cell 1998; 92: 725–734.
Kamijo T, Weber JD, Zambetti G, Zindy F, Roussel MF, Sherr CJ . Functional and physical interactions of the ARF tumor suppressor with p53 and Mdm2. Proc Natl Acad Sci USA 1998; 95: 8292–8297.
Honda R, Yasuda H . Association of p19ARF with Mdm2 inhibits ubiquitin ligase activity of Mdm2 for tumor suppressor p53. EMBO J 1999; 18: 22–27.
Dornan D, Wertz I, Shimizu H, Arnott D, Frantz GD, Dowd P et al. The ubiquitin ligase COP1 is a critical negative regulator of p53. Nature 2004; 429: 86–92.
Chen D, Kon N, Li M, Zhang W, Qin J, Gu W . ARF-BP1/Mule is a critical mediator of the ARF tumor suppressor. Cell 2005; 121: 1071–1083.
Kelly-Spratt KS, Gurley KE, Yasui Y, Kemp CJ . p19Arf suppresses growth, progression, and metastasis of H-ras-driven carcinomas through p53-dependent and-independent pathways. PLoS Biol 2004; 2: 1138–1149.
Carnero A, Hudson JD, Price CM, Beach DH . p16INK4A and p19ARF act in overlapping pathways in cellular immortalization. Nat Cell Biol 2000; 2: 148–155.
Eischen CM, Weber JD, Roussel MF, Sherr C, Cleveland JL . Disruption of the ARF-Mdm2-p53 tumor suppressor pathway in Myc-induced lymphomagenesis. Genes Dev 1999; 13: 2658–2669.
Weber JD, Jeffers JR, Rehg JE, Randle DH, Lozano G, Roussel MF et al. p53-independent functions of the p19ARF tumor suppressor. Genes Dev 2000; 14: 2358–2365.
Sugimoto M, Kuo ML, Roussel MF, Sherr CJ . Nucleolar Arf tumor suppressor inhibits ribosomal RNA processing. Mol Cell 2003; 11: 415–424.
Itahana K, Bhat KP, Jin A, Itahana Y, Hawke D, Kobayashi R et al. Tumor suppressor ARF degrades B23, a nucleolar protein involved in ribosome biogenesis and cell proliferation. Mol Cell 2003; 12: 1151–1164.
Bertwistle D, Sugimoto M, Sherr CJ . Physical and functional interactions of the Arf tumor suppressor protein with nucleophosmin/B23. Mol Cell Biol 2004; 24: 985–996.
Martelli F, Hamilton T, Silver DP, Sharpless NE, Bardeesy N, Rokas M et al. p19ARF targets certain E2F species for degradation. Proc Natl Acad Sci USA 2001; 98: 4455–4460.
Eymin B, Karayan L, Seite P, Brambilla C, Brambilla E, Larsen CJ et al. Human ARF binds E2F1 and inhibits its transcriptional activity. Oncogene 2001; 20: 1033–1041.
Datta A, Nag A, Pan W, Hay N, Gartel AL, Colamonici O et al. Myc-ARF (alternate reading frame) interaction inhibits the functions of Myc. J Biol Chem 2004; 279: 36698–36707.
Qi Y, Gregory MA, Li Z, Brousal JP, West K, Hann SR . p19ARF directly and differentially controls the functions of c-Myc independently of p53. Nature 2004; 431: 712–717.
Rocha S, Campbell KJ, Perkins ND . p53- and Mdm2-independent repression of NF-κB transactivation by the ARF tumor suppressor. Mol Cell 2003; 12: 15–25.
Xirodimas DP, Chisholm J, Desterro JM, Lane DP, Hay RT . p14ARF promotes accumulation of SUMO-1 conjugated (H) Mdm2. FEBS Lett 2002; 528: 207–211.
Chen L, Chen J . MDM2-ARF complex regulates p53 sumoylation. Oncogene 2003; 22: 5348–5357.
Kuo ML, den Besten W, Thomas MC, Sherr CJ . Arf-induced turnover of the nucleolar nucleophosmin-associated SUMO-2/3 protease Senp3. Cell Cycle 2008; 7: 3378–3387.
Lane DP, Hoeffler WK . SV40 large T shares an antigenic determinant with a cellular protein of molecular weight 68 000. Nature 1980; 288: 167–170.
Hirling H, Scheffner M, Restle T, Stahl H . RNA helicase activity associated with the human p68 protein. Nature 1989; 339: 562–564.
Heinlein UA . Dead box for the living. J Pathol 1998; 184: 345–347.
Stevenson RJ, Hamilton SJ, MacCallum DE, Hall PA, Fuller-Pace FV . Expression of the ‘dead box’ RNA helicase p68 is developmentally and growth regulated and correlates with organ differentiation/maturation in the fetus. J Pathol 1998; 184: 351–359.
Dubey P, Hendrickson RC, Meredith SC, Siegel CT, Shabanowitz J, Skipper JC et al. The immunodominant antigen of an ultraviolet-induced regressor tumor is generated by a somatic point mutation in the DEAD box helicase p68. J Exp Med 1997; 185: 695–705.
Causevic M, Hislop RG, Kernohan NM, Carey FA, Kay RA, Steele RJ et al. Overexpression and poly-ubiquitylation of the DEAD-box RNA helicase p68 in colorectal tumours. Oncogene 2001; 20: 7734–7743.
Yang L, Lin C, Liu ZR . Phosphorylations of DEAD box p68 RNA helicase are associated with cancer development and cell proliferation. Mol Cancer Res 2005; 3: 355–363.
Yang L, Lin C, Liu ZR . P68 RNA helicase mediates PDGF-induced epithelial mesenchymal transition by displacing Axin from beta-catenin. Cell 2006; 127: 139–155.
Saporita AJ, Chang HC, Winkeler CL, Apicelli AJ, Kladney RD, Wang J et al. RNA Helicase DDX5 Is a p53-Independent Target of ARF That Participates in Ribosome Biogenesis. Cancer Res 2011; 71: 6708–6717.
Eilers M, Schirm S, Bishop JM . The MYC protein activates transcription of the α-prothymosin gene. EMBO J 1991; 10: 133–141.
Littlewood TD, Hancock DC, Danielian PS, Parker MG, Evan GI . A modified oestrogen receptor ligand-binding domain as an improved switch for the regulation of heterologous proteins. Nucleic Acids Res 1995; 23: 1686–1690.
Packham G, Cleveland JL . Induction of ornithine decarboxylase by IL-3 is mediated by sequential c-Myc-independent and c-Myc-dependent pathways. Oncogene 1997; 15: 1219–1232.
Chang DW, Claassen GF, Hann SR, Cole MD . The c-Myc transactivation domain is a direct modulator of apoptotic versus proliferative signals. Mol Cell Biol 2000; 20: 4309–4319.
Tago K, Chiocca S, Sherr CJ . Sumoylation induced by the Arf tumor suppressor: a p53-independent function. Proc Natl Acad Sci USA 2005; 102: 7689–7694.
Jacobs AM, Nicol SM, Hislop RG, Jaffray EG, Hay RT, Fuller-Pace FV . SUMO modification of the DEAD box protein p68 modulates its transcriptional activity and promotes its interaction with HDAC1. Oncogene 2007; 26: 5866–5876.
Bates GJ, Nicol SM, Wilson BJ, Jacobs AM, Bourdon JC, Wardrop J et al. The DEAD box protein p68: a novel transcriptional coactivator of the p53 tumour suppressor. EMBO J 2005; 24: 543–553.
Reinhardt HC, Yaffe MB . Kinases that control the cell cycle in response to DNA damage: Chk1, Chk2, and MK2. Curr Opin Cell Biol 2009; 21: 245–255.
Dang CV . MYC on the path to cancer. Cell 2012; 149: 22–35.
Gao P, Tchernyshyov I, Chang TC, Lee YS, Kita K, Ochi T et al. c-Myc suppression of miR-23a/b enhances mitochondrial glutaminase expression and glutamine metabolism. Nature 2009; 458: 762–765.
Wise DR, DeBerardinis RJ, Mancuso A, Sayed N, Zhang XY, Pfeiffer HK et al. Myc regulates a transcriptional program that stimulates mitochondrial glutaminolysis and leads to glutamine addiction. Proc Natl Acad Sci USA 2008; 105: 18782–18787.
Martin DE, Powers T, Hall MN . Regulation of ribosome biogenesis: where is TOR? Cell Metab 2006; 4: 259–260.
Durán RV, Oppliger W, Robitaille AM, Heiserich L, Skendaj R, Gottlieb E et al. Glutaminolysis activates Rag-mTORC1 signaling. Mol Cell 2012; 47: 349–358.
Zindy F, Eischen CM, Randle DH, Kamijo T, Cleveland JL, Sherr CJ et al. Myc signaling via the ARF tumor suppressor regulates p53-dependent apoptosis and immortalization. Genes Dev 1998; 12: 2424–2433.
Washburn MP, Wolters D, Yates JR 3rd . Large-scale analysis of the yeast proteome by multidimensional protein identification technology. Nat Biotechnol 2001; 19: 242–247.
Frank SR, Schroeder M, Fernandez P, Taubert S, Amati B . Binding of c-Myc to chromatin mediates mitogen-induced acetylation of histone H4 and gene activation. Genes Dev 2001; 15: 2069–2082.
Acknowledgements
We thank Charles Sherr and Martine Roussel for critical advice and reagents, and Michael Washburn of Stowers Institute for Medical Research Proteomics Center for performing MudPIT analysis. This work was supported by a grant-in-aid for scientific research from the MEXT (20770103), MEXT-supported program for the strategic research foundation at private universities (2013–2017) and the Takeda Science Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Tago, K., Funakoshi-Tago, M., Itoh, H. et al. Arf tumor suppressor disrupts the oncogenic positive feedback loop including c-Myc and DDX5. Oncogene 34, 314–322 (2015). https://doi.org/10.1038/onc.2013.561
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2013.561
Keywords
This article is cited by
-
Regulation of apoptotic pathways during endometriosis: from the molecular basis to the future perspectives
Archives of Gynecology and Obstetrics (2016)