Abstract
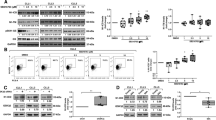
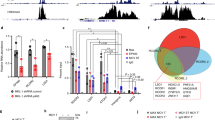
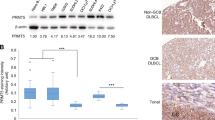
Inappropriate activation of the NOTCH signaling pathway, for example, by activating mutations, contributes to the pathogenesis of various human malignancies. Here, we demonstrate that aberrant expression of an essential NOTCH coactivator of the Mastermind-like (MAML) family provides an alternative mechanism to activate NOTCH signaling in human lymphoma cells. We detected high-level MAML2 expression in several B cell-derived lymphoma types, including classical Hodgkin lymphoma (cHL) cells, relative to normal B cells. Inhibition of MAML-protein activity by a dominant negative form of MAML or by small hairpin RNAs targeting MAML2 in cHL cells resulted in downregulation of the NOTCH target genes HES7 and HEY1, which we identified as overexpressed in cHL cells, and in reduced proliferation. Furthermore, a NOTCH gene-expression signature in cHL cells confirmed their cell-autonomous NOTCH activity. Finally, in line with the essential role of MAML proteins for assembly and activity of the NOTCH transcriptional complex (NTC), we show that MAML-derived small-peptide constructs block NOTCH activity and disrupt NTC formation in vitro. These data strongly suggest direct targeting of the NTC as treatment strategy for NOTCH-dependent malignancies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Bessho Y, Miyoshi G, Sakata R, Kageyama R . (2001). Hes7: a bHLH-type repressor gene regulated by Notch and expressed in the presomitic mesoderm. Genes Cells 6: 175–185.
Brummelkamp TR, Bernards R, Agami R . (2002). A system for stable expression of short interfering RNAs in mammalian cells. Science 296: 550–553.
Del Bianco C, Aster JC, Blacklow SC . (2008). Mutational and energetic studies of Notch 1 transcription complexes. J Mol Biol 376: 131–140.
Ellisen LW, Bird J, West DC, Soreng AL, Reynolds TC, Smith SD et al. (1991). TAN-1, the human homolog of the Drosophila notch gene, is broken by chromosomal translocations in T lymphoblastic neoplasms. Cell 66: 649–661.
Fryer CJ, Lamar E, Turbachova I, Kintner C, Jones KA . (2002). Mastermind mediates chromatin-specific transcription and turnover of the Notch enhancer complex. Genes Dev 16: 1397–1411.
Grabher C, von Boehmer H, Look AT . (2006). Notch 1 activation in the molecular pathogenesis of T-cell acute lymphoblastic leukaemia. Nat Rev Cancer 6: 347–359.
Iso T, Kedes L, Hamamori Y . (2003). HES and HERP families: multiple effectors of the Notch signaling pathway. J Cell Physiol 194: 237–255.
Janz M, Hummel M, Truss M, Wollert-Wulf B, Mathas S, Jöhrens K et al. (2006). Classical Hodgkin lymphoma is characterized by high constitutive expression of activating transcription factor 3 (ATF3), which promotes viability of Hodgkin/Reed-Sternberg cells. Blood 107: 2536–2539.
Jeffries S, Robbins DJ, Capobianco AJ . (2002). Characterization of a high-molecular-weight Notch complex in the nucleus of Notch(ic)-transformed RKE cells and in a human T-cell leukemia cell line. Mol Cell Biol 22: 3927–3941.
Jin B, Shen H, Lin S, Li JL, Chen Z, Griffin JD et al. (2010). The mastermind-like 1 (MAML1) co-activator regulates constitutive NF-kappaB signaling and cell survival. J Biol Chem 285: 14356–14365.
Jundt F, Acikgoez O, Kwon SH, Schwarzer R, Anagnostopoulos I, Wiesner B et al. (2008). Aberrant expression of Notch1 interferes with the B-lymphoid phenotype of neoplastic B cells in classical Hodgkin lymphoma. Leukemia 22: 1587–1594.
Jundt F, Anagnostopoulos I, Förster R, Mathas S, Stein H, Dörken B . (2002). Activated Notch1 signaling promotes tumor cell proliferation and survival in Hodgkin and anaplastic large cell lymphoma. Blood 99: 3398–3403.
Jundt F, Pröbsting KS, Anagnostopoulos I, Muehlinghaus G, Chatterjee M, Mathas S et al. (2004). Jagged1-induced Notch signaling drives proliferation of multiple myeloma cells. Blood 103: 3511–3515.
Kapp U, Yeh WC, Patterson B, Elia AJ, Kagi D, Ho A et al. (1999). Interleukin 13 is secreted by and stimulates the growth of Hodgkin and Reed-Sternberg cells. J Exp Med 189: 1939–1946.
Koch U, Radtke F . (2007). Notch and cancer: a double-edged sword. Cell Mol Life Sci 64: 2746–2762.
Kopan R, Ilagan MX . (2009). The canonical Notch signaling pathway: unfolding the activation mechanism. Cell 137: 216–233.
Lee SY, Kumano K, Nakazaki K, Sanada M, Matsumoto A, Yamamoto G et al. (2009). Gain-of-function mutations and copy number increases of Notch2 in diffuse large B-cell lymphoma. Cancer Sci 100: 920–926.
Li X, Gounari F, Protopopov A, Khazaie K, von Boehmer H . (2008). Oncogenesis of T-ALL and nonmalignant consequences of overexpressing intracellular NOTCH1. J Exp Med 205: 2851–2861.
Lietz A, Janz M, Sigvardsson M, Jundt F, Dörken B, Mathas S . (2007). Loss of bHLH transcription factor E2A activity in primary effusion lymphoma confers resistance to apoptosis. Br J Haematol 137: 342–348.
Lin SE, Oyama T, Nagase T, Harigaya K, Kitagawa M . (2002). Identification of new human mastermind proteins defines a family that consists of positive regulators for notch signaling. J Biol Chem 277: 50612–50620.
Lubman OY, Ilagan MX, Kopan R, Barrick D . (2007). Quantitative dissection of the Notch:CSL interaction: insights into the Notch-mediated transcriptional switch. J Mol Biol 365: 577–589.
Maillard I, Weng AP, Carpenter AC, Rodriguez CG, Sai H, Xu L et al. (2004). Mastermind critically regulates Notch-mediated lymphoid cell fate decisions. Blood 104: 1696–1702.
Mathas S, Janz M, Hummel F, Hummel M, Wollert-Wulf B, Lusatis S et al. (2006). Intrinsic inhibition of transcription factor E2A by HLH proteins ABF-1 and Id2 mediates reprogramming of neoplastic B cells in Hodgkin lymphoma. Nat Immunol 7: 207–215.
McElhinny AS, Li JL, Wu L . (2008). Mastermind-like transcriptional co-activators: emerging roles in regulating cross talk among multiple signaling pathways. Oncogene 27: 5138–5147.
Moellering RE, Cornejo M, Davis TN, Del Bianco C, Aster JC, Blacklow SC et al. (2009). Direct inhibition of the NOTCH transcription factor complex. Nature 462: 182–188.
Murre C . (2005). Helix-loop-helix proteins and lymphocyte development. Nat Immunol 6: 1079–1086.
Nam Y, Sliz P, Song L, Aster JC, Blacklow SC . (2006). Structural basis for cooperativity in recruitment of MAML coactivators to Notch transcription complexes. Cell 124: 973–983.
Nam Y, Weng AP, Aster JC, Blacklow SC . (2003). Structural requirements for assembly of the CSL. intracellular Notch1. Mastermind Master-like 1 transcriptional activation complex. J Biol Che 278: 21232–21239.
Nie L, Xu M, Vladimirova A, Sun XH . (2003). Notch-induced E2A ubiquitination and degradation are controlled by MAP kinase activities. EMBO J 22: 5780–5792.
O'Neil J, Grim J, Strack P, Rao S, Tibbitts D, Winter C et al. (2007). FBW7 mutations in leukemic cells mediate NOTCH pathway activation and resistance to gamma-secretase inhibitors. J Exp Med 204: 1813–1824.
Oyama T, Harigaya K, Muradil A, Hozumi K, Habu S, Oguro H et al. (2007). Mastermind-1 is required for Notch signal-dependent steps in lymphocyte development in vivo. Proc Natl Acad Sci USA 104: 9764–9769.
Rosati E, Sabatini R, Rampino G, Tabilio A, Di Ianni M, Fettucciari K et al. (2009). Constitutively activated Notch signaling is involved in survival and apoptosis resistance of B-CLL cells. Blood 113: 856–865.
Stanelle J, Döring C, Hansmann ML, Küppers R . (2010). Mechanisms of aberrant GATA-3 expression in classical Hodgkin lymphoma and its consequences for the cytokine profile of Hodgkin and Reed/Sternberg cells. Blood (doi:10.1182/blood2010-01-265827).
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA et al. (2005). Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102: 15545–15550.
Tonon G, Modi S, Wu L, Kubo A, Coxon AB, Komiya T et al. (2003). t(11;19)(q21;p13) translocation in mucoepidermoid carcinoma creates a novel fusion product that disrupts a Notch signaling pathway. Nat Genet 33: 208–213.
Weng AP, Ferrando AA, Lee W, Morris IV JP, Silverman LB, Sanchez-Irizarry C et al. (2004). Activating mutations of NOTCH1 in human T cell acute lymphoblastic leukemia. Science 306: 269–271.
Weng AP, Nam Y, Wolfe MS, Pear WS, Griffin JD, Blacklow SC et al. (2003). Growth suppression of pre-T acute lymphoblastic leukemia cells by inhibition of notch signaling. Mol Cell Biol 23: 655–664.
Wilson JJ, Kovall RA . (2006). Crystal structure of the CSL-Notch-Mastermind ternary complex bound to DNA. Cell 124: 985–996.
Wu L, Aster JC, Blacklow SC, Lake R, Artavanis-Tsakonas S, Griffin JD . (2000). MAML1, a human homologue of Drosophila mastermind, is a transcriptional co-activator for NOTCH receptors. Nat Genet 26: 484–489.
Wu L, Maillard I, Nakamura M, Pear WS, Griffin JD . (2007). The transcriptional coactivator Maml1 is required for Notch2-mediated marginal zone B-cell development. Blood 110: 3618–3623.
Wu L, Sun T, Kobayashi K, Gao P, Griffin JD . (2002). Identification of a family of mastermind-like transcriptional coactivators for mammalian notch receptors. Mol Cell Biol 22: 7688–7700.
Acknowledgements
We thank Caroline Gärtner (Berlin), Melanie Manzke (Berlin) and Simone Kressmann (Berlin) for outstanding technical assistance, and Peter Rahn (Berlin) for cell sorting. We thank Raphael Kopan (Washington) for the Hes1-pGL2 reporter construct, Lizi Wu (Gainesville, Florida) for the MAML3 expression construct, Karin Zimmermann (Berlin) and Ulf Leser (Berlin) for helpful suggestions in regard to gene set analysis, and Ariane Buchal and Benedikt Sedlmaier (both Berlin, Germany) for providing human tonsil material. This work was supported in part by grants from the Deutsche Forschungsgemeinschaft, the Berliner Krebsgesellschaft and the Wilhelm Sander-Stiftung.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Rights and permissions
About this article
Cite this article
Köchert, K., Ullrich, K., Kreher, S. et al. High-level expression of Mastermind-like 2 contributes to aberrant activation of the NOTCH signaling pathway in human lymphomas. Oncogene 30, 1831–1840 (2011). https://doi.org/10.1038/onc.2010.544
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2010.544
Keywords
This article is cited by
-
Advances in targeted therapy for malignant lymphoma
Signal Transduction and Targeted Therapy (2020)
-
Inactivation of the putative ubiquitin-E3 ligase PDLIM2 in classical Hodgkin and anaplastic large cell lymphoma
Leukemia (2017)
-
A roadmap of constitutive NF-κB activity in Hodgkin lymphoma: Dominant roles of p50 and p52 revealed by genome-wide analyses
Genome Medicine (2016)
-
NOTCH1 Signaling as a Therapeutic Target in Sézary Syndrome
Journal of Investigative Dermatology (2012)