Abstract
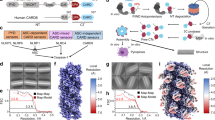
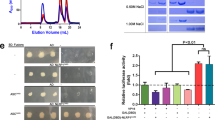
Inflammasomes are cytosolic caspase-1-activation complexes that sense intrinsic and extrinsic danger signals, and trigger inflammatory responses and pyroptotic cell death. Homotypic interactions among Pyrin domains and caspase recruitment domains (CARDs) in inflammasome-complex components mediate oligomerization into filamentous assemblies. Several cytosolic proteins consisting of only interaction domains exert inhibitory effects on inflammasome assembly. In this study, we determined the structure of the human caspase-1 CARD domain (caspase-1CARD) filament by cryo-electron microscopy and investigated the biophysical properties of two caspase-1-like CARD-only proteins: human inhibitor of CARD (INCA or CARD17) and ICEBERG (CARD18). Our results reveal that INCA caps caspase-1 filaments, thereby exerting potent inhibition with low-nanomolar Ki on caspase-1CARD polymerization in vitro and inflammasome activation in cells. Whereas caspase-1CARD uses six complementary surfaces of three types for filament assembly, INCA is defective in two of the six interfaces and thus terminates the caspase-1 filament.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Lamkanfi, M. & Dixit, V.M. Mechanisms and functions of inflammasomes. Cell 157, 1013–1022 (2014).
Latz, E., Xiao, T.S. & Stutz, A. Activation and regulation of the inflammasomes. Nat. Rev. Immunol. 13, 397–411 (2013).
Martinon, F., Mayor, A. & Tschopp, J. The inflammasomes: guardians of the body. Annu. Rev. Immunol. 27, 229–265 (2009).
Ferrao, R. & Wu, H. Helical assembly in the death domain (DD) superfamily. Curr. Opin. Struct. Biol. 22, 241–247 (2012).
Lu, A. et al. Unified polymerization mechanism for the assembly of ASC-dependent inflammasomes. Cell 156, 1193–1206 (2014).
Cai, X. et al. Prion-like polymerization underlies signal transduction in antiviral immune defense and inflammasome activation. Cell 156, 1207–1222 (2014).
Zhang, L. et al. Cryo-EM structure of the activated NAIP2-NLRC4 inflammasome reveals nucleated polymerization. Science 350, 404–409 (2015).
Hu, Z. et al. Structural and biochemical basis for induced self-propagation of NLRC4. Science 350, 399–404 (2015).
Qu, Y. et al. Phosphorylation of NLRC4 is critical for inflammasome activation. Nature 490, 539–542 (2012).
Kofoed, E.M. & Vance, R.E. Innate immune recognition of bacterial ligands by NAIPs determines inflammasome specificity. Nature 477, 592–595 (2011).
Vezzani, A., Maroso, M., Balosso, S., Sanchez, M.A. & Bartfai, T. IL-1 receptor/Toll-like receptor signaling in infection, inflammation, stress and neurodegeneration couples hyperexcitability and seizures. Brain Behav. Immun. 25, 1281–1289 (2011).
Stehlik, C. et al. The PAAD/PYRIN-only protein POP1/ASC2 is a modulator of ASC-mediated nuclear-factor-kappa B and pro-caspase-1 regulation. Biochem. J. 373, 101–113 (2003).
Bedoya, F., Sandler, L.L. & Harton, J.A. Pyrin-only protein 2 modulates NF-kappaB and disrupts ASC:CLR interactions. J. Immunol. 178, 3837–3845 (2007).
Dorfleutner, A. et al. A Shope Fibroma virus PYRIN-only protein modulates the host immune response. Virus Genes 35, 685–694 (2007).
Lamkanfi, M. et al. INCA, a novel human caspase recruitment domain protein that inhibits interleukin-1beta generation. J. Biol. Chem. 279, 51729–51738 (2004).
Humke, E.W., Shriver, S.K., Starovasnik, M.A., Fairbrother, W.J. & Dixit, V.M. ICEBERG: a novel inhibitor of interleukin-1beta generation. Cell 103, 99–111 (2000).
Druilhe, A., Srinivasula, S.M., Razmara, M., Ahmad, M. & Alnemri, E.S. Regulation of IL-1beta generation by Pseudo-ICE and ICEBERG, two dominant negative caspase recruitment domain proteins. Cell Death Differ. 8, 649–657 (2001).
Le, H.T. & Harton, J.A. Pyrin- and CARD-only proteins as regulators of NLR functions. Front. Immunol. 4, 275 (2013).
Kersse, K., Vanden Berghe, T., Lamkanfi, M. & Vandenabeele, P. A phylogenetic and functional overview of inflammatory caspases and caspase-1-related CARD-only proteins. Biochem. Soc. Trans. 35, 1508–1511 (2007).
Lee, S.H., Stehlik, C. & Reed, J.C. Cop, a caspase recruitment domain-containing protein and inhibitor of caspase-1 activation processing. J. Biol. Chem. 276, 34495–34500 (2001).
da Cunha, J.P., Galante, P.A. & de Souza, S.J. Different evolutionary strategies for the origin of caspase-1 inhibitors. J. Mol. Evol. 66, 591–597 (2008).
Wu, B. et al. Molecular imprinting as a signal-activation mechanism of the viral RNA sensor RIG-I. Mol. Cell 55, 511–523 (2014).
Egelman, E.H. The iterative helical real space reconstruction method: surmounting the problems posed by real polymers. J. Struct. Biol. 157, 83–94 (2007).
Lin, S.C., Lo, Y.C. & Wu, H. Helical assembly in the MyD88–IRAK4–IRAK2 complex in TLR/IL-1R signalling. Nature 465, 885–890 (2010).
Park, H.H. et al. Death domain assembly mechanism revealed by crystal structure of the oligomeric PIDDosome core complex. Cell 128, 533–546 (2007).
Lu, A. et al. Plasticity in PYD assembly revealed by cryo-EM structure of the PYD filament of AIM2. Cell Discov. 1, 15013 (2015).
Guimaraes, C.P. et al. Site-specific C-terminal and internal loop labeling of proteins using sortase-mediated reactions. Nat. Protoc. 8, 1787–1799 (2013).
de Alba, E. Structure and interdomain dynamics of apoptosis-associated speck-like protein containing a CARD (ASC). J. Biol. Chem. 284, 32932–32941 (2009).
Tsao, K.L., DeBarbieri, B., Michel, H. & Waugh, D.S. A versatile plasmid expression vector for the production of biotinylated proteins by site-specific, enzymatic modification in Escherichia coli. Gene 169, 59–64 (1996).
Wu, H. Higher-order assemblies in a new paradigm of signal transduction. Cell 153, 287–292 (2013).
Hou, F. et al. MAVS forms functional prion-like aggregates to activate and propagate antiviral innate immune response. Cell 146, 448–461 (2011).
Qiao, Q. et al. Structural architecture of the CARMA1/Bcl10/MALT1 signalosome: nucleation-induced filamentous assembly. Mol. Cell 51, 766–779 (2013).
López-Castejón, G. & Pelegrín, P. Current status of inflammasome blockers as anti-inflammatory drugs. Expert Opin. Investig. Drugs 21, 995–1007 (2012).
Brusselle, G.G., Provoost, S., Bracke, K.R., Kuchmiy, A. & Lamkanfi, M. Inflammasomes in respiratory disease: from bench to bedside. Chest 145, 1121–1133 (2014).
Aguilera, M., Darby, T. & Melgar, S. The complex role of inflammasomes in the pathogenesis of inflammatory bowel diseases: lessons learned from experimental models. Cytokine Growth Factor Rev. 25, 715–730 (2014).
Ridker, P.M. & Lüscher, T.F. Anti-inflammatory therapies for cardiovascular disease. Eur. Heart J. 35, 1782–1791 (2014).
Esser, N., Legrand-Poels, S., Piette, J., Scheen, A.J. & Paquot, N. Inflammation as a link between obesity, metabolic syndrome and type 2 diabetes. Diabetes Res. Clin. Pract. 105, 141–150 (2014).
Robbins, G.R., Wen, H. & Ting, J.P. Inflammasomes and metabolic disorders: old genes in modern diseases. Mol. Cell 54, 297–308 (2014).
Caldwell, J.E., Heiss, S.G., Mermall, V. & Cooper, J.A. Effects of CapZ, an actin capping protein of muscle, on the polymerization of actin. Biochemistry 28, 8506–8514 (1989).
Lin, Y.H., Li, J., Swanson, E.R. & Russell, B. CapZ and actin capping dynamics increase in myocytes after a bout of exercise and abates in hours after stimulation ends. J. Appl. Physiol. (1985) 114, 1603–1609 (2013).
Maun, N.A., Speicher, D.W., DiNubile, M.J. & Southwick, F.S. Purification and properties of a Ca2+-independent barbed-end actin filament capping protein, CapZ, from human polymorphonuclear leukocytes. Biochemistry 35, 3518–3524 (1996).
Cooper, J.A. Effects of cytochalasin and phalloidin on actin. J. Cell Biol. 105, 1473–1478 (1987).
Wakatsuki, T., Schwab, B., Thompson, N.C. & Elson, E.L. Effects of cytochalasin D and latrunculin B on mechanical properties of cells. J. Cell Sci. 114, 1025–1036 (2001).
Carlier, M.F., Criquet, P., Pantaloni, D. & Korn, E.D. Interaction of cytochalasin D with actin filaments in the presence of ADP and ATP. J. Biol. Chem. 261, 2041–2050 (1986).
Shaikh, T.R. et al. SPIDER image processing for single-particle reconstruction of biological macromolecules from electron micrographs. Nat. Protoc. 3, 1941–1974 (2008).
Mindell, J.A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003).
Scheres, S.H. & Chen, S. Prevention of overfitting in cryo-EM structure determination. Nat. Methods 9, 853–854 (2012).
Biasini, M. et al. SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res. 42, W252–W258 (2014).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Pettersen, E.F. et al. UCSF Chimera: a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Winn, M.D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 67, 235–242 (2011).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
DiMaio, F. et al. Atomic-accuracy models from 4.5-Å cryo-electron microscopy data with density-guided iterative local refinement. Nat. Methods 12, 361–365 (2015).
Hirakawa, H., Ishikawa, S. & Nagamune, T. Design of Ca2+-independent Staphylococcus aureus sortase A mutants. Biotechnol. Bioeng. 109, 2955–2961 (2012).
Meerbrey, K.L. et al. The pINDUCER lentiviral toolkit for inducible RNA interference in vitro and in vivo. Proc. Natl. Acad. Sci. USA 108, 3665–3670 (2011).
Holm, L. & Sander, C. Dali: a network tool for protein structure comparison. Trends Biochem. Sci. 20, 478–480 (1995).
Acknowledgements
The work was supported by US National Institutes of Health (NIH) grants to H.W. (Pioneer Award DP1-HD-087988), to H.L.P. (Pioneer Award DP1-GM-106409), and to Q.Y. (R00: 4R00AI108793-02). F.I.S. was supported by an Advanced Postdoc.Mobility Fellowship from the Swiss National Science Foundation. The cryo-EM facility was funded through the NIH grant AI100645, Center for HIV/AIDS Vaccine Immunology and Immunogen Design (CHAVI-ID). The experiments were performed in part at the Center for Nanoscale Systems at Harvard University, a member of the National Nanotechnology Infrastructure Network (NNIN), which is supported by the US National Science Foundation (NSF) under award no. ECS-0335765. We thank E. Egelman for generous guidance in methods of helical reconstruction.
Author information
Authors and Affiliations
Contributions
A.L., Y.L., F.I.S., Q.Y., S.C., T.-M.F., and Y.M. performed the experiments and analyzed the data. A.L., Q.Y., and T.-M.F. purified the recombinant proteins and performed biochemical experiments. S.C. and Y.M. collected the cryo-EM data, and Y.L. processed the data and completed the helical reconstruction. F.I.S. generated stable cell lines and performed cellular assays, and H.L.P. supervised the experiments. A.B.T. performed Rosetta refinement. A.L. and H.W. conceived the study and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Cryo-EM structure determination of the caspase-1CARD filament.
(a) Enlarged experimental and simulated power spectra showing the position of the meridional layer line. (b-c) A magnified view of the cryo-EM density for helix 1 and helix 4. (d-e) Gold standard and model versus map Fourier shell correlation plots showing the estimated resolution at 4.8 Å. (f) Summary of MolProbity validation report for the caspase-1CARD filament model. (g) Superposition of the caspase-1CARD structure in the filament with the NMR structure of ICEBERG and a homology model of INCA based on the ICEBERG structure.
Supplementary Figure 3 Comparison with other helical-DD-fold structures.
(a-c) Overall best alignment of the type I, II, and III dimers in caspase-1CARD filament (cyan, magenta, and green) with those in MAVSCARD filament (gray). (d) Structural alignment of caspase-1CARD and MAVSCARD as performed by the Dali server (Holm, L. & Sander, C.,Trends Biochem. Sci. 20, 478-480, 1995). Buried interfacial residues identified by PISA (Krissinel, E. & Henrick, K., J Mol Biol 372, 774-97, 2007) for each asymmetric dimer types were highlighted to show the similarity in their relative locations. (e) Comparison of relative angular differences in the caspase-1CARD filament (cyan, magenta, and green) and in the MAVSCARD filament, the PIDDosome complex, the Myddosome complex, and the ASCPYD filament. The left subunits (Ib, IIb, and IIIb monomers) are first aligned in the respective dimers, and the rotation angles required to bring the Ia, IIa, and IIIa monomers in superposition are indicated.
Supplementary Figure 4 ICEBERG does not interact with ASCCARD.
(a) ICEBERG did not nucleate ASCCARD polymerization. (b) ICEBERG did not inhibit GFP-ASCCARD nucleated ASCCARD polymerization.
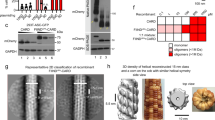
Supplementary Figure 5 Gel-filtration profiles of INCA alone, caspase-1CARD-SUMO alone, and a mixture of the two proteins.
INCA does not interact with monomeric caspase-1CARD. Size-exclusion chromatography was done at concentrations comparable to FP assay for caspase-1CARD labeled with TAMRA fluorophore and its mixture with slightly over-stoichiometric amount of INCA.
Supplementary Figure 6 A model of how INCA caps growing caspase-1CARD oligomer.
Conserved type Ia, IIa, IIIa, and IIIb interfaces allow stochastic incorporation of INCA into growing caspase-1CARD filament. Defective type Ib and IIb interfaces prevent caspase-1 recruitment and filament elongation. C1: caspase-1; I: INCA. The composite electrostatic surface and ribbon diagram at the right is complemented with a top view schematic and a side view helical plot at the right. (a) Normal activation. (b) Caspase-1CARD capped by INCA.
Supplementary Figure 7 Effect of R55E mutation on the inhibition potency of INCA.
This charge-reversal mutation at the type Ia interface greatly reduced INCA's inhibition potency on nucleated caspase-1CARD polymerization. Notably, the Ki is increased to ~1.7 μM.
Supplementary Figure 8 Mechanistic model for the inhibition of inflammasome activation by INCA.
Steps: 1. Activation of sensor proteins such as ALRs (e.g. AIM2) and NLRs (e.g. NLRP3) forms an initial PYD layer. 2. The sensor PYDs nucleate the formation of ASCPYD filaments that further recruit ASC monomers. 3. ASCCARD clusters outside of these PYD filament core. 4. Pro-caspase-1 monomers are recruited by CARD-CARD interaction to form CARD filaments. 5a. During normal activation, caspase-1 filaments elongate by recruitment of pro-caspase-1 monomers. 6. Proximity-driven dimerization and autoprocessing lead to the production of active caspase-1 dimers. 5b. In the presence of INCA, capsase-1 filaments are capped to prevent recruitment of pro-caspase-1 thereby repressing inflammasome activation.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 (PDF 7612 kb)
Supplementary Data Set 1
Original gel images for Figure 2h (PDF 3613 kb)
Rights and permissions
About this article
Cite this article
Lu, A., Li, Y., Schmidt, F. et al. Molecular basis of caspase-1 polymerization and its inhibition by a new capping mechanism. Nat Struct Mol Biol 23, 416–425 (2016). https://doi.org/10.1038/nsmb.3199
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3199
This article is cited by
-
The role of inflammasomes in human diseases and their potential as therapeutic targets
Signal Transduction and Targeted Therapy (2024)
-
Preparation and characterization of monoclonal antibodies against porcine gasdermin D protein
Applied Microbiology and Biotechnology (2024)
-
Oligomeric states of ASC specks regulate inflammatory responses by inflammasome in the extracellular space
Cell Death Discovery (2023)
-
Cryo-EM structures of the active NLRP3 inflammasome disc
Nature (2023)
-
A Review on Caspases: Key Regulators of Biological Activities and Apoptosis
Molecular Neurobiology (2023)