Abstract
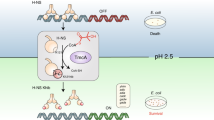
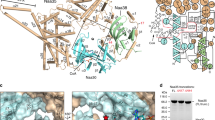
Protein lysine acetylation networks can regulate central processes such as carbon metabolism and gene expression in bacteria. In Escherichia coli, cyclic AMP (cAMP) regulates protein lysine acetyltransferase (PAT) activity at the transcriptional level, but in Mycobacterium tuberculosis, fusion of a cyclic nucleotide-binding domain to a Gcn5-like PAT domain enables direct cAMP control of protein acetylation. Here we describe the allosteric activation mechanism of M. tuberculosis PAT. The crystal structures of the autoinhibited and cAMP-activated PAT reveal that cAMP binds to a cryptic site in the regulatory domain that is over 32 Å from the catalytic site. An extensive conformational rearrangement relieves this autoinhibition by means of a substrate-mimicking lid that covers the protein-substrate binding surface. A steric double latch couples the domains by harnessing a classic, cAMP-mediated conformational switch. The structures suggest general features that enable the evolution of long-range communication between linked domains.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Frankel, A.D. & Kim, P.S. Modular structure of transcription factors: implications for gene regulation. Cell 65, 717–719 (1991).
Dorit, R.L. & Gilbert, W. The limited universe of exons. Curr. Opin. Genet. Dev. 1, 464–469 (1991).
Long, M., de Souza, S.J. & Gilbert, W. Evolution of the intron-exon structure of eukaryotic genes. Curr. Opin. Genet. Dev. 5, 774–778 (1995).
Geer, L.Y., Domrachev, M., Lipman, D.J. & Bryant, S.H. CDART: protein homology by domain architecture. Genome Res. 12, 1619–1623 (2002).
Marcotte, E.M., Pellegrini, M., Thompson, M.J., Yeates, T.O. & Eisenberg, D. A combined algorithm for genome-wide prediction of protein function. Nature 402, 83–86 (1999).
Kuriyan, J. & Eisenberg, D. The origin of protein interactions and allostery in colocalization. Nature 450, 983–990 (2007).
Kuriyan, J. & Cowburn, D. Modular peptide recognition domains in eukaryotic signaling. Annu. Rev. Biophys. Biomol. Struct. 26, 259–288 (1997).
Hu, L.I., Lima, B.P. & Wolfe, A.J. Bacterial protein acetylation: the dawning of a new age. Mol. Microbiol. 77, 15–21 (2010).
Kim, G.W. & Yang, X.J. Comprehensive lysine acetylomes emerging from bacteria to humans. Trends Biochem. Sci. 36, 211–220 (2011).
Schwer, B., Bunkenborg, J., Verdin, R.O., Andersen, J.S. & Verdin, E. Reversible lysine acetylation controls the activity of the mitochondrial enzyme acetyl-CoA synthetase 2. Proc. Natl. Acad. Sci. USA 103, 10224–10229 (2006).
Hirschey, M.D., Shimazu, T., Huang, J.Y., Schwer, B. & Verdin, E. SIRT3 regulates mitochondrial protein acetylation and intermediary metabolism. Cold Spring Harb. Symp. Quant. Biol. 76, 267–277 (2011).
Yu, B.J., Kim, J.A., Moon, J.H., Ryu, S.E. & Pan, J.G. The diversity of lysine-acetylated proteins in Escherichia coli. J. Microbiol. Biotechnol. 18, 1529–1536 (2008).
Wang, Q. et al. Acetylation of metabolic enzymes coordinates carbon source utilization and metabolic flux. Science 327, 1004–1007 (2010).
Zhang, J. et al. Lysine acetylation is a highly abundant and evolutionarily conserved modification in Escherichia coli. Mol. Cell. Proteomics 8, 215–225 (2009).
Thao, S. & Escalante-Semerena, J.C. Control of protein function by reversible Nɛ-lysine acetylation in bacteria. Curr. Opin. Microbiol. 14, 200–204 (2011).
Castaño-Cerezo, S., Bernal, V., Blanco-Catala, J., Iborra, J.L. & Canovas, M. cAMP-CRP co-ordinates the expression of the protein acetylation pathway with central metabolism in Escherichia coli. Mol. Microbiol. 82, 1110–1128 (2011).
Nanchen, A., Schicker, A., Revelles, O. & Sauer, U. Cyclic AMP-dependent catabolite repression is the dominant control mechanism of metabolic fluxes under glucose limitation in Escherichia coli. J. Bacteriol. 190, 2323–2330 (2008).
Nambi, S., Basu, N. & Visweswariah, S.S. cAMP-regulated protein lysine acetylases in mycobacteria. J. Biol. Chem. 285, 24313–24323 (2010).
Xu, H., Hegde, S.S. & Blanchard, J.S. Reversible acetylation and inactivation of Mycobacterium tuberculosis acetyl-CoA synthetase is dependent on cAMP. Biochemistry 50, 5883–5892 (2011).
Nyström, T. & Neidhardt, F.C. Expression and role of the universal stress protein, UspA, of Escherichia coli during growth arrest. Mol. Microbiol. 11, 537–544 (1994).
Crosby, H.A., Pelletier, D.A., Hurst, G.B. & Escalante-Semerena, J.C. System-wide studies of N-lysine acetylation in Rhodopseudomonas palustris reveals substrate specificity of protein acetyltransferases. J. Biol. Chem. 287, 15590–15601 (2012).
Vetting, M.W. et al. Structure and functions of the GNAT superfamily of acetyltransferases. Arch. Biochem. Biophys. 433, 212–226 (2005).
Fraser, J.S. et al. Accessing protein conformational ensembles using room-temperature X-ray crystallography. Proc. Natl. Acad. Sci. USA 108, 16247–16252 (2011).
Lang, P.T. et al. Automated electron-density sampling reveals widespread conformational polymorphism in proteins. Protein Sci. 19, 1420–1431 (2010).
Garrity, J., Gardner, J.G., Hawse, W., Wolberger, C. & Escalante-Semerena, J.C. N-lysine propionylation controls the activity of propionyl-CoA synthetase. J. Biol. Chem. 282, 30239–30245 (2007).
Nambi, S., Badireddy, S., Visweswariah, S.S. & Anand, G.S. Cyclic AMP–induced conformational changes in mycobacterial protein acetyltransferases. J. Biol. Chem. 287, 18115–18129 (2012).
Kornev, A.P., Taylor, S.S. & Ten Eyck, L.F. A generalized allosteric mechanism for cis-regulated cyclic nucleotide binding domains. PLoS Comput. Biol. 4, e1000056 (2008).
Popovych, N., Tzeng, S.R., Tonelli, M., Ebright, R.H. & Kalodimos, C.G. Structural basis for cAMP-mediated allosteric control of the catabolite activator protein. Proc. Natl. Acad. Sci. USA 106, 6927–6932 (2009).
Passner, J.M., Schultz, S.C. & Steitz, T.A. Modeling the cAMP-induced allosteric transition using the crystal structure of CAP-cAMP at 2.1 Å resolution. J. Mol. Biol. 304, 847–859 (2000).
Sharma, H., Yu, S., Kong, J., Wang, J. & Steitz, T.A. Structure of apo-CAP reveals that large conformational changes are necessary for DNA binding. Proc. Natl. Acad. Sci. USA 106, 16604–16609 (2009).
Kim, C., Xuong, N.H. & Taylor, S.S. Crystal structure of a complex between the catalytic and regulatory (RIα) subunits of PKA. Science 307, 690–696 (2005).
Berman, H.M. et al. The cAMP binding domain: an ancient signaling module. Proc. Natl. Acad. Sci. USA 102, 45–50 (2005).
Shenoy, A.R. & Visweswariah, S.S. New messages from old messengers: cAMP and mycobacteria. Trends Microbiol. 14, 543–550 (2006).
Bai, G., Knapp, G.S. & McDonough, K.A. Cyclic AMP signalling in mycobacteria: redirecting the conversation with a common currency. Cell Microbiol. 13, 349–358 (2011).
Muñoz-Elias, E.J. et al. Replication dynamics of Mycobacterium tuberculosis in chronically infected mice. Infect. Immun. 73, 546–551 (2005).
Gill, W.P. et al. A replication clock for Mycobacterium tuberculosis. Nat. Med. 15, 211–214 (2009).
Wayne, L.G. Synchronized replication of Mycobacterium tuberculosis. Infect. Immun. 17, 528–530 (1977).
Brent, M.M., Iwata, A., Carten, J., Zhao, K. & Marmorstein, R. Structure and biochemical characterization of protein acetyltransferase from Sulfolobus solfataricus. J. Biol. Chem. 284, 19412–19419 (2009).
Van Duyne, G.D., Standaert, R.F., Karplus, P.A., Schreiber, S.L. & Clardy, J. Atomic structures of the human immunophilin FKBP-12 complexes with FK506 and rapamycin. J. Mol. Biol. 229, 105–124 (1993).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Leslie, A.G.W. Recent changes to the MOSFLM package for processing film and image plate data. Joint CCP4 ESF-EAMCB Newslett. Protein Crystallogr. 26 (1992).
Collaborative Computational Project, No. 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Langer, G., Cohen, S.X., Lamzin, V.S. & Perrakis, A. Automated macromolecular model building for X-ray crystallography using ARP/wARP version 7. Nat. Protoc. 3, 1171–1179 (2008).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Murshudov, G.N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D Biol. Crystallogr. 67, 355–367 (2011).
Cowtan, K. The Buccaneer software for automated model building. 1. Tracing protein chains. Acta Crystallogr. D Biol. Crystallogr. 62, 1002–1011 (2006).
Chen, V.B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Larkin, M.A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
Kabsch, W. & Sander, C. Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22, 2577–2637 (1983).
Pettersen, E.F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Acknowledgements
We thank J. Holton, G. Meigs and J. Tanamachi at Beamline 8.3.1 at Lawrence Berkeley National Laboratory for help with X-ray data collection. We appreciate the support of the TB Structural Genomics Consortium. This work was supported by a postdoctoral research fellowship from the Canadian Institutes of Health Research to H.J.L. and US National Institutes of Health grants R01GM70962 and P01AI068135 to T.A.
Author information
Authors and Affiliations
Contributions
H.J.L. conducted all the biochemical and crystallographic studies. P.T.L. conducted the computational studies with Ringer. T.A., H.J.L., P.T.L. and C.M.S. wrote the manuscript. All authors designed analyses, discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 (PDF 4404 kb)
Rights and permissions
About this article
Cite this article
Lee, H., Lang, P., Fortune, S. et al. Cyclic AMP regulation of protein lysine acetylation in Mycobacterium tuberculosis. Nat Struct Mol Biol 19, 811–818 (2012). https://doi.org/10.1038/nsmb.2318
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2318
This article is cited by
-
Negative regulation of the acsA1 gene encoding the major acetyl-CoA synthetase by cAMP receptor protein in Mycobacterium smegmatis
Journal of Microbiology (2022)
-
Lysine acetylation of DosR regulates the hypoxia response of Mycobacterium tuberculosis
Emerging Microbes & Infections (2018)
-
Novel protein acetyltransferase, Rv2170, modulates carbon and energy metabolism in Mycobacterium tuberculosis
Scientific Reports (2017)