Abstract
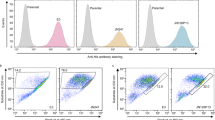
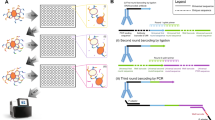
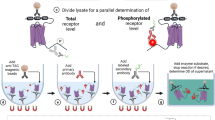
The B-cell lymphoma-2 (Bcl-2) family contains six antiapoptotic members, each with a hydrophobic pocket in which Bcl-2 homology region 3 (BH3) helices bind. This binding quenches apoptotic signals from activated BH3 family members. Many tumor cells either have increased expression of one of these six proteins or become overexpressed under treatment. Six fusion proteins made up of glutathione-S-transferase and each of the Bcl-2 members are bound individually to six glutathione bead sets, each set being easily distinguished by its different intensity of red fluorescence. The coated bead sets are washed, combined and incubated with green fluorescent Bim-BH3 peptide and a small molecule in 10-μl wells for 1 h. The green fluorescence signal for each bead set is resolved, and selective inhibitors are expected to reduce the signal for individual bead sets. Each 384-well plate is analyzed in 12 min, measuring 200 of 2,000 beads (∼10%) of each type per well.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Gad, S.C. Drug Discovery Handbook (John Wiley and Sons, 2005).
Edwards, B.S. et al. Biomolecular screening of formylpeptide receptor ligands with a sensitive, quantitative, high-throughput flow cytometry platform. Nat. Protoc. 1, 59–66 (2006).
Tsujimoto, Y., Cossman, J., Jaffe, E. & Croce, C.M. Involvement of the bcl-2 gene in human follicular lymphoma. Science 228, 1440–1443 (1985).
Bakhshi, A. et al. Cloning the chromosomal breakpoint of t(14;18) human lymphomas: clustering around JH on chromosome 14 and near a transcriptional unit on 18. Cell 41, 899–906 (1985).
Cleary, M.L., Smith, S.D. & Sklar, J. Cloning and structural analysis of cDNAs for bcl-2 and a hybrid bcl-2/immunoglobulin transcript resulting from the t(14;18) translocation. Cell 47, 19–28 (1986).
Youle, R.J. & Strasser, A. The BCL-2 protein family: opposing activities that mediate cell death. Nat. Rev. Mol. Cell Biol. 9, 47–59 (2008).
Sattler, M. et al. Structure of Bcl-xL-Bak peptide complex: recognition between regulators of apoptosis. Science 275, 983–986 (1997).
Chen, L. et al. Differential targeting of prosurvival Bcl-2 proteins by their BH3-only ligands allows complementary apoptotic function. Mol. Cell 17, 393–403 (2005).
Adams, J.M. & Cory, S. The Bcl-2 apoptotic switch in cancer development and therapy. Oncogene 26, 1324–1337 (2007).
Reed, J.C. & Pellecchia, M. Apoptosis-based therapies for hematologic malignancies. Blood 106, 408–418 (2005).
Zhai, D., Jin, C., Satterthwait, A.C. & Reed, J.C. Comparison of chemical inhibitors of antiapoptotic Bcl-2-family proteins. Cell Death. Differ. 13, 1419–1421 (2006).
Nakamura, K. et al. Homogeneous time-resolved fluorescence resonance energy transfer assay for measurement of Phox/Bem1p (PB1) domain heterodimerization. J. Biomol. Screen. 13, 396–405 (2008).
Owicki, J.C. Fluorescence polarization and anisotropy in high throughput screening: perspectives and primer. J. Biomol. Screen. 5, 297–306 (2000).
Carson, R.T. & Vignali, D.A. Simultaneous quantitation of 15 cytokines using a multiplexed flow cytometric assay. J. Immunol. Methods 227, 41–52 (1999).
Curpan, R.F. et al. High-throughput screen for chemical inhibitors of anti-apoptotic Bcl-2 family proteins by multiples flow cytometry. Assay Drug Dev. Technol. (2011).
Simons, P.C. et al. Duplexed, bead-based competitive assay for inhibitors of protein kinases. Cytometry A 71, 451–459 (2007).
Tessema, M. et al. Glutathione-S-transferase-green fluorescent protein fusion protein reveals slow dissociation from high site density beads and measures free GSH. Cytometry A 69, 326–334 (2006).
Bartsch, J.W. et al. An investigation of liquid carryover and sample residual for a high-throughput flow cytometer sample delivery system. Anal. Chem. 76, 3810–3817 (2004).
Acknowledgements
This work was supported by NIH grants 1X01 MHO79850-01, CA113318 and U54 MH074425/MH084690, the New Mexico Molecular Libraries Screening Center/University of New Mexico Center for Molecular Discovery (NMMLSC/UNMCMD), and the University of New Mexico Shared Flow Cytometry Resource and Cancer Research and Treatment Center (P30 CA118100). We acknowledge the contributions of R. Halip, C. Bologa and T. Oprea to the analysis of primary and dose-response screens.
Author information
Authors and Affiliations
Contributions
P.C.S. devised the method of GSH bead synthesis and assay development and prepared the manuscript; S.M.Y. contributed to assay development and implementation; M.B.C. was involved in instrument programming and maintenance; A.W. developed and analyzed the spreadsheets for screening and dose response; D.Z. was involved in fusion-protein creation and purification; J.C.R. was the leader of the Burnham team investigating Bcl-2 family and helped in conceptualizing the assay; B.S.E. was the leader of NMMLSC/UNMCMD high-throughput screening core and also a co-developer of HyperCyt platform; L.A.S. was the principal investigator of NMMLSC/UNMCMD, a co-developer of the HyperCyt platform, and contributed to target assay conceptualization and manuscript preparation.
Corresponding author
Ethics declarations
Competing interests
B.S.E. and L.A.S. declare competing financial interests as co-inventors of HyperCyt and co-founders of IntelliCyt Corporation.
Rights and permissions
About this article
Cite this article
Simons, P., Young, S., Carter, M. et al. Simultaneous in vitro molecular screening of protein-peptide interactions by flow cytometry, using six Bcl-2 family proteins as examples. Nat Protoc 6, 943–952 (2011). https://doi.org/10.1038/nprot.2011.339
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2011.339
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.