Abstract
The fusion peptide of influenza virus hemagglutinin (HA) has a critical role in mediating the entry of the virus into the cells and is also the only universally conserved sequence in the HAs of all strains of influenza A and influenza B viruses. Therefore, it could be an attractive target for new vaccine development and a potency marker for existing influenza vaccines. The fusion peptide epitope is hidden inside the HA proteins, making it inaccessible for quantitative antibody binding. Our simple slot blot protocol highlights pre-treatment of HA samples with moderate concentrations of denaturant to maximally expose the fusion peptide on the protein surface, followed by detection using universal antibodies targeting the fusion peptide. The method is highly reliable, inexpensive and easy to follow. The entire procedure takes only 5 h and can be applied to the quantitative determination of virtually all influenza viral HAs using a single antibody targeting the fusion peptide.
Similar content being viewed by others
Introduction
Hemagglutinin (HA) is the principle surface protein of influenza virus that induces neutralizing antibodies1. There are 16 subtypes of influenza A viral HAs2, but in the same subtype of HAs, the amino acid sequences can vary widely3,4. Currently, there are thousands of HA sequences submitted in the public database and the number of sequence is constantly increasing3. The ever-increasing number of HA sequences reflects the rapidly evolving nature of the virus, making it a daunting challenge for vaccine development. Indeed, the seasonal influenza vaccines have to be annually updated4,5,6,7,8. Specifically, the World Health Organization (WHO) recommends the updating of strains of the seasonal influenza vaccines about 6–8 months before the upcoming flu season. This approach has inherent disadvantages, including the uncertainty about the actual circulating strain, the need for annual updating of the manufacturing process in addition to the preparation of reagents for vaccine lot release4,5,6,7,8. Obviously, these shortcomings would be significantly exacerbated in the event of a pandemic influenza outbreak. Such problems associated with influenza vaccines are largely due to frequent mutations in the genes for the viral surface proteins (HA and neuraminidase). As HA is the major surface protein that induces protective immune responses, it is measured as a vaccine potency marker by single radial immunodiffusion (SRID) method9.
The current SRID method depends on the availability of HA antigen references and the corresponding antiserum for the standardization of vaccine quantification. These reagents are updated and distributed annually by the WHO collaborating centers. Owing to the fact that at least 2–3 months are required to make these reagents from the time when a new strain is identified, the WHO recommends that the manufacturers develop alternative methods to quantify the HAs in the vaccine preparations in the event of a pandemic outbreak10. A reliable alternative method could allow faster release of the vaccine for mass immunization, when the time is of essence to save numerous human lives11,12. It is also important to note that the rapidly evolving nature of the influenza virus will continue to pose tremendous challenge to public health and scientific community12. Clearly, the rapid development of vaccines against emerging influenza viruses and facilitating faster release of vaccines is of unquestionable importance.
Comparison of various methods for HA quantification
SRID, albeit rather tedious and time consuming, is able to determine the antigenic conformation and quantity of HA proteins in vaccine preparations9. The assay has been successfully used for over 20 years to quantify HA in influenza vaccines. However, it requires subtype-specific antisera, which must be generated annually. Therefore, alternative assays for the quantification of influenza viral HAs aimed at shortening the time required to produce vaccines could facilitate more rapid release of vaccines for human immunization, which would be urgently required in the event of a pandemic influenza outbreak.
Two categories of alternative assays, i.e., physicochemical and immunological assays, have been described recently. The physicochemical assays are mainly based on chromatographic separation alone or in conjunction with mass spectrometry13,14,15,16. These assays have shown good repeatability and accuracy in quantifying HA proteins in vaccine preparation. It should be noted that ultra-performance liquid chromatography is used in combination with isotope dilution tandem mass spectrometry; the assay is quite fast and allows analysis of more than 100 samples per day15,16. In addition, selected peptides representing the conserved sequences of several subtypes are used as standards for vaccine quantification15,16. Yet challenges remain, including the lack of methods for determining their three-dimensional structure and analyzing elusive components of the vaccines; the necessity to develop subtype-specific protein standards; high equipment costs; the need for highly trained staff; and the demand for high through-put3,13,14,15,16. Although several groups have reported physicochemical methods, alternative immunoassay for HA quantification is much less documented, largely because of the constantly changing nature of the HAs. Indeed, there are some 6,000 of HA sequences reported in public domain (http://www.ncbi.nlm.nih.gov/genomes/FLU/FLU.html).
The immunoassay based on universal antibody targeting the fusion peptide of virally all influenza viral HAs
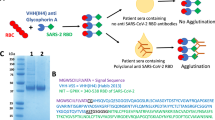
Universal antibodies against the most conserved sequences of all influenza viral HAs. We recently reported the development of mono-specific antibodies exclusively binding to the fusion peptide, the only universally conserved regions of all HAs3. The mono-specific antibodies were raised using fusion peptide (the amino terminal 1–14 amino acid of the HA2 subunit) chemically modified and then conjugated to keyhole limpet haemocyanin (KLH), followed by injection into the rabbits3. The antibodies showed remarkable specificity against the viral sequences (the fusion peptide) as no cross-reactivity to the proteins derived from allantoic fluids could be detected. We also proved that the universal antibodies can bind to HA from 13 different subtypes (H1–H13) of influenza A virus and virus from type B available at our institutions. It is interesting to note that the fusion peptide region targeted by the universal antibodies was found to have only two significant amino acid substitutions based on bioinformatics analyses of all the publicly available sequences (regardless of the hosts from which the viruses were isolated). Importantly, the two substitutions were later found not to affect the binding between the HA and the antibodies3. As this sequence is universally conserved, we conclude that all HAs identified so far should be recognized by the antibodies even though, understandably, it is impossible to actually test all 6,000 HA sequences reported in the public domain3.
Advantages and limitations of universal antibody-based assay. Given that the fusion peptide has a critical role in viral life cycle by mediating the fusion between the virus and host cells1,17 and that antibodies targeting the fusion peptide or part of this fusion peptide could protect the animals from challenge with a broad spectrum of influenza virus strains18,19,20, the application of a universal antibody targeting this region for practical issue (vaccine quantification, especially when the subtype/strain-specific anti-serum is not available) or fundamental research (receptor binding and conformational epitope of HA) would be of interest. Moreover, the antibody can bind to essentially all influenza type A and B viral HAs, and there is no need to generate antibodies against every single subtype of viral HAs. The assay is simple, reproducible and inexpensive.
There are some limitations of the universal antibody-based immunoassay or the antibody itself, i.e., it is not designed to distinguish the different strains of viral HAs. As such, it cannot be used for the identification of the strain, for which SRID using strain-specific antisera should be used. In addition, pre-treatment of the HA-containing samples with moderate concentration of denaturant (4 M urea) results in partial unfolding of the protein, exposing the fusion peptide for more effective binding to the antibodies. Interestingly, destabilizing the HA proteins with 4 M urea resulted in a conformational change of the HA to the fusogenic state indistinguishable from that induced by low pH17. Therefore, interested investigators might wish to optimize experimental conditions in their studies when using antibodies against the fusion peptide in various forms of vaccine preparations as only recombinant HA and seasonal influenza vaccines have been tested in our work. The conditions for the testing of other forms of vaccine preparations (pseudotyped virus, virus-like particle or recombinant adenovirus expressing HA) have not yet been studied. Furthermore, it is noteworthy that there is no standardized SRID method for the quantification of these forms of vaccines either.
In short, all newly reported methods (physicochemical or immunological) have their own strength and weakness compared with the traditional SRID method. One of the most striking difference is that the alternative methods may allow rapid quantification of HA proteins, thus reducing the time needed for the production and release of the vaccine, but they are unable to detect the 'antigenic' conformation and/or determine the subtype specificity3,13,14,15,16.
The availability of the universal antibodies. Interested investigators could generate their own antibodies by following the procedure described previously3 or they may contact the authors for the antibodies to be used in their exploratory research. Currently, the supplies of the mono-specific antibodies to research community are limited as the monoclonal antibodies are still being characterized. Preliminary results indicate that the monoclonal antibody can be used to detect the fusion protein. The monoclonal antibodies will be made available for the research community after the full characterization has been completed.
In this report, we detail the protocols for the determination of influenza virus HAs and provide troubleshooting advice for commonly encountered problems.
Experimental design
Samples: two types of samples have been evaluated in our studies.
-
1
Influenza virus or vaccine samples: the annual human vaccines are obtainable in most countries. Currently, influenza viruses used for vaccines are mainly produced from allantoic fluids derived from seed virus-inoculated embryonated eggs or cell cultures based on the classical gradient centrifugation21. The vaccine samples are usually supplied as liquid solutions and should be stored at 4 °C. The vaccines often appear opaque and may sediment over time even within expiry date. Unless instructed by the vaccine suppliers, storage of influenza vaccines by direct freezing the samples at −20° or −70 °C is not recommended as it could cause aggregation of the samples. Researchers interested in laboratory studies and not human vaccine manufacture could also prepare the vaccine-like samples by themselves by purifying the virus from egg or cell cultures using the previously reported procedure21.
-
2
Recombinant HA proteins: Recombinant HA (rHA) proteins are being studied in clinical trials now3,4,5,6,7. rHA produced in insect expression systems could be used in this protocol. The rHA in liquid form needs to be stored at 4 °C and should normally appear clear.
Increased sensitivity by exposing the fusion peptide. The fusion peptide is the only universally conserved sequence in all strains of the influenza virus HAs. It is only partially exposed on the surface of the HA protein in solution at neutral pH (refs. 1,3,17). Exposure of the HAs to acidic condition or moderate concentration of denaturant (i.e., 4 M urea) resulted in irreversible conformation changes in the viral HA, accompanied by exposure of the fusion peptide at the surface of the proteins3,17. To improve the sensitivity and accuracy of the assay, it is necessary to pre-treat the samples before the immunoassay using 4 M urea. This step has two specific purposes: (1) maximize the exposure of the fusion peptide; (2) ensure the solubilization of any precipitates of viral proteins in human vaccine preparation, which is usually opaque and can sediment over storage time, thus interfering with the immunoassay.
Antigen quantification. It should be noted that the amounts of antigens on the membrane can also be directly determined without the use of X-ray film exposure. In this case, after adding enhanced chemiluminescent (ECL) substrate at Step 17, analyze the quantity of the HA using the ChemiDoc-It 410 Imaging System. Alternatively, the investigator could also chose to use fluorescently labeled secondary antibodies rather than an enzymatic reaction; the Odyssey System can then be used to quantify the HA contents. It seems that the fluorescently labeled antibody/Odyssey imager offers a broad linear dynamic range to accurately detect strong and weak signals on the same blot, please refer to http://www.licor.com.
Linear regression versus logistic regression. In addition to sample pre-treatment, using appropriate calibration curve fitting in immunoassays is crucial for quantifying HA accurately3. Linear regression is the simplest method for the determination of the analytes by plotting the concentration (x) versus the response (y) using the linear portion of the response curve (y = mx + b). If a broader range of analytes is considered for concentration calculation, the four parameter logistic (4PL) for immunoassay currently accepted can be used3. A variety of software is available. We routinely use SigmaPlot (Systat Software, Chicago, Illinois, USA).
Materials
Reagents
-
Tris buffered saline (TBS) buffer (Invitrogen, cat. no. R017R.0000)
-
Urea (Sigma, cat. no. U1250)
-
Tween-20 (Sigma, cat. no. P7949) or equivalent
-
Skim milk powder, Blotting Grade Non-Fat Dry Milk (Bio-Rad cat. no. 1706404)
-
Mono-specific rabbit antibody against the fusion peptide of influenza virus HA4. (The antibodies are available upon request, see above)
-
Sheep-anti-Rabbit IgG horseradish peroxidase conjugate (Amersham, cat. no. RPN 2108).
-
ECL substrate, Amersham ECL Western Blotting Analysis system (Amersham/GE health care, cat. no. RPN 2108)
-
Viral vaccine samples (A/Solomon Islands/3/2006) as described3. Seasonable influenza vaccines are commonly available in most countries.
-
Positive controls for subtype-specific vaccines and their corresponding antiserum could be obtained from the National Institute for Biological Standards and Control (http://www.nibsc.ac.uk). The controls should be treated the same way as the samples would be (see below for details).
Equipment
-
Bio-Dot SF Microfiltration Apparatus (Bio-Rad, cat. no. 170-6542)
-
Immobilon-FL Transfer Membrane (Polyvinylidene Fluoride, PVDF), Millipore (methanol activate) (cat. no. IPFL 00010). Note: we routinely use PVDF; nitrocellulose or nylon membrane could also be used but we did not compare them in terms of immobilizing efficiency for this study.
-
Filter papers (Bio-Rad cat. no. 1620161)
-
Vacuum pump (Millipore, model # WP 6111560)
-
X-ray film (BioMax Light Film) (Kodak, cat. no. 178 8207)
-
FluorChem Gel Documentation System (Alpha Innotech, cat. no. 29-008-1896X)
-
ChemiDoc-It 410 Imaging System (UVP, LLC, cat. no. ChemiDoc-It 410)
-
Various types of pipettes suitable for 1–10, 10–50, 50–100, 100–200, 200–1,000 μl (Fisherbrand EX)
-
Filter forceps (for safe handling of filter membranes, Fisher types or equivalent)
-
Latex gloves (Kimberly-Clark, Latex or Nitrile gloves or equivalent)
-
Sterile flasks for solution preparations (VWR or equivalent)
-
Stovall Belly Dancer Shaker (Stovall Life Science cat. no. BDRAA 115S or equivalent)
-
Kleenex facial tissue or equivalent
Reagent setup
Blocking buffer
-
We add 5 g of skim milk powder to 100 ml of TBS buffer and dissolve the powder by gently shaking the flask. The blocking buffer should be stored at 4 °C and discarded after one week.
Washing buffer
-
We use 1× TBS containing 0.1% Tween-20. Use the buffer for each washing step (see below in the Procedure) and also to dilute primary and secondary antibodies at final concentration of 1/2,000 and 1:10,000, respectively. Store the buffer at room temperature (20–25 °C) for at least 3 months. Periodically inspect the solution for any opaqueness or the appearance of a precipitate.
8 M Urea in PBS
-
Dissolve 480.48 g urea in a final volume of 1,000 ml of phosphate buffered saline (PBS). The solution can be stored for up to 1 week at room temperature.
Preparation of PVDF and filter paper
-
Pre-treat the PVDF membrane by cutting the PVDF membrane to a 9 cm × 12 cm size and activating the membrane for 15 s in methanol. Rinse the membrane in double distilled water (ddH2O) and remove the excess of ddH2O using Kleenex facial tissue. Also pre-wet three sheets of Bio-Dot filter paper in TBS.
Procedure
pre-treatment of samples:
Timing 45 min
-
1
Pre-treatment of HA samples: treat the HA samples (human vaccine) by combining equal volume of samples with equal volume of 8 M urea. Leave the mixture at room temperature for 30 min before it is used for subsequent analyses.
Critical Step
Pre-treatment of the samples with 4 M urea exposes the fusion peptide of the HA and enhances the binding of the antibodies to the fusion peptide3.
setup of apparatus and pre-treatment of membrane:
Timing 10 min
-
2
Assemble the Bio-Dot SF apparatus with the pre-wet filter paper and membrane (see Reagent Setup) using forceps, according to the manufacturer's instructions; tighten the screws.
-
3
Turn on the vacuum and set the three-way valve to vacuum position.
-
4
Re-tighten the screw to prevent any leakage due to the vacuum application.
Critical Step
Re-tightening the screw is important as vacuum application can loosen the apparatus, resulting in leakage of samples.
-
5
Change the flow valve to the atmospheric position; apply 100 μl TBS to each well to re-hydrate the membrane.
-
6
Change the flow to the gentle vacuum (apply vacuum by putting finger over atmospheric valve).
-
7
After solution has drained change to atmospheric pressure, disconnect vacuum.
sample applications:
Timing 10–15 min
-
8
Apply sample from Step 1 (at a volume of 200 μl); check for air bubbles; remove bubbles by pipetting up and down. The un-used wells are filled with 200 μl of TBS to facilitate the vacuum suction.
Critical Step
Air bubbles must be removed by gently pipetting the samples up and down a few times.
-
9
Apply gentle vacuum.
-
10
After samples are drained, remove the membrane.
Pause point
For long-term storage, the membranes can be air-dried, sealed in Saran Wrap and stored at room temperature for future use.
-
11
After removing the membrane, place it in a plastic tray containing the blocking solution (5% fat-free milk in TBS); gently shake the tray on a platform shaker for 1 h.
blocking of the membrane:
Timing 60 min for blocking; 10 min for washing
-
12
Remove the membrane from the blocking solution and wash twice for 5 min in TBS or 0.1% Tween-20.
binding of the primary antibodies:
Timing 2 h for primary antibody incubation; 10 min for washing
-
13
Incubate the membrane with 25 ml of primary antibody (anti-HA 1:2,000) at room temperature for 2 h in a plastic tray (10 cm × 11.5 cm).
binding of the secondary antibodies:
Timing 1 h for 2nd antibody incubation; 15 min for washing
-
14
Wash the membrane twice for 5 min to remove the unbound first antibody using TBS or Tween-20 in the plastic tray.
-
15
Incubate the membrane with 25 ml of secondary antibody (sheep anti-rabbit IgG conjugated with horseradish peroxidase at 1:10,000 dilution) at room temperature for 1 h on shaker with gentle shaking.
-
16
Wash as in Step 14; followed by another wash for 5 min using TBS to remove Tween-20.
development of reaction and data acquisition:
Timing 20 min
-
17
Add 7 ml of ECL substrate to membrane (or sufficient to cover the whole membrane) and allow the enzymatic reaction to proceed for 1 min, followed by removal of excess ECL substrate by gently blotting the membrane in tissue paper.
-
18
Expose the membrane to X-ray film in the dark at room temperature. (It should be noted that depending on the amounts of the antigens on the blots, the exposure time varies. In general, 10 s is used to start with, whereas exposure beyond 10 min will not increase the signals.)
-
19
Quantify the amount of HA using the Alpha Innotech Imaging system by placing the developed film on the detection screen of the instrument and carry out analyses of densitometry following the user's manual.
calculation of HA concentrations:
Timing 20 min
-
20
Calculation of HA concentration from standard curve: calculate the HA concentration in the vaccine using linear regression for the linear portion of the plot of the concentration versus the response using the linear portion of the curve. Should a wider range be desired, use the 4PL regression for accurate determination of the HA concentration in the sample.
Troubleshooting
Troubleshooting advice can be found in Table 1.
Timing
Step 1, pre-treatment of samples: 45 min
Steps 2–7, setup of apparatus and pre-treatment of membrane: 10 min
Steps 8–10, sample applications: 10–15 min
Steps 11 and 12, blocking of the membrane: 60 min for blocking; 10 min for washing
Steps 13 and 14, binding of the primary antibodies: 2 h for primary antibody incubation; 10 min for washing
Steps 15 and 16, binding of the secondary antibodies: 1 h for 2nd antibody incubation; 15 min for washing
Steps 17–19, development of reaction and data acquisition: 20 min
Step 20, calculation of HA concentrations: 20 min
Anticipated results
Reproducible results can be obtained by following this protocol.
As shown in Figure 1a, the intensities of the blot decrease proportionally with decreasing concentrations of the vaccine without any nonspecific binding being detected in the negative control in which normal rabbit serum control was used in place of the antibodies.
(a) The HA samples at various concentrations were analyzed using the protocol detailed in the text. The pre-bleed sample served as negative control. Similar results were obtained using affinity-purified mono-specific antibodies3. (b) The figure shows that the concentration of the antigen in solution (competing antigen), plotted on x axis, is inversely proportional to the O.D. values (y axis). The 4-parameter logistic (4PL) model was used for the calibration curve fitting in the competitive ELISA. The curve demonstrates a very good fitting as revealed by R = 0.9975.
Figure 1b shows that the concentrations of the HA antigens were proportionally corresponding to the densitometry values. It is noted that linear dose response was observed at lower concentrations (6.5–200 ng), whereas a higher concentration range (up to 600 ng) can be calculated using the widely accepted 4PL model for immunoassay, with a good fitting of the curve (R = 0.976).
References
Skehel, J.J. & Wiley, D.C. Receptor binding and membrane fusion in virus entry: the influenza HA. Annu. Rev. Biochem. 69, 531–569 (2000).
Fouchier, R.A., Munster, V., Wallensten, A., Bestebroer, T.M., Herfst, S. & Smith, D. Characterization of a novel influenza A virus hemagglutinin subtype (H16) obtained from black-headed gulls. J. Virol. 79, 2814–2222 (2005).
Chun, S. et al. Universal antibodies and their applications to the quantitative determination of virtually all subtypes of the influenza A viral hemagglutinins. Vaccine 26, 6068–6076 (2008).
Poland, G.A., Rottinghaus, S.T. & Jacobson, R.M. Influenza vaccines: a review and rationale for use in developed and underdeveloped countries. Vaccine 19, 2216–2220 (2001).
Simonsen, L., Taylor, R.J., Viboud, C., Miller, M.A. & Jackson, L.A. Mortality benefits of influenza vaccination in elderly people: an ongoing controversy. Lancet Infect. Dis. 7, 658–666 (2007).
Carrat, F. & Flahault, A. Influenza vaccine: the challenge of antigenic drift. Vaccine 25, 6852–6862 (2007).
Bridges, C.B. et al. Effectiveness and cost–benefit of influenza vaccination of healthy working adults: a randomized controlled trial. JAMA 284, 1655–1663 (2000).
De Filette, M. et al. Universal influenza A vaccine: optimization of M2-based constructs. Virology 337, 149–161 (2005).
Wood, J.M. et al. The influence of the host cell on standardisation of influenza vaccine potency. Dev. Biol. Stand. 98, 183–188 (1999).
World Health Organization. WHO Guidelines on the Use of Vaccines and Antivirals during Influenza Pandemics WHO/CDS/CSR/RMD/2004.8. (World Health Organization, Department of Communicable Disease Surveillance and Response, Geneva, Switzerland, 2004).
Belshe, R.B. Implications of the emergence of a novel H1 influenza virus. N. Engl. J. Med. 360, 2667–2668 (2009).
Neumann, G., Noda, T. & Kawaoka, Y. Emergence and pandemic potential of swine-origin H1N1 influenza virus. Nature 459, 931–939 (2009).
Kapteyn, J.C. et al. HPLC-based quantification of haemagglutinin in the production of egg- and MDCK cell-derived influenza virus seasonal and pandemic vaccines. Vaccine. 27, 1468–1477 (2009).
García-Cañas, V., Lorbetskie, B., Bertrand, D., Cyr, T.D. & Girard, M. Selective and quantitative detection of influenza virus proteins in commercial vaccines using two-dimensional high-performance liquid chromatography and fluorescence detection. Anal. Chem. 79, 3164–3172 (2007).
Williams, T.L. et al. Quantification of influenza virus hemagglutinins in complex mixtures using isotope dilution tandem mass spectrometry. Vaccine 26, 2510–2520 (2008).
Luna, L.G., Williams, T.L., Pirkle, J.L. & Barr, J.R. Ultra performance liquid chromatography isotope dilution tandem mass spectrometry for the absolute quantification of proteins and peptides. Anal. Chem. 80, 2688–2693 (2008).
Carr, C.M., Chaudhry, C. & Kim, P.S. Influenza hemagglutinin is spring-loaded by a metastable native conformation. Proc. Natl. Acad. Sci. USA 94, 14306–14313 (1997).
Prabhu, N. et al. Monoclonal antibodies against the fusion peptide of hemagglutinin protect mice from lethal influenza A virus H5N1 infection. J. Virol. 83, 2553–2562 (2009).
Sui, J. et al. Structural and functional bases for broad-spectrum neutralization of avian and human influenza A viruses. Nat. Struct. Mol. Biol. 16, 265–273 (2009).
Ekiert, D.C. et al. Antibody recognition of a highly conserved influenza virus epitope. Science 324, 246–251 (2009).
Reimer, C.B., Baker, R.S., Van Frank, R.M., Newlin, T.E., Cline, G.B. & Anderson, N.G. Purification of large quantities of influenza virus by density gradient centrifugation. J. Virol. 1, 1207–1216 (1967).
Acknowledgements
We thank Dr. Michel Girard, Dr. Aline Rinfret, Helen Macdonald-Piquard (Health Canada) for helpful discussions. We also appreciate critical reviews of the papers by Dr. Michael Rosu-Myles, Dr. Shiv Prasad and editorial review by Mrs. Monika Tocchi (Health Canada). We also thank Alex Bliu (Health Canada) for statistical analyses. This work was supported by the Canadian Regulatory Strategy for Biotechnology and by Pandemic Preparedness Funds of Health Canada (to X.L., R.H., T.D.C.), Ministry of Science and Technology, P.R. China (2004BA519A69 to J.W., C.L.), and Public Health Agency of Canada Research Funds (to R.H.).
Author information
Authors and Affiliations
Contributions
C.L., B.J., J.W., T.D.C., R.H. and X.L. designed the experiments, interpreted results and prepared the paper. C.L., B.J., G.V.D. and Y.S. conducted the experiments.
Corresponding author
Rights and permissions
About this article
Cite this article
Li, C., Jaentschke, B., Song, Y. et al. A simple slot blot for the detection of virtually all subtypes of the influenza A viral hemagglutinins using universal antibodies targeting the fusion peptide. Nat Protoc 5, 14–19 (2010). https://doi.org/10.1038/nprot.2009.200
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2009.200
This article is cited by
-
Hemagglutinin Quantitative ELISA-based Potency Assay for Trivalent Seasonal Influenza Vaccine Using Group-Specific Universal Monoclonal Antibodies
Scientific Reports (2019)
-
VaxArray potency assay for rapid assessment of “pandemic” influenza vaccines
npj Vaccines (2018)
-
Universal type/subtype-specific antibodies for quantitative analyses of neuraminidase in trivalent influenza vaccines
Scientific Reports (2018)
-
Regulatory science accelerates the development of biotechnology drugs and vaccines by NIFDC
Emerging Microbes & Infections (2014)
-
Analytical technologies for influenza virus-like particle candidate vaccines: challenges and emerging approaches
Virology Journal (2013)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.