Abstract
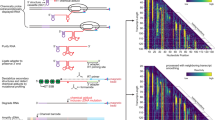
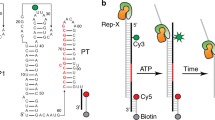
RNA selective 2′-hydroxyl acylation analyzed by primer extension (SHAPE) chemistry exploits the discovery that conformationally dynamic nucleotides preferentially adopt configurations that facilitate reaction between the 2′-OH group and a hydroxyl-selective electrophile, such as benzoyl cyanide (BzCN), to form a 2′-O-adduct. BzCN is ideally suited for quantitative, time-resolved analysis of RNA folding and ribonucleoprotein (RNP) assembly mechanisms because this reagent both reacts with flexible RNA nucleotides and also undergoes auto-inactivating hydrolysis with a half-life of 0.25 s at 37 °C. RNA folding is initiated by addition of Mg2+ or protein, or other change in solution conditions, and nucleotide resolution structural images are obtained by adding aliquots of the evolving reaction to BzCN and then 'waiting' for 1 second. Sites of the 2′-O-adduct formation are subsequently scored as stops to primer extension using reverse transcriptase. This time-resolved SHAPE protocol makes it possible to obtain 1-second structural snapshots in time-resolved kinetic studies for RNAs of arbitrary length and complexity in a straightforward and concise experiment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Tinoco, I. Jr. & Bustamante, C. How RNA folds. J. Mol. Biol. 293, 271–281 (1999).
Leontis, N.B. & Westhof, E. Analysis of RNA motifs. Curr. Opin. Struct. Biol. 13, 300–308 (2003).
Gesteland, R.F., Cech, T.R. & Atkins, J.F. (eds.). The RNA World (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, 2004).
Webb, A.E. & Weeks, K.M. A collapsed state functions to self-chaperone RNA folding into a native ribonucleoprotein complex. Nat. Struct. Biol. 8, 135–140 (2001).
Furtig, B. et al. Time-resolved NMR studies of RNA folding. Biopolymers 86, 360–383 (2007).
Williamson, J.R. Biophysical studies of bacterial ribosome assembly. Curr. Opin. Struct. Biol. 18, 299–304 (2008).
Woodson, S.A. RNA folding and ribosome assembly. Curr. Opin. Chem. Biol. 12, 667–673 (2008).
Tijerina, P., Mohr, S. & Russell, R. DMS footprinting of structured RNAs and RNA–protein complexes. Nat. Protoc. 2, 2608–2623 (2007).
Hennelly, S.P. et al. A time-resolved investigation of ribosomal subunit association. J. Mol. Biol. 346, 1243–1258 (2005).
Sclavi, B., Woodson, S., Sullivan, M., Chance, M. & Brenowitz, M. Following the folding of RNA with time-resolved synchrotron X-ray footprinting. Methods Enzymol. 295, 379–402 (1998).
Shcherbakova, I. & Brenowitz, M. Monitoring structural changes in nucleic acids with single residue spatial and millisecond time resolution by quantitative hydroxyl radical footprinting. Nat. Protoc. 3, 288–302 (2008).
Brenowitz, M., Chance, M.R., Dhavan, G. & Takamoto, K. Probing the structural dynamics of nucleic acids by quantitative time-resolved and equilibrium hydroxy radical 'footprinting'. Curr. Opin. Struct. Biol. 12, 648–653 (2002).
Mortimer, S.A. & Weeks, K.M. Time-resolved RNA SHAPE chemistry. J. Am. Chem. Soc. 130, 16178–16180 (2008).
Merino, E.J., Wilkinson, K.A., Coughlan, J.L. & Weeks, K.M. RNA structure analysis at single nucleotide resolution by selective 2′-hydroxyl acylation and primer extension (SHAPE). J. Am. Chem. Soc. 127, 4223–4231 (2005).
Mortimer, S.A. & Weeks, K.M. A fast-acting reagent for accurate analysis of RNA secondary and tertiary structure by SHAPE chemistry. J. Am. Chem. Soc. 129, 4144–4145 (2007).
Gherghe, C.M., Shajani, Z., Wilkinson, K.A., Varani, G. & Weeks, K.M. Strong correlation between SHAPE chemistry and the generalized NMR order parameter (S2) in RNA. J. Am. Chem. Soc. 130, 12244–12245 (2008).
Wilkinson, K.A., Merino, E.J. & Weeks, K.M. Selective 2′-hydroxyl acylation analyzed by primer extension (SHAPE): quantitative RNA structure analysis at single nucleotide resolution. Nat. Protoc. 1, 1610–1616 (2006).
Wilkinson, K.A. et al. High-throughput SHAPE analysis reveals structures in HIV-1 genomic RNA strongly conserved across distinct biological states. PLoS Biol. 6, e96 (2008).
Wilkinson, K.A., Merino, E.J. & Weeks, K.M. RNA SHAPE chemistry reveals nonhierarchical interactions dominate equilibrium structural transitions in tRNAAsp transcripts. J. Am. Chem. Soc. 127, 4659–4667 (2005).
Wang, B., Wilkinson, K.A. & Weeks, K.M. Complex ligand-induced conformational changes in tRNAAsp revealed by single-nucleotide resolution SHAPE chemistry. Biochemistry 47, 3454–3461 (2008).
Duncan, C.D.S. & Weeks, K.M. SHAPE analysis of long-range interactions reveals extensive and thermodynamically preferred misfolding in a fragile group I intron RNA. Biochemistry 47, 8504–8513 (2008).
Leaderach, A., Shcherbakova, I., Liang, M.P., Brenowitz, M. & Altman, R.B. Local kinetic measures of macromolecular structure reveal partitioning among multiple parallel pathways from the earliest steps in the folding of a large RNA molecule. J. Mol. Biol. 358, 1179–1190 (2006).
Gherghe, C.M., Mortimer, S.A., Krahn, J.M., Thompson, N.L. & Weeks, K.M. Slow conformational dynamics at C2′-endo nucleotides in RNA. J. Am. Chem. Soc. 130, 8884–8885 (2008).
Vasa, S.M., Guex, N., Wilkinson, K.A., Weeks, K.M. & Giddings, M.C. ShapeFinder: a software system for high-throughput quantitative analysis of nucleic acid reactivity information resolved by capillary electrophoresis. RNA 14, 1979–1990 (2008).
Krasilnikov, A.S., Yang, X., Pan, T. & Mondragon, A. Crystal structure of the specificity domain of ribonuclease P. Nature 421, 760–764 (2003).
Baird, N.J., Westhof, E., Qin, H., Pan, T. & Sosnick, T.R. Structure of a folding intermediate reveals the interplay between core and peripheral elements in RNA folding. J. Mol. Biol. 352, 712–722 (2005).
Baird, N.J., Fang, X., Srividya, N., Pan, T. & Sosnick, T.R. Folding of a universal ribozyme: the ribonuclease P RNA. Q. Rev. Biophys. 40, 113–161 (2007).
Acknowledgements
This work was supported by a grant from the National Science Foundation (MCB-0416941 to K.M.W.).
Author information
Authors and Affiliations
Contributions
S.A.M. and K.M.W. collaborated on all aspects of the conception, design, presentation and writing of this paper.
Corresponding author
Rights and permissions
About this article
Cite this article
Mortimer, S., Weeks, K. Time-resolved RNA SHAPE chemistry: quantitative RNA structure analysis in one-second snapshots and at single-nucleotide resolution. Nat Protoc 4, 1413–1421 (2009). https://doi.org/10.1038/nprot.2009.126
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2009.126
This article is cited by
-
Programmable antivirals targeting critical conserved viral RNA secondary structures from influenza A virus and SARS-CoV-2
Nature Medicine (2022)
-
New windows into retroviral RNA structures
Retrovirology (2018)
-
Ribosome-dependent conformational flexibility changes and RNA dynamics of IRES domains revealed by differential SHAPE
Scientific Reports (2018)
-
A 3′-end structure in RNA2 of a crinivirus is essential for viral RNA synthesis and contributes to replication-associated translation activity
Scientific Reports (2016)
-
Cotranscriptional folding of a riboswitch at nucleotide resolution
Nature Structural & Molecular Biology (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.