Abstract
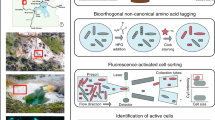
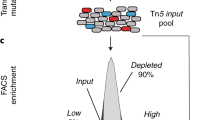
Substrate-induced gene-expression screening (SIGEX) has been developed for isolating novel catabolic genes from environmental metagenomes, particularly genes that are difficult to obtain using conventional gene-cloning methods. In SIGEX, restriction enzyme-digested metagenome fragments are ligated into an operon-trap vector (e.g., p18GFP), and a library is constructed in a liquid culture by transforming a cloning host (e.g., Escherichia coli). The library is subjected to a substrate-dependent gene-induction assay, and positive cells are selected by detecting activity of a co-expressed marker (e.g., GFP) encoded in the vector. High-throughput screening is possible if fluorescence-activated cell sorting (FACS) is used to select GFP-expressing cells. The abovementioned SIGEX procedure requires ∼17 d. In this protocol, a widely applicable SIGEX scheme is presented along with typical experimental results.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Amann, R.I., Ludwig, W. & Schleifer, K.H. Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol. Rev. 59, 143–169 (1995).
Cowan, D.A. Microbial genomes—the untapped resource. Trends Biotechnol. 18, 14–16 (2000).
Handelsman, J., Rondon, M.R., Brady, S.F., Clardy, J. & Goodman, R.M. Molecular biological access to the chemistry of unknown soil microbes: a new frontier for natural products. Chem. Biol. 5, R245–R249 (1998).
Lorenz, P., Liebeton, K., Niehaus, F. & Eck, J. Screening for novel enzymes for biocatalytic processes: accessing the metagenome as a resource of novel functional sequence space. Curr. Opin. Biotechnol. 13, 572–577 (2002).
Knietsch, A., Waschkowitz, T., Bowien, S., Henne, A. & Daniel, R. Construction and screening of metagenomic libraries derived from enrichment cultures: generation of a gene bank for genes conferring alcohol oxidoreductase activity on Escherichia coli. Appl. Environ. Microbiol. 69, 1408–1416 (2003).
Henne, A., Daniel, R., Schmitz, R.A. & Gottschalk, G. Construction of environmental DNA libraries in Escherichia coli and screening for the presence of genes conferring utilization of 4-hydroxybutyrate. Appl. Environ. Microbiol. 65, 3901–3907 (1999).
Okuta, A., Ohnishi, K. & Harayama, S. PCR isolation of catechol 2,3-dioxygenase gene fragments from environmental samples and their assembly into functional genes. Gene 212, 221–228 (1998).
Uchiyama, T., Abe, T., Ikemura, T. & Watanabe, K. Substrate-induced gene-expression screening of environmental metagenome libraries for isolation of catabolic genes. Nat. Biotechnol. 23, 88–93 (2005).
de Lorenzo, V. Problems with metagenomic screening. Nat. Biotechnol. 23, 1045 (2005).
Uchiyama, T. & Watanabe, K. The SIGEX scheme: high throughput screening of environmental metagenomes for the isolation of novel catabolic genes. Biotechnol. Genet. Eng. Rev. 24, 107–116 (2007).
Sambrook, J. & Russell, D.W. Transformation of E. coli by electroporation. in Molecular Cloning: A Laboratory Manual 3rd edn. Vol. 1 (eds. Sambrook, J. & Russell, D.W.) 1.119–1.122 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, 2001).
Uchiyama, T. & Watanabe, K. Improved inverse PCR scheme for metagenome walking. Biotechniques 41, 183–188 (2006).
Sambrook, J. & Russell, D.W. Agarose gel electrophoresis. in Molecular Cloning: A Laboratory Manual 3rd edn. Vol. 3 (eds. Sambrook, J. & Russell, D.W.) 5.4–5.13 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, 2001).
Sambrook, J. & Russell, D.W. Detection of DNA in agarose gels. in Molecular Cloning: A Laboratory Manual 3rd edn. Vol. 3 (eds. Sambrook, J. & Russell, D.W.) 5.14–5.17 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, 2001).
Sambrook, J. & Russell, D.W. Commonly used techniques in molecular cloning. in Molecular Cloning: A Laboratory Manual 3rd edn. Vol. 3 (eds. Sambrook, J. & Russell, D.W.) A8.12–A8.16 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, 2001).
Sambrook, J. & Russell, D.W. Preparation of plasmid DNA by alkaline lysis with SDS. in Molecular Cloning: A Laboratory Manual 3rd edn. Vol. 1 (eds. Sambrook, J. & Russell, D.W.) 1.31–1.42 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, 2001).
Sambrook, J. & Russell, D.W. DNA sequencing. in Molecular Cloning: A Laboratory Manual 3rd edn. Vol. 2 (eds. Sambrook, J. & Russell, D.W.) 12.1–12.120 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, 2001).
Cowles, C.E., Nichols, N.N. & Harwood, C.S. BenR, a XylS homologue, regulates three different pathways of aromatic acid degradation in Pseudomonas putida. J. Bacteriol. 182, 6339–6346 (2000).
Watanabe, K., Watanabe, K., Kodama, Y., Syutsubo, K. & Harayama, S. Molecular characterization of bacterial populations in petroleum-contaminated groundwater discharged from underground crude oil storage cavities. Appl. Environ. Microbiol. 66, 4803–4809 (2000).
Acknowledgements
This work was supported by the Japan Society for Promotion of Science. We thank Robert A. Kanaly for critical reading of this manuscript.
Author information
Authors and Affiliations
Contributions
T.U. prepared the experimental data and wrote the paper, and K.W. wrote the paper.
Corresponding author
Rights and permissions
About this article
Cite this article
Uchiyama, T., Watanabe, K. Substrate-induced gene expression (SIGEX) screening of metagenome libraries. Nat Protoc 3, 1202–1212 (2008). https://doi.org/10.1038/nprot.2008.96
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2008.96
This article is cited by
-
A plasmid-based genomic screening system for transcriptional regulators of non-adjacent xenobiotic catabolism genes
Applied Microbiology and Biotechnology (2020)
-
Functional Metagenomic Technologies for the Discovery of Novel Enzymes for Biomass Degradation and Biofuel Production
BioEnergy Research (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.