Abstract
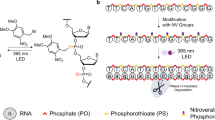
The activity of a 20-mer antisense oligodeoxynucleotide (asODN) is transiently blocked by attaching a partially complementary sense strand (sODN) via a heterobifunctional photocleavable linker (PL). The asODN-PL-sODN conjugate forms a DNA hairpin-like structure that is considerably more stable than the corresponding asODN/sODN duplex. In conjugate form, the asODN is prevented from hybridizing to exogenous RNA or DNA molecules. Activity is restored after modest exposure to UV light (λ ≈ 365 nm). Here, we provide a detailed procedure for synthesizing photoactive asODNs in good yields. Synthesis, purification and analysis of the light-activated asODN can be completed within 1–2 weeks.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Zheng, L. et al. An approach to genomewide screens of expressed small interfering RNAs in mammalian cells. Proc. Natl. Acad. Sci. USA 101, 135–140 (2004).
Peterson, R.T., Link, B.A., Dowling, J.E. & Schreiber, S.L. Small molecule developmental screens reveal the logic and timing of vertebrate development. Proc. Natl. Acad. Sci. USA 97, 12965–12969 (2000).
Ewald, A.J., Peyrot, S.M., Tyszka, J.M., Fraser, S.E. & Wallingford, J.B. Regional requirements for dishevelled signaling during Xenopus gastrulation: separable effects on blastopore closure, mesendoderm internalization and archenteron formation. Development 131, 6195–6209 (2004).
Hayashi, S. & McMahon, A.P. Efficient recombination in diverse tissues by a tamoxifen-inducible form of Cre: a tool for temporally regulated gene activation/inactivation in the mouse. Dev. Biol. 244, 305–318 (2002).
Pasquinelli, A.E. & Ruvkun, G. Control of developmental timing by microRNAs and their targets. Annu. Rev. Cell Dev. Biol. 18, 495–513 (2002).
Yuh, C.-H., Dorman, E.R., Howard, M.L. & Davidson, E.H. An otx cis-regulatory module: a key node in the sea urchin endomesoderm gene regulatory network. Dev. Biol. 269, 536–551 (2004).
McCray, J.A. & Trentham, D.R. Properties and uses of photoreactive caged compounds. Annu. Rev. Biophys. Biophys. Chem. 18, 239–270 (1989).
Goeldner, M. & Givens, R. (eds.) Dynamic Studies in Biology (Wiley-VCH, Weinheim, 2005).
Pelliccioli, A.P. & Wirz, J. Photoremovable protecting groups: reaction mechanisms and applications. Photochem. Photobiol. Sci. 1, 441–458 (2002).
Mayer, G. & Heckel, A. Biologically active molecules with a 'light switch'. Angew. Chem. Int. Ed. Engl. 45, 4900–4921 (2006).
Adams, S.R. & Tsien, R.Y. Controlling cell chemistry with caged compounds. Annu. Rev. Physiol. 55, 755–784 (1993).
Lima, S.Q. & Miesenbock, G. Remote control of behavior through genetically targeted photostimulation of neurons. Cell 121, 141–152 (2005).
Carter, A.G. & Sabatini, B.L. State-dependent calcium signaling in dendritic spines of striatal medium spiny neurons. Neuron 44, 483–493 (2004).
Piston, D.W., Summers, R.G., Knobel, S.M. & Morrill, J.B. Characterization of involution during sea urchin gastrulation using two-photon excited photorelease and confocal microscopy. Microsc. Microanal. 4, 404–414 (1998).
Shoham, S., O'Connor, D.H., Sarkisov, D.V. & Wang, S.S.-H. Rapid neurotransmitter uncaging in spatially defined patterns. Nat. Methods 2, 837–843 (2005).
Ando, H. et al. Lhx2 mediates the activity of Six3 in zebrafish forebrain growth. Dev. Biol. 287, 456–468 (2005).
Ando, H., Furuta, T., Tsien, R.Y. & Okamoto, H. Photo-mediated gene activation using caged RNA/DNA in zebrafish embryos. Nat. Genet. 28, 317–325 (2001).
Monroe, W.T., McQuain, M.M., Chang, M.S., Alexander, J.S. & Haselton, F.R. Targeting expression with light using caged DNA. J. Biol. Chem. 274, 20895–20900 (1999).
Shah, S., Rangarajan, S. & Friedman, S.H. Light-activated RNA interference. Angew. Chem. Int. Ed. Engl. 44, 1328–1332 (2005).
Ghosn, B., Haselton, F.R., Gee, K.R. & Monroe, W.T. Control of DNA hybridization with photocleavable adducts. Photochem. Photobiol. 81, 953–959 (2005).
Tang, X. & Dmochowski, I.J. Phototriggering of caged fluorescent oligodeoxynucleotides. Org. Lett. 7, 279–282 (2005).
Matsunaga, D., Asanuma, H. & Komiyama, M. Photoregulation of RNA digestion by RNase H with azobenzene-tethered DNA. J. Am. Chem. Soc. 126, 11452–11453 (2004).
Tang, X. & Dmochowski, I.J. Controlling RNA digestion by RNase H with a light-activated DNA hairpin. Angew. Chem. Int. Ed. Engl. 45, 3523–3526 (2006).
Goldmacher, V.S., Senter, P.D., Lambert, J.M. & Blattler, W.A. Photoactivation of toxin conjugates. Bioconjug. Chem. 3, 104–107 (1992).
Senter, P.D., Tansey, M.J., Lambert, J.M. & Blattler, W.A. Novel photocleavable protein crosslinking reagents and their use in the preparation of antibody-toxin conjugates. Photochem. Photobiol. 42, 231–237 (1985).
Kwart, H. & Burchuk, I. Isomerism and adduct stability in the Diels-Alder reaction. I. The adducts of furan and maleimide. J. Am. Chem. Soc. 74, 3094–3097 (1952).
Ishii, Y. & Lehrer, S.S. Effects of the state of the succinimido-ring on the fluorescence and structural properties of pyrene maleimide-labeled alpha alpha-tropomyosin. Biophys. J. 50, 75–80 (1986).
Sambrook, J. & Russell, D.W. In Molecular Cloning: A Laboratory Manual (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, 2001) 9.66–9.69.
Acknowledgements
Support for this work came from grants from the Penn Genomics Institute (PGI) and Penn Institute for Medicine and Engineering (IME), and startup funds provided to I.J.D. by the Department of Chemistry at the University of Pennsylvania.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Tang, X., Dmochowski, I. Synthesis of light-activated antisense oligodeoxynucleotide. Nat Protoc 1, 3041–3048 (2006). https://doi.org/10.1038/nprot.2006.462
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.462
This article is cited by
-
Transcriptome in vivo analysis (TIVA) of spatially defined single cells in live tissue
Nature Methods (2014)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.