Abstract
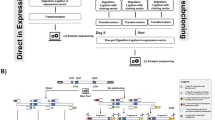
A simple protocol to introduce random mutations, named error-prone rolling circle amplification (RCA), is described. A template plasmid is amplified by RCA in the presence of MnCl2 and used for transformation of a host strain to give a mutant library with three to four random point mutations per kilobase throughout the entire plasmid. The prime advantage of this method is its simplicity. This protocol requires neither the design of specific primers nor the exploration of thermal cycling conditions. It takes just 10 min to prepare the reaction mixture, followed by overnight incubation and transformation of a host strain. This method permits rapid preparation of randomly mutated plasmid libraries, and will enable the wider adoption of random mutagenesis.
NOTE: In the PDF version of this article initially published online, the publication date was shown as 29 December 2007 instead of 29 December 2006. The error has been corrected in the PDF version of the article.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
22 February 2007
changed 2007 to 2006
References
Bloom, J.D. et al. Evolving strategies for enzyme engineering. Curr. Opin. Struct. Biol. 15, 447–452 (2005).
Jaeger, K.E. & Eggert, T. Enantioselective biocatalysis optimized by directed evolution. Curr. Opin. Biotechnol. 15, 305–313 (2004).
Arnold, F.H., Wintrode, P.L., Miyazaki, K. & Gershenson, A. How enzymes adapt: lessons from directed evolution. Trends Biochem. Sci. 26, 100–106 (2001).
Johannes, T.W. & Zhao, H. Directed evolution of enzymes and biosynthetic pathways. Curr. Opin. Microbiol. 9, 261–267 (2006).
Reetz, M.T. Directed evolution of enantioselective enzymes as catalysts for organic synthesis. Adv. Catal. 49, 1–69 (2006).
Aharoni, A., Griffiths, A.D. & Tawfik, D.S. High-throughput screens and selections of enzyme-encoding genes. Curr. Opin. Chem. Biol. 9, 210–216 (2005).
Goddard, J.P. & Reymond, J.L. Enzyme assays for high-throughput screening. Curr. Opin. Biotechnol. 15, 314–322 (2004).
Taylor, S.V., Kast, P. & Hilvert, D. Investigating and engineering enzymes by genetic selection. Angew. Chem. Int. Ed. Engl. 40, 3310–3335 (2001).
Lin, H. & Cornish, V.W. Screening and selection methods for large-scale analysis of protein function. Angew. Chem. Int. Ed. Engl. 41, 4402–4425 (2002).
Reetz, M.T. & Jaeger, K.E. Superior biocatalysts by directed evolution. In Biocatalysis - From Discovery to Application 31–57 (Springer-Verlag, Berlin, 1999).
Leung, D.W., Chen, E. & Goeddel, D.W. A method for random mutagenesis of a defined DNA segment using a modified polymerase chain reaction. Techniques 1, 11–15 (1989).
Greener, A., Callahan, M. & Jerpseth, B. In in vitro Mutagenesis Protocols (ed. Trower, M.K.) (Humana press, New Jersey, 1996).
Kornberg, A. & Baker, T. In DNA Replication (W.H. Freeman & Company, NY, 1992).
Fire, A. & Xu, S.Q. Rolling replication of short DNA circles. Proc. Natl. Acad. Sci. USA 92, 4641–4645 (1995).
Liu, D.Y., Daubendiek, S.L., Zillman, M.A., Ryan, K. & Kool, E.T. Rolling circle DNA synthesis: small circular oligonucleotides as efficient templates for DNA polymerases. J. Am. Chem. Soc. 118, 1587–1594 (1996).
Lizardi, P.M. et al. Mutation detection and single-molecule counting using isothermal rolling-circle amplification. Nat. Genet. 19, 225–232 (1998).
Dean, F.B., Nelson, J.R., Giesler, T.L. & Lasken, R.S. Rapid amplification of plasmid and phage DNA using Phi 29 DNA polymerase and multiply-primed rolling circle amplification. Genome Res. 11, 1095–1099 (2001).
Fujii, R., Kitaoka, M. & Hayashi, K. One-step random mutagenesis by error-prone rolling circle amplification. Nucleic Acids Res. 32, e145 (2004).
Ding, X., Snyder, A.K., Shaw, R., Farmerie, W.G. & Song, W.Y. Direct retransformation of yeast with plasmid DNA isolated from single yeast colonies using rolling circle amplification. BioTechniques 35, 774–779 (2003).
Camps, M., Naukkarinen, J., Johnson, B.P. & Loeb, L.A. Targeted gene evolution in Escherichia coli using a highly error-prone DNA polymerase I. Proc. Natl. Acad. Sci. USA 100, 9727–9732 (2003).
Henke, E. & Bornscheuer, U.T. Directed evolution of an esterase from Psueudomonas fluorescens. Random mutagenesis by error-prone PCR or a mutator strain and identification of mutants showing enhanced enantioselectivity by a resorufin-based fluorescence assay. Biol. Chem. 380, 1029–1033 (1999).
Bornscheuer, U.T., Altenbuchner, J. & Meyer, H.H. Directed evolution of an esterase for the stereoselective resolution of a key intermediate in the synthesis of epothilones. Biotechnol. Bioeng. 58, 554–559 (1998).
Reetz, M.T., Zonta, A., Schimossek, K., Liebeton, K. & Jaeger, K.E. Creation of enantioselective biocatalysts for organic chemistry by in vitro evolution. Angew. Chem. Int. Ed. Engl. 36, 2830–2832 (1997).
de Vega, M., Lazaro, J.M. & Salas, M. Phage φ29 DNA polymerase residues involved in the proper stabilisation of the primer-terminus at the 3′-5′ exonuclease active site. J. Mol. Biol. 304, 1–9 (2000).
Voss, C., Schmidt, T., Schleef, M., Friehs, K. & Flaschel, E. Production of supercoiled multimeric plasmid DNA for biopharmaceutical application. J. Biotechnol. 105, 205–213 (2003).
Vakulenko, S.B. et al. Effects on substrate profile by mutational substitutions at positions 164 and 179 of the class A TEMpUC19 β-lactamase from Escherichia coli. J. Biol. Chem. 274, 23052–23060 (1999).
Vakulenko, S.B., Toth, M., Taibi, P., Mobashery, S. & Lerner, S.A. Effects of Asp-179 mutations in TEMpUC19 β-lactamase on susceptibility to β-lactams. Antimicrob. Agents Chemother. 39, 1878–1880 (1995).
Blanco, L. et al. Highly efficient DNA-synthesis by the phage φ29 DNA polymerase. J. Biol. Chem. 264, 8935–8940 (1989).
Acknowledgements
This study was supported in part by a grant from the Program for Promotion of Basic Research Activities for Innovative Biosciences (PROBRAIN).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Fujii, R., Kitaoka, M. & Hayashi, K. Error-prone rolling circle amplification: the simplest random mutagenesis protocol. Nat Protoc 1, 2493–2497 (2006). https://doi.org/10.1038/nprot.2006.403
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.403
This article is cited by
-
In vitro self-replication and multicistronic expression of large synthetic genomes
Nature Communications (2020)
-
Biomolecular engineering for nanobio/bionanotechnology
Nano Convergence (2017)
-
Development of simple random mutagenesis protocol for the protein expression system in Pichia pastoris
Biotechnology for Biofuels (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.