Abstract
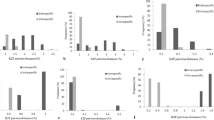
A DNA barcode consists of a standardized short sequence of DNA (400-800bp) used to identify the taxonomic species a small organic fragment belongs to. Even though it has been easy to discriminate animal species by using the mitochondrial gene cox1, this is still difficult for plants seeing that the mitochondrial genome is not variable enough on the species level. During the Second International Barcode of Life Conference in Tapei (September 2007), different plastid regions were proposed as potential plant DNA barcodes, such as atpF-atpH and psbK-psbI, but no consensus on which region to use was reached during the meeting. The largest plant DNA barcoding study to date proposed matK as the best candidate and suggested that in combination with trnH-psbA a slight increase in performance could be achieved. However, no study has tested the suitability of the newly proposed psbK-psbI and atpF-atpH for plant barcoding purposes. Four potential DNA barcodes, matK, trnH-psbA, atpF-atpH, and psbK-psbI, were amplified and sequenced for a selective sampling including mainly trees and shrubs of the flora of the Kruger National Park Africa (South Africa). The performance of each region and also each possible combination of these were tested by applying a battery of metrics and statistical tests. Our results confirm that the second half (5’ end) of matK is the best candidate in a single locus barcoding approach reaching 87.5% of species correctly identified. Combining matK with trnH-psbA and psbK-psbI increased only slightly the performance in discriminating species. The results from this study show that the use of a ‘three-region barcode’ does not significantly outperform matK in a single-locus barcoding approach. We therefore argue against the ‘multiple barcode approach’ proposed by the plant working group, and instead propose to keep barcoding plants in line with the approach taken for animals, i.e. using one barcode: cox1 for animals and matK for plants.
Similar content being viewed by others
Article PDF
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lahaye, R., Savolainen, V., Duthoit, S. et al. A test of psbK-psbI and atpF-atpH as potential plant DNA barcodes using the flora of the Kruger National Park (South Africa) as a model system. Nat Prec (2008). https://doi.org/10.1038/npre.2008.1896.1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/npre.2008.1896.1
Keywords
This article is cited by
-
Development of chloroplast marker for identification of Ulva species
Journal of Oceanology and Limnology (2022)
-
Re-evaluation of the phylogenetic relationships and species delimitation of two closely related families (Lamiaceae and Verbenaceae) using two DNA barcode markers
Journal of Biosciences (2020)
-
Viola woosanensis, a recurrent spontaneous hybrid between V. ulleungdoensis and V. chaerophylloides (Violaceae) endemic to Ulleung Island, Korea
Journal of Plant Research (2016)
-
Phylogeographical structure inferred from cpDNA sequence variation of Zygophyllum xanthoxylon across north-west China
Journal of Plant Research (2015)
-
DNA Barcoding in Endangered Mesoamerican Groups of Plants
The Botanical Review (2013)