Abstract
Aberrant biochemical processes in the brain frequently go along with subtle shifts of the cellular epigenetic profile that might support the pathogenic progression of psychiatric disorders. Although recent reports have implied the ability of certain antidepressants and mood stabilizers to modulate epigenetic parameters, studies comparing the actions of these compounds under the same conditions are lacking. In this study, we screened amitriptyline (AMI), venlafaxine, citalopram, as well as valproic acid (VPA), carbamazepine, and lamotrigine for their potential actions on global and local epigenetic modifications in rat primary astrocytes. Among all drugs, VPA exposure evoked the strongest global chromatin modifications, including histone H3/H4 hyperacetylation, 2MeH3K9 hypomethylation, and DNA demethylation, as determined by western blot and luminometric methylation analysis, respectively. CpG demethylation occurred independently of DNA methyltransferase (DNMT) suppression. Strikingly, AMI also induced slight cytosine demethylation, paralleled by the reduction in DNMT enzymatic activity, without affecting the global histone acetylation status. Locally, VPA-induced chromatin modifications were reflected at the glutamate transporter (GLT-1) promoter as shown by bisulfite sequencing and acetylated histone H4 chromatin immunoprecipitation analysis. Distinct CpG sites in the distal part of the GLT-1 promoter were demethylated and enriched in acetylated histone H4 in response to VPA. For the first time, we could show that these changes were associated with an enhanced transcription of this astrocyte-specific gene. In contrast, AMI failed to stimulate GLT-1 transcription and to alter promoter methylation levels. In conclusion, VPA and AMI globally exerted chromatin-modulating activities using different mechanisms that divergently precipitated at an astroglial gene locus.
Similar content being viewed by others
INTRODUCTION
The vertebrate brain is exposed to a number of environmental cues that are processed in an adaptive manner. These adaptive processes form the basis of neuroplasticity, and are characterized by often-persistent changes in neural gene expression. Epigenetics represents an essential element for transmitting environmental cues into altered neural molecular pathways (Szyf et al, 2008). Frequently, epigenetic chromatin modifications affect the acetylation or methylation status of histones and the methylation of cytosine residues within CpG dinucleotides. Histone acetylation and DNA methylation show some degree of interdependency, although a detailed analysis of this interaction is still pending (D'Alessio and Szyf, 2006). Epigenetic modifications subsequently alter chromatin packaging, and, if occurring in the proximity of gene promoters, exons or introns, either promote or suppress gene transcription (Herman and Baylin, 2003). Enzymes catalyzing these epigenetic reactions belong to the classes of histone acetyltransferases (HATs) and methyltransferases (HMTs), histone deacetylases (HDACs), and DNA methyltransferases (DNMTs) (Ng and Bird, 1999).
Recently, it has been shown how external stimuli can be translated into modified epigenetic marks for the example of the brain-derived neurotrophic factor (BDNF) promoter (Chen et al, 2003). After membrane depolarization on isolated neurons, specific CpG dinucleotides within the BDNF promoter underwent demethylation, accompanied by the upregulation of gene transcription. Similarly, mRNA and protein levels of distinct DNMT subtypes were reduced under similar conditions, which resulted in a diminished activity of the respective enzymes (Sharma et al, 2008).
The consequence of an intrinsically false or inadequate adaptation process in response to detrimental environmental factors such as stress might go along with the establishment of aberrant epigenetic signatures. In humans, there is mounting evidence that epigenetic mechanisms are involved in the pathophysiology of stress-related disorders, including unipolar and bipolar depression (Mill and Petronis, 2007; Petronis, 2003). Postmortem studies of suicide victims with a history of depression or childhood abuse showed differential expression of DNMT subtypes (Poulter et al, 2008) and promoter-wide hypermethylation of ribosomal RNA gene promoters (McGowan et al, 2008), respectively. Humans who experienced childhood abuse and committed suicide also showed increased methylation of the neuron-specific glucocorticoid receptor (GR) promoter in the hippocampus paralleled by reduced levels of the corresponding mRNA transcripts (McGowan et al, 2009). In general, GR has a pivotal role in balancing the hypothalamic pituitary axis (HPA) in response to eg, stressful situations, and a HPA imbalance is observed in various psychiatric disorders. In rodents, it was recently shown that poor maternal care of the offspring increased DNA methylation of the GR exon 17 promoter in the hippocampus and, thereby impaired negative feedback regulation of HPA axis due to lower GR protein levels (Weaver et al, 2004). Interestingly, application of an HDAC inhibitor (HDAC-I) restored normal HPA response to stress in the offspring. In addition, chronic social stress in rodents led to distinct biochemical histone modifications within the BDNF gene promoter, paralleled by a reduced BDNF transcription (Tsankova et al, 2006).
Hence, pharmacological interference with the epigenetic setup of the cell might represent an avenue to normalize the aberrantly installed epigenetic profiles. So far, drug therapy of patients suffering from bipolar and major depression is based on the stabilization of neurotransmitter circuits that involve the serotonergic, noradrenergic, glutamatergic, and GABAergic system. Most of the therapeutic compounds exert their beneficial effects after a latency period, indicating the necessity for preceding adaptive processes. This raises the question, whether psychoactive substances would potentially interfere directly or indirectly with the epigenetic makeup of exposed cells. Indeed, the mood stabilizer, valproic acid (VPA), was the first psychoactive drug shown to exhibit HDAC inhibitory properties (Gottlicher et al, 2001; Phiel et al, 2001). Another study investigated antiepileptic drugs and identified topiramate as an inhibitor of HDAC activity (Eyal et al, 2004).
In this study, we aimed to comprehensively analyze the potential of frequently prescribed antidepressants, including amitriptyline (AMI), venlafaxine (VEN), citalopram (CIT), and mood stabilizers such as VPA, carbamazepine (CBZ), and lamotrigine (LTG) to affect epigenetic parameters in astrocytes. Astrocytes represent the majority of cells within the CNS, and modulate synaptic strength by their association with synapses (Schipke and Kettenmann, 2004). In addition, astrocytes support neuronal function, as they produce neurotrophic factors, including BDNF and glial-derived neurotrophic factor, also in response to VPA exposure (Chen et al, 2006; Wu et al, 2008c). We monitored drug-induced global epigenetic changes by measuring histone H3 and H4 acetylation/methylation, as well as by DNA methylation. To explore whether global changes would be also reflected at gene loci that are important for astrocytic function, we selected the glutamate transporter (GLT-1) promoter as a model gene.
MATERIALS AND METHODS
Primary Astrocyte Cell Culture
Enriched astroglial cultures were prepared from postnatal day 1 rat pups (Sprague–Dawley, Charles River, Sulzfeld, Germany) as described previously (Franke et al, 1998). In these cultures, 90% of all cells represent type-1 astrocytes. Briefly, the dissected cortical hemispheres and hippocampus were trypsinized (Invitrogen, Merelbeke, Belgium); trypsin action was terminated by adding Hanks' balanced salt solution (Invitrogen), supplemented with 10% fetal calf serum. The tissue was passed through a serological pipette, spun, and resuspended in modified Eagle's medium (MEM, Invitrogen) supplemented with 10% horse serum (Invitrogen). After the third passage, the cells were maintained in a serum-free (MEM/ Ham's F-12, 1 : 1) N2-supplemented medium.
Pharmacological Treatment of Astrocytes
All experiments were carried out using third-passage astrocytic cells. Sodium butyrate (NaB), VPA, CBZ, LTG, AMI, CIT, and trichostatin A (TSA) were purchased from Sigma (Deisenhof, Germany). VEN was an industrial donation (Wyeth Pharma GmbH, Münster, Germany). Stock solutions of the drugs were prepared by dissolving the substances in distilled water (VPA, NaB, AMI, CIT, VEN), 100% ethanol (TSA), or DMSO (CBZ, LTG). When treating cells for 72 h, the drug-containing medium was renewed after 48 h. For washout experiments, after 72 h of treatment, the cells were rinsed twice with the medium, and thereafter kept in the absence of the drug for an additional 48 h.
Cellular Protein Extraction and Western Blot Analysis
Cells were lysed in the buffer containing 62.5 mM Tris, 2% SDS, and 10% sucrose, supplemented with protease inhibitor cocktail (Sigma). Samples were sonicated and heated at 95°C for 5 min. Proteins were separated by SDS-PAGE and electrotransferred onto nitrocellulose membranes. Blots were placed in Tris-buffered saline, supplemented with 0.05% Tween (Sigma) and 5% non-fat milk for 1 h at room temperature, and then incubated with primary antibody (diluted in TBS/0.05% Tween) overnight at 4°C. Subsequently, the blots were washed and probed with the respective horseradish peroxidase-conjugated secondary antibody for 1 h at room temperature. The immunoreactive bands were visualized using the ECL detection reagent (Millipore, Billerica, MA, USA). The following primary antibodies were used: anti-acetyl-Histone H3 antibody (recognizes acetylation at lysine 9; 1 : 2000; no. 06–942, Upstate, Temecula, CA, USA), anti-acetyl-Histone H4 antibody (recognizes acetylation at lysines 5, 8, 12, and 16; 1 : 4000; no. 06–866, Upstate), anti-DNMT-1 (1 : 1000; no. IMG-261A, IMGENEX, San Diego, CA, USA), anti-GADD 45α (1 : 250; no. sc-6850, Santa Cruz Biotechnology, Heidelberg, Germany), anti-dimethyl-Histone H3 antibodies (react with methylated histones on lysine 9 or 27; 1 : 4000; a kind gift from Dr Thomas Jenuwein, Max-Planck-Institute of Immunobiology, Freiburg, Germany), anti-α-actin (1 : 2500; no. A2066, Sigma), and anti-hsp90 (1 : 2000; no. sc-7947, Santa Cruz Biotechnology). Determination of the relative optical density and quantification of band intensities were performed using the Kodak Image Analysis Software.
DNMT Activity Assay
Nuclear extracts from cortical astrocytes were obtained using the EpiQuik Nuclear Extraction Kit (Epigentek, Brooklyn, NY) according to the manufacturer's instructions. DNMT activity was determined by incubating 4 μg of protein nuclear extract with 0.5 μg Poly (deoxyinosinic-deoxycytidylic) acid (Poly(dI-dC)·Poly(dI-dC); Sigma) in reaction buffer (20 mM Tris pH 7.8, 10% glycerol, 5 mM EDTA, 1 mM DTT, 0.2 mM PMSF) containing 3 μM S-adenosyl-L-[methyl-3H]methionine (3H-SAM, specific activity: 15 Ci/mmol; GE Healthcare, Munich, Germany) for 3 h at 37°C. Thereafter, DNA was isolated using the GeneClean Kit (MP Biomedicals). To reduce background signals, three washing steps were conducted and the DNA binding step was performed in the presence of a 1000 times excess of unlabeled SAM to reduce nonspecific binding of the labeled SAM. To minimize the loss of DNA, genomic DNA (3 μg per sample) was added to the binding buffer.
Isolation of Genomic DNA
Isolation of genomic DNA was carried out by salting out proteins and precipitating the genomic DNA with isopropanol as described previously (Zschocke et al, 2007).
Luminometric Methylation Analysis
Luminometric methylation analysis (LUMA) was performed according to Karimi et al (2006), with minor modifications. Briefly, 2 μg of genomic DNA was cleaved with 4 Units of Eco RI (Fermentas, St Leon-Rot, Germany) in a buffer containing 66 mM Tris-acetate, 20 mM Mg-acetate, 0.2 mg/ml BSA, 132 mM K-acetate pH 7.9, for 2 h at 37°C. Eco RI was inactivated at 65°C for 20 min. Subsequently, the reaction content was diluted by adding an equal amount of dH2O. In two separate 20 μl reactions, DNA (1 μg each) was subjected to enzymatic digestion by adding 2 Units of Hpa II or Msp I (New England Biolabs, Frankfurt, Germany), and the mixture was incubated 4 h at 37°C. After heat inactivation of Hpa II and Msp I at 65°C for 20 min, samples were processed using a pyrosequencer (Biotage, Uppsala, Sweden). The Hpa II/Eco RI and Msp I/Eco RI ratios were calculated, and the result of the measurement was expressed as the percentage of CCpGG methylation.
Methylation Analysis by Bisulfite Genomic Sequencing
Bisulfite treatment of genomic DNA (2 μg) was performed using the EpiTech Bisulfite Kit (Qiagen, Hilden, Germany) according to the manufacturer's instructions. Bisulfite-specific primers were designed using Methyl Primer Express Software (Applied Biosystems). The PCR mixture was composed of 1 × PCR Buffer (Invitrogen), 1.5 mM MgCl2 (Invitrogen), 0.2 mM of each dNTP (peqLAB, Erlangen, Germany), 0.5–1 μM of each primer, and 4 Units of DNA Taq polymerase (Invitrogen). Sequences of primers are listed in Table 1. Methylation of individual CpG sites was detected by direct sequencing (Qiagen). All sequencing reaction mixtures were based on the BigDye 3.1 Terminator chemistry (Applied Biosystems). Data collection was carried out on a 3730 × l DNA Analyzer (Applied Biosystems).
Quantitative Real-Time RT-PCR
Total RNA was prepared from astroglial cells according to the manufacturer's protocol for NucleoSpin RNA II (Macherey-Nagel, Düren, Germany). A total of 500 ng of purified RNA was transcribed with random primers and Omniscript Reverse Transcriptase (Qiagen). Quantitative real-time PCR analysis of GLT-1 cDNA was performed using the LightCycler Carousel-Based System (Roche Applied Science, Mannheim, Germany) and SYBR green (QantiFast SYBR Green, Qiagen). β-Actin was used as an endogenous control for normalization. The primers used are listed in Table 1. Relative changes of gene expression were calculated using the comparative ΔΔCT method.
Transient Transfection and Luciferase Reporter Assay
Approximately 3−5 × 106 primary cortical rat astrocytes were transiently transfected with human EAAT2 promoter reporter plasmid (a kind gift from Dr Jürgen Engele, University of Leipzig, Leipzig, Germany) using Amaxa's Nucleofector device (Amaxa Biosystem, Cologne, Germany). Simian virus 40 promoter-driven nonsecretory Gaussia luciferase expression vector was cotransfected to correct for transfection efficiency. Twenty-four hours after transfection, drug treatment was carried out for 24 h. Luciferase assays were performed by applying the Dual Luciferase Assay System (Promega, Mannheim, Germany) using the TriStar LB941 Luminometer (Berthold Technologies, Wildbad, Germany).
Chromatin Immunoprecipitation
After pharmacological treatment, 2 × 106 astrocytes were fixed in vivo with 1% formaldehyde for 10 min, and the reaction was finally quenched with 0.125 M glycine for 5 min at RT. From then on, all steps were carried out at 4°C, if not stated otherwise. Nuclei were extracted in 1 ml hypotonic buffer composed of 0.5% Igepal-CA630, 10 mM KCl, 1.5 mM MgCl2, and 10 mM HEPES, pH 7.9, followed by 15 min of incubation on ice and brief homogenization. Nuclei were recovered by 5 min by centrifugation at 800 g. Nuclear pellets were dissolved in 400 μl lysis buffer (containing 1% SDS, 10 mM EDTA, 50 mM Tris-HCl, pH 8.1) and sonicated using a Branson sonifier (12 × 20 s bursts, 90% pulse, output control 4). The average length of DNA fragments generated by sonication was 500 bp. Two 200 μl aliquots (for IgG control and 2 μg of anti-acetyl-H4 antibody) were 1 : 10 diluted in ChIP dilution buffer (0.01% SDS, 1.1% Triton, 1.2 mM EDTA, 167 mM NaCl, and 16.7 mM Tris-HCl, pH 8.1) and precleared for 3 h with blocked Dynabeads (Promega, protein A coupled, blocked with 0.1% BSA and 0.05% salmon sperm DNA). Anti-acetyl-H4 antibody was added to the samples, preincubated for another 2 h, and rotated overnight with blocked Dynabeads. Four subsequent washing steps were carried out using low salt (0.1% SDS, 1% Triton, 2 mM EDTA, 150 mM NaCl, 20 mM Tris-HCl, pH 8.1), high salt (see low salt, except 500 mM NaCl), LiCl (0.25 M LiCl, 1% NP-40, 1% deoxy-cholate, 1 mM EDTA, 10 mM Tris-HCl, ph 8.1), and TE buffer. Elution and reversion of cross-links were performed in buffer containing 1% SDS, 50 mM NaHCO3, 1 mM EDTA, 50 mM Tris-HCl pH 8.0, for 2 h at 62°C and a final 95°C heating step for 5 min. DNA was purified using the ultra-clean PCR Clean-up Kit according to the manufacturer's instructions (MoBio Laboratories, Carlsbad, CA, USA), and later on used in real-time PCR using primers listed in Table 1.
RESULTS
VPA, but not Other Psychoactive Drugs, Induce Global Histone Hyperacetylation
In this study, we set out to explore the effects of different classes of antidepressants (such as VEN, CIT, AMI) and mood stabilizers (such as VPA, LTG, CBZ) on the epigenetic machinery of primary rat astrocytes, which express the classical targets of antidepressants such as serotonin and noradrenaline transporters, and different 5 HT-receptor subtypes (Azmitia, 2001; Inazu et al, 2001; Inazu et al, 2003). To further evaluate mechanistic aspects, we also included the established HDAC-I TSA and NaB in some assays.
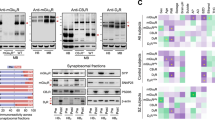
First, we measured the degree of global histone H3 and H4 acetylation (AcH3, AcH4), which is indicative of cellular HAT and HDAC activity, after 24 h of drug exposure at two different concentrations. The lower concentrations are in the range of therapeutic plasma levels, whereas higher concentrations represent a value that might be reached in the brain assuming cumulative processes and enrichment of the drugs at the loci of action. In addition, TSA was used as a pan-inhibitor of HDACs. Western blot analysis showed that out of all psychoactive drugs, only VPA induced hyperacetylation of both histones H3/H4 in a dose-dependent manner (Figure 1). Moreover, there was no profound difference in the pattern of induction when comparing astrocytes from the hippocampus and cortex.
Screen of psychoactive drugs for their global effects on histone acetylation. Cortical (CTX; a, b) and hippocampal astrocytes (HPC; c, d) were treated with VPA (1 and 10 mM), LTG (10 and 100 μM), CBZ (10 and 100 μM), VEN (1 and 10 μM), CIT (1 and 10 μM), AMI (1 and 10 μM), or with the HDAC inhibitor TSA (0.2 μM) for 24 h. After drug treatment, cell lysates were prepared and subjected to western blot analysis using polyclonal antibodies against AcH3K9 and AcH4. AcH3K9 and AcH4 band intensities were normalized to α-actin. The bars are depicted as the percentage of optical band densities (mean±SE, n=2) calculated relative to the value that corresponds to histone acetylation induced by TSA (set to 100%); t-test: *p<0.05 vs untreated cells, **p<0.005 vs untreated cells.
Dynamic Changes of Histone Hyperacetylation After VPA Exposure
To evaluate the dynamics of VPA-mediated histone acetylation during and after drug exposure, we conducted a time-course analysis of 24 and 72 h VPA treatment, followed by a drug-removal step for 48 h. In both, cortical and hippocampal astrocytes, VPA-induced hyperacetylation of H3K9 was stronger after 24 h than after 72 h, whereas this difference was less pronounced in the case of H4 acetylation (Figure 2). A similar trend was observed in cells treated with 0.2 μM TSA. Forty-eight hours after VPA withdrawal, AcH3K9 and AcH4 returned to basal levels. Our results indicate that the effects of VPA on histone acetylation are only transient, peak during the first hours of treatment, and are fully reversible after drug removal.
VPA induces dynamic changes of AcH3K9 and AcH4 patterns in astrocytes from different brain regions. Cortical (a, b) and hippocampal (c, d) astrocytes were exposed to 1 and 10 mM VPA for 24, 72, and 72 h, followed by a 48 h washout period, whereas a single dose of TSA (0.2 μM) was applied for 24 and 72 h. The level of AcH3K9 and AcH4 was monitored by western blot analysis and detected with AcH3K9- and AcH4-specific antibodies. α-Actin immunoreactivity served as a control for protein loading. Optical densities of AcH3K9- and AcH4-specific bands were normalized to α-actin. Data represent mean±SE (n=2) of the percentage of optical band densities calculated relative to the value that corresponds to histone acetylation induced by 24 h treatment with 0.2 μM TSA (set to 100%); t-test: *p<0.05 vs untreated cells, **p<0.005 vs untreated cells, #p<0.05 vs 1 mM VPA depletion treatment, §p<0.05 vs 10 mM VPA depletion treatment.
DNA Demethylation at CCpGG Sites Exerted by VPA is Reversible
Histone acetylation and DNA methylation operate in a concerted manner. In agreement with this hypothesis, HDAC-Is are capable of inducing DNA demethylation in human cancer cell lines (Detich et al, 2003; Ou et al, 2007) and in vivo (Dong et al, 2007). Therefore, we examined whether VPA/TSA trigger CpG demethylating in nontransformed primary astrocytes. In total, three different methods were used to determine the degree of global CpG methylation. LUMA measures the average methylation status of all CCpGG sites by Hpa II/Msp I restriction analysis, coupled to a Taq polymerase-dependent luminometric reaction. The second, capillary electrophoresis technique depicts the average methylation status of all cytosines. Third, we analyzed long interspersed nuclear elements (LINE-1), serving as surrogate markers of global DNA methylation changes (Yang et al, 2004), by applying direct bisulfite sequencing of a certain stretch of the LINE-1 ORFII.
First, we validated the accuracy of LUMA and bisulfite sequencing with respect to CpG methylation. There was a high degree of similarity between results obtained by LUMA and direct bisulfite sequencing (r2=0.98, Supplementary Figure S1a).
As determined by LUMA, untreated cortical and hippocampal astrocytes displayed 64±1% and 67±1% methylation of CCpGG sites, respectively (Supplementary Figure S1b). In cells from the cortex, treatment with 1 mM VPA induced significant demethylation after 72 h (−4.9±1.8%), whereas 48 h after drug withdrawal, the level of CCpGG methylation returned to baseline (Table 2). At a concentration of 10 mM, VPA triggered DNA demethylation by −7.5±1.3% already after 24 h, and reached −17.2±2.0% after 72 h. Again, demethylation was reversed after 48 h of drug washout. In cells from the hippocampus, results obtained by LUMA were slightly less coherent. After 24 and 72 h of 1 mM VPA exposure, significant hypomethylation (−9.7±2.4% and −5.2±0.6%, respectively) was observed. Higher doses of VPA of 10 mM induced −5.0±1.4% hypomethylation after 72 h. Although the reversibility of methylation changes was not as pronounced as in the cortex, it appeared to be in this direction. Finally, 0.2 μM TSA induced significant DNA hypomethylation at CCpGG sites in the cortex and hippocampus. Collectively, our results suggest that VPA-induced hypomethylation of CCpGG profiles are present in both brain regions, although magnitude and kinetics seem to differ slightly.
DNA demethylation processes can occur passively by eg, retention of DNMT enzymes in the cytoplasm during cell-cycle transitions. In addition, active mechanisms of DNA demethylation occurring in quiescent cells in short periods of time are discussed, requiring a putative demethylating enzyme or DNA repair machinery. To determine the percentage of cells present in different stages of the cell cycle, we conducted flow cytometry analysis (Supplementary Figure S2). Prolonged treatment with 10 mM VPA for 72 h increased the percentage of G1/G0 cells by 9% (92±1% of the total cells), paralleled by a reduction of cells in the S- and G2/M-phase (2±1% and 6±1% of the total cells, respectively). The cell-cycle data illustrate that very few of the untreated astrocytes were in the S-phase and VPA reduced this number even further. Thus, it seems very unlikely that the VPA-mediated reduction in DNA methylation by 18% (LUMA) can be accounted for by the few dividing cells. Thus, VPA might rather trigger active demethylation processes as already claimed by other groups for eg, HEK293 cells (Detich et al, 2003).
Amitriptylin Induces Genome-Wide CCpGG Hypomethylation Independent of HDAC Inhibition
We further examined whether drugs that do not possess HDAC inhibitory activities might also alter the degree of DNA methylation. For that purpose, we chose AMI from the group of antidepressants, and LTG as a mood stabilizer, and treated cortical astrocytes with 100 μM LTG for 72 h and with 1 or 10 μM AMI (for 24 and 72 h). Surprisingly, AMI (1 μM) exposure led to a significant decrease of methylation at CCpGG sites after 24 and 72 h (−7.4±4.2% and −9.7±4.0%, respectively) (Table 2). A higher dose was also effective after 72 h of the treatment (−9.2±2.0%). In contrast, LTG exposure did not produce changes of DNA methylation. Thus, AMI induced significant DNA hypomethylation (Table 2) in the absence of global histone hyperacetylation (Figure 1), which was partially reversible after drug withdrawal.
Total Cytosine Methylation is not Altered by Drug Exposure
Of all CpG sites in the genome, CCpGG tetranucleotides represent only a minor part of approximately 7–8%, where 35.25% of these sites are located in transposable elements, 64% in unique sequences, from which 14% are in CpG islands, as calculated for the mouse genome (Fazzari and Greally, 2004). As LUMA exclusively detects CCpGG sites, we additionally performed capillary electrophoresis analysis covering the methylation status of all CpG sites. The equivalent LUMA samples of (1) the time course of high dose VPA and (2) the 72 h time point of high dose AMI were subjected to capillary electrophoresis analysis. Strikingly, differences in total cytosine methylation levels between control and treated astrocytes could not be detected by capillary electrophoresis (Supplementary Table 1). Possibly, smaller changes in cytosine methylation that might not be evenly distributed over the whole genome, but rather located at certain CCpGG-rich DNA regions, could be better resolved by LUMA than by the capillary electrophoresis technique.
Finally, genomic DNA derived from cortical astrocytes treated with VPA (10 mM), TSA (0.2 μM), AMI (10 μM), and LTG (100 μM) for 72 h showed no differences in the average methylation levels of total LINE-1 CpGs sites compared with control cells (Supplementary Figure S3a-c). Together with the results obtained from capillary electrophoresis and LUMA analysis, we propose that VPA, TSA, and AMI do not cause ample DNA hypomethylation randomly at any CpG site of the genome, but rather at distinct regions that are likely enriched in CCpGG tetranucleotides and prone to DNA demethylating events.
Modulation of the Epigenetic Signature of an Astrocyte-Specific Promoter by VPA and TSA
On the basis of our observations that VPA triggers global histone hyperacetylation and DNA hypomethylation events, and that AMI, to a certain extent, influences the genomic DNA methylation status as well, we speculated whether genes crucial for the physiology of astrocytes are among the affected loci. As a paradigm, we chose the glutamate transporter subtype, GLT-1 (EAAT2, human homolog), the gene product ensuring low resting glutamate concentrations in the synaptic cleft (Choi, 1988; Rothstein et al, 1996). Previously, we collected data of an epigenetic component of human EAAT2 regulation (Zschocke et al, 2007), and there is increasing evidence that a disturbed glutamatergic neurotransmission in psychiatric disorders might also be affected by pharmacological interventions (Hashimoto et al, 2007; Javitt, 2004).
Hence, we determined the methylation status of the rat GLT-1 gene covering a substantial part of the 5′-UTR and two parts of distinct promoter regions by bisulfite sequencing. The 5′-UTR of GLT-1 is composed of a classical CpG island (Zschocke et al, 2005). Sequencing of approximately two-thirds of the CpG island (Figure 3a) showed that all inspected CpG dinucleotides were unmethylated in untreated cortical astrocytes. Furthermore, a region outside the CpG island located ∼900 bp upstream of the ATG start codon was also completely unmethylated (data not shown). Nevertheless, a region ∼1950 bp relative to the ATG start codon that contains putative binding sites for two chromatin-associated transcriptional regulators, proofed to be differentially methylated before and after drug exposure. Three CpG sites at positions −1978, −1958, and −1929 were methylated to 34±1%, 33±1%, and 36±2%, respectively, in untreated cells, as measured by direct bisulfite sequencing (data not shown). A 72 h treatment with 10 mM VPA led to a reduction of methylation at all three sites (−18.3±7.8%, −16.8±0.3%, and −27.7±4.6%, respectively) (Figure 3b). Similarly, TSA noticeably decreased cytosine methylation by −27.7 to −32.5%. In contrast to VPA/TSA, AMI (10 μM, 72 h) did not induce significant changes at any of the examined CpG dinucleotides. Similarly, LTG (100 μM, 72 h) did not change the methylation at positions −1978 and −1958, although a small increase in cytosine methylation was detected at position −1929 (+9.2±1.8%). Our findings show that global CCpGG DNA hypomethylation induced by VPA, TSA, and AMI have differential impacts at the single gene level.
Epigenetic modification of the GLT-1 promoter of rat cortical astrocytes by VPA and TSA treatment. (a) The scheme depicts the 5′-UTR of GLT-1 and adjacent promoter region. Empty circles represent unmethylated CpG regions, whereas filled circles represent moderately methylated individual CpGs (∼34%), as measured in untreated cortical astrocytes. The genomic sequence of GLT-1 distal promoter region is shown and putative binding sites for myb-like transcriptional regulator (myb TR) and for nuclear factor-1 (NF-1) are depicted. (b) CpG methylation changes were monitored at individual CpG sites (−1978, −1958, and −1929) after drug exposure. Cortical astrocytes were treated for 72 h with 10 mM VPA, 0.2 μM TSA, 10 μM AMI, and 100 μM LTG. After treatment, genomic DNA was isolated, bisulfite-converted, PCR amplified, and submitted to direct sequencing. Bars represent percentage change of CpG methylation (mean±SE, n=3) compared with untreated cells; t-test: *p<0.05 vs control, **p<0.005 vs control, ***p<0.0005 vs control. (c) Cortical astrocytes were exposed to 10 mM VPA for 72 h, genomic DNA was isolated, fragmented and immunoprecipitated using an anti-AcH4 antibody or IgG control. Precipitated DNA was subjected to real-time PCR analysis with a GLT-1 promoter-specific set of primers. AcH4 enrichment is depicted as fold-increase±SE (n=2) compared with untreated cells (set to 1). The results are normalized to input; t-test: ***p<0.0005 vs control.
To further characterize the epigenetic signature at the GLT-1 promoter after drug exposure, we investigated the composition of acetylated histone H4 at the differentially methylated promoter stretch by chromatin immunoprecipitation (ChIP). Indeed, we observed a 3.5-fold enrichment of acetylated H4 at the GLT-1 promoter in VPA (10 mM, 72 h) treated cortical astrocytes, as compared with control cells (Figure 3c).
HDAC-Is Increase GLT-1 Gene Transcription in Cortical Astrocytes
As VPA and TSA treatment resulted in an altered epigenetic composition of the GLT-1 promoter that potentially leads to a more relaxed chromatin structure, we tested whether these compounds are able to enhance GLT-1 mRNA transcription. In untreated cortical astrocytes, GLT-1 mRNA was detected at fairly moderate levels as determined by real-time PCR. After stimulation with VPA (low millimolar range) or TSA (low nanomolar range) for 24 h, we observed a dose dependent ∼4-fold increase of GLT-1 mRNA, respectively (Figure 4a and b). Moreover, neither AMI nor LTG influenced GLT-1 mRNA expression (data not shown). A reporter gene assay with the human GLT-1 promoter ortholog coupled to a luciferase gene confirmed the activation of the promoter by VPA and TSA, but not by LTG or AMI treatment (Figure 4c). This alludes to a role of transacting factors that might be triggered by VPA/TSA, as chromatin modifications at the reporter plasmids might be less relevant. We further tested whether the HDAC-I NaB also exerts an effect on GLT-1 expression in cortical astrocytes. Application of 1 mM NaB increased GLT-1 mRNA transcript levels after 72 h, but not at 24 h of treatment as determined by real-time PCR analysis (Figure 4d). Compared with VPA and TSA, this regulatory effect was weaker with regard to the kinetics of gene activation. In conclusion, CpG demethylation and histone hyperacetylation of a distinct region of the GLT-1 promoter induced by VPA or TSA was associated with an increased transcriptional activity of the gene. Conversely, AMI and LTG, failing to modify epigenetic marks at the promoter, did not enhance the basal transcription of GLT-1.
HDAC inhibitors VPA and TSA increase GLT-1 gene transcription in a dose-dependent manner. GLT-1 transcript levels were quantified by real-time PCR after 24 h treatment with 2.5, 5, and 10 mM VPA (a) and with 5, 15, or 30 nM TSA (b) and 24 and 72 h after 1 mM NaB treatment (d). Real-time PCR analysis was carried out with total RNA derived from treated and untreated (Co) cortical astrocytes and with GLT-1- and actin-specific primers. (control is set to 1; bars depict mean±SE (n=3) of the fold-increase; t-test: *p<0.05 vs control, **p<0.005 vs control.) (c) Reporter gene assays were conducted to directly assess promoter activity. Cortical primary astrocytes were transiently transfected with reporter plasmid containing a homolog of the human GLT-1 promoter coupled to a luciferase gene. The cells were cotransfected with nonsecretory Gaussia luciferase expression vector for normalization. The luciferase activity was assayed 24 h after applying drugs at the indicated concentrations. Bars present mean±SE (n=3–5) of the fold-increase compared with untreated cells (set to 1); t-test: *p<0.05 vs control.
Cellular DNMT-1 and GADD 45α Protein Levels are Unaffected by VPA/TSA and AMI
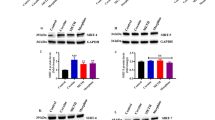
The molecular mechanisms of DNA demethylation are still unclear. We focused our further analysis on two key factors that are responsible for propagation, maintenance, or de novo establishment of DNA methylation patterns, namely DNMT-1 and GADD 45α (growth arrest and DNA damage-inducible protein 45α). In cortical astroglial cells, treatment with 1 and 10 mM VPA or with 200 nM TSA for 24 and 72 h, did not significantly decrease the levels of DNMT-1 protein under either condition (Figure 5a). In addition, AMI did not influence DNMT-1 levels under the specified treatment regiment (Figure 5b). Recently, it has been proposed that GADD 45α might have an active role in demethylating DNA (Barreto et al, 2007). As shown in Figure 5a and b, VPA/TSA and AMI did not robustly alter GADD 45α expression levels, although TSA slightly reduced protein levels after prolonged exposure. Our data imply that global changes in DNA methylation are not correlated with major differences in DNMT-1 or GADD 45α protein levels after VPA/TSA and AMI administration.
AMI, but not VPA, reduces DNA methyltransferase activity of cortical astrocytes independent of DNMT-1 downregulation. (a, b) Drug treatment has no impact on DNMT-1 and GADD 45α expression levels. Cortical astroglial cells were treated with VPA, AMI, and TSA ranging from 24 to 72 h, including drug washout as specified. Western blot analysis was carried out using cellular extracts prepared from cortical astrocytes, which were probed with DNMT-1-, GADD 45α-, α-actin-, or hsp90-specific antibodies. (c) Cells were treated with VPA (10 mM), AMI (1,10 μM), and 5-Aza (8 μM) for 3d or 6–11d (5-Aza, long-term). Nuclear extracts were isolated and supplemented with the respective drugs. DNA methyltransferase activity was quantified by measuring the incorporation of S-adenosyl-L-[methyl-3H]methionine into the substrate Poly(dI-dC)·Poly(dI-dC). Data are presented as relative DNA methyltransferase activity±SE (n=2 for VPA/5-Aza, n=4 for AMI). Control is set to 100%.
DNMT Activity is Differentially Affected by VPA and AMI
To test the possibility that DNMT activity is directly modified by VPA or AMI, we performed an in vitro methylation assay using poly dIdC DNA as template and tritium-labeled S-adenosyl-methionine (3H-SAM) as substrate. Nuclear extracts of VPA (10 mM, 72 h)- and AMI (1, 10 μM; 72 h)-treated cortical astrocytes were subjected to methylation reaction, and the amount of incorporated 3H-SAM was quantified (Figure 5c). In parallel, aliquots of nuclear extracts were analyzed for DNMT-1 levels by western blot (data not shown). VPA did not reduce the amount of incorporated SAM, nor were nuclear DNMT protein levels significantly reduced in the nuclear extracts, indicating that VPA does not impair DNMT function. Conversely, 10 μM AMI led to a dramatic reduction in DNMT enzymatic activity of ∼55% (Figure 5c), concomitantly with unchanged DNMT-1 levels (data not shown). Application of the specific DNMT inhibitor 5-aza-deoxycytidine (8 μM; 3–11 days) reduced the activity of DNMT by approximately 50–90%, which proofs the reliability of the assay to monitor DNMT activity changes. Altogether, our results indicate different modes of actions by VPA and AMI to reduce DNA methylation.
VPA and AMI Differentially Modulate Histone H3K9 Methylation Marks
So far, our results endorsed the idea of differing actions of VPA and AMI with regard to histone acetylation patterns and induction of GLT-1 expression. To corroborate these findings with other putative differences, we focused on further histone marks, the dimethylation of histone H3 at lysine residues 9 (2MeH3K9) and 27 (2MeH3K27). Psychoactive drugs possess the potential to influence the level of histone methylation in vivo (Huang et al, 2007; Tsankova et al, 2006; Wilkinson et al, 2009). We exposed cortical and hippocampal astrocytes to VPA (1 and 10 mM) and AMI (1 and 10 μM) for 24 h, and performed western blot analysis using 2MeH3K9- and 2MeH3K27-specific antibodies. VPA slightly, but significantly, decreased the levels of 2MeH3K9, whereas AMI showed a minor and nonsignificant increase of this repressive histone mark (Figure 6a and b). The applied drugs exerted no effect on 2MeH3K27 in both brain regions (Figure 6c and d). It is noteworthy that although the slight increase in 2MeH3K9 levels induced by NaB (applied at 0.1 and 1 mM for 24 h) was not significant compared with controls, these levels were significantly different from the reduced levels in response to the structurally related drug, VPA (Figure 6a and b). In this context, it is worth mentioning that NaB exposure did not result in global CCpGG methylation changes, although both substances VPA and NaB increased acetylation of H3 and H4 to a similar extent (data not shown). Collectively, our data support the idea that VPA and AMI affect different chromatin components as reflected at the level of 2MeH3K9 or AcH3/4. Furthermore, also structurally similar HDAC-I like VPA and NaB do not necessarily match in their epigenetic modes of action. This is in agreement with other studies showing various HDAC-I differentially affecting target genes (Hauke et al, 2009).
Differential effects of VPA and AMI on H3K9 dimethylation levels. Cortical (CTX) and hippocampal (HPC) astrocytes were treated with VPA (1 and 10 mM), AMI (1 and 10 μM) and NaB (0.1 and 1 mM) for 24 h. Western blot analysis was performed using antibodies directed against (a, b) dimethyl-H3K9 (α-2MeH3K9) or (c, d) dimethyl-H3K27 (α-2MeH3K9). As a control for protein loading, α-actin-specific antibody was used. Data represent mean±SE (n=3) of the percentage of optical band densities normalized to α-actin immunoreactivity. Control is set to 1; t-test: *p<0.05.
DISCUSSION
There is mounting evidence that psychiatric illnesses, major depression in particular, are associated with compromised glial function. For example, a reduced number of glial cells was reported in the prefrontal, orbitofrontal, and cingulate cortex (Cotter et al, 2001; Rajkowska et al, 1999). A number of animal studies replicated similar findings, for example, describing a 25% reduction of glial cells in the hippocampal formation of psychosocially stressed male tree shrews (Czeh et al, 2006). It is noteworthy that fluoxetin treatment for 28 days prevented the numerical decrease in astrocytes of the animals. Hence, abnormal astrocytic function might add to the pathogenesis and progression of mood disorders by deteriorating neuroplasticity and related processes. It has become increasingly evident, that beside neurons being the primary targets of antidepressant/mood stabilizer action, astrocytes also respond to these treatments (Manev et al, 2003; Pavone and Cardile, 2003).
In this study, we evaluated the effects of CIT, AMI, VEN, as well as VPA, LTG, and CBZ on various aspects of astroglial epigenetic parameters. For mechanistic insight, we also applied broad-spectrum HDAC-I, including TSA and NaB. With regard to HDAC subtypes, TSA is generally considered to inhibit both class I and class II HDACs, whereas VPA and NaB are preferentially class I inhibitors (Guardiola and Yao, 2002; Gurvich et al, 2004). Of all psychoactive drugs tested, only VPA led to a pronounced and transient global hyperacetylation of histones H3 and H4, paralleled by a small and transient demethylation of genomic CCpGG tetranucleotides. These results are consistent with previous studies showing that VPA directly inhibits the catalytic center of HDAC 2 with an EC50 value of 0.52 mM and of HDAC 5/6 with an EC50 of 2.4 mM (Gottlicher et al, 2001). HDAC-Is such as MS-275 and TSA also diminish DNMT-1, −3a and −3b protein levels and enzymatic activity in NT-2 precursor cells, tentatively explaining the mechanism of DNA demethylation events (Kundakovic et al, 2009). In primary astrocytes, we found no changes in DNMT-1 levels upon VPA administration. GADD 45α, a protein involved in cell-cycle control and DNA repair, represents a candidate as a mediator for active DNA demethylation as the overexpression of GADD 45α activates methylation-silenced reporter plasmids and promotes global DNA demethylation. VPA upregulates GADD 45α in N1E-115 neuroblastoma cells, thereby inducing growth arrest (Yamauchi et al, 2007), whereas in astroglial cells, GADD 45α levels were not elevated after VPA and TSA exposure in our experimental setup. Hence, we conclude that mechanisms other than DNMT-1 downregulation and GADD 45α upregulation are accountable for VPA-mediated CCpGG demethylation in quiescent cells. It is noteworthy that LINE-1 retrotransposons were spared from demethylation events, and methylation changes were not measurable at total cytosine residues, possibly due to resolution limits of capillary electrophoresis. In that context, VPA presumably does not reactivate silenced retrotransposons, which would harbor a mutagenic risk for the cell, and also does not mediate the demethylation of the gross of genomic CpG sites. We further observed small brain region-specific differences in responsiveness to DNA demethylation triggered by VPA, with generally lower magnitudes in the hippocampus.
Although it has been shown that CBZ inhibits representatives of class I and class II HDACs leading to slightly increased AcH4 levels in human HepG2 and HEK cell lines (Beutler et al, 2005), CBZ did not exert similar effects in primary astrocytes. Possibly, cells treated with CBZ need to actively proliferate to be more susceptible for these minor acetylation changes, as opposed to VPA. Moreover, LTG did not mediate any changes in the global histone acetylation profile in astrocytes. Similarly, acutely administered antidepressants CIT, AMI, and VEN did not show any significant influence on the global H3 and H4 acetylation status in hippocampal and cortical astrocytes. At this point in time, we cannot exclude the possibility that chronic administration of the drugs over several weeks might, nevertheless, change the degree of histone acetylation. Daily i.p. injections of fluoxetine, a substance related to CIT, over 10 days leads to an increase in HDAC2 immunoreactivity in coronal tissue sections, accompanied by reduction in global acetylated histone H3 (Cassel et al, 2006). Furthermore, chronic, but not acute imipramine treatment of socially stressed animals induces hyperacetylation of H3 at distinct regions of the BDNF gene potentially mediated by the downregulation of HDAC5 expression.
Surprisingly, although AMI did not display any HDAC inhibitory properties, it induced DNA hypomethylation at CCpGG sites in cortical astrocytes, similarly to VPA. This observation raised the possibility that DNA hypomethylation does not depend on histone hyperacetylation. Indeed, DNMT enzymatic activity was markedly reduced in AMI-treated astroglial cells from the cortex. To our knowledge, only very few studies deal with psychoactive drug-induced changes of the global degree of DNA methylation. For instance, 3-weeks administration of haloperidol, an antipsychotic drug, to rats was documented to result in a small, but significant reduction in total methyl-cytosine content (Shimabukuro et al, 2009).
DNA methylation and histone acetylation are complemented by combinatorial monomethylation, dimethylation, or trimethylation of histones H3/H4 to form a complex epigenetic code-controlling gene transcription (Lachner and Jenuwein, 2002). There exists a functional interplay of enzymes establishing various histone and DNA modifications. For instance, HDAC and H3K9 HMT G9a are recruited to chromatin sites together with transcriptional coactivators (Roopra et al, 2004), and DNMT-1 was reported to interact with SUV39H1, a histone H3K9 methyltransferase (Fuks et al, 2003). Our data show that VPA exposure caused a slight but specific reduction in the levels of dimethyl-H3K9, leaving dimethyl-H3K27 unchanged. In line with our results, HDAC-I desipeptide decreases the level of H3K9 methylation and induces DNA demethylation by reducing G9A and SUV39H1 activity and DNMT-1 binding, respectively (Wu et al, 2008b). In contrast to VPA, AMI did not change dimethyl-H3K9 levels.
Global epigenetic alterations caused by VPA hit a selected gene promoter, the activity of which is crucial for astrocytic functionality, GLT-1. At glutamatergic synapses, GLTs recycle released extracellular glutamate back into astrocytes to terminate postsynaptic and presynaptic receptor activation (Schousboe and Waagepetersen, 2006; Vandenberg, 1998). Distinct parts of the GLT-1 promoter were demethylated and enriched in acetylated histone H4 after VPA treatment, whereas the CpG island in proximity of the translational start site was completely unmethylated in treated and untreated cells. Concomitantly, the activity of the GLT-1 gene was profoundly increased, analogously to the actions of TSA on GLT-1 gene transcription, indicating that HDAC inhibitory properties of VPA are attributable to the documented effects. The differentially methylated promoter region contains putative consensus sequences (as predicted by MatInspector software) for the recruitment of two chromatin regulatory proteins, namely nuclear factor 1 and myb-like transcriptional regulator (Chikhirzhina et al, 2008; Ko et al, 2008). A number of epigenetically regulated genes such as myoD are differentially methylated at distal enhancer elements, whereas CpG islands close to transcription start sites are protected from methylation (Brunk et al, 1996) and, hence, are less susceptible to epigenetic alterations.
Chronic VPA exposure increases GLT-1 expression in vivo in the rodent hippocampus (Hassel et al, 2001). In addition, I-phenylpyridinium-challenged primary astrocytes, simultaneously treated with TSA, showed improvement in glutamate uptake as reported by Wu et al (2008a). Very recently, Allritz et al (2009) showed stimulatory effects of TSA on GLT-1 mRNA and protein levels in rat astrocytes of various brain regions. We complement the findings that HDAC-I likely induces GLT-1 expression in part by chromatin changes at the promoter region, probably in concert with the induction of transacting factors. Conversely, AMI neither targeted GLT-1 gene transcription nor the respective CpG sites within the GLT-1 promoter.
Taken together, we could show for the first time that of all tested psychoactive substances, VPA exerted the strongest influence on chromatin remodeling events in astrocytes involving CpG demethylation and histone modifications, targeting GLT-1, an astroglial gene important for glutamate homeostasis in the CNS. Similarly, AMI also brought about changes in global DNA methylation, possibly by functional impairment of DNMTs, without targeting the GLT-1 gene (Figure 7). The underlying molecular mechanisms and biological conditions including different neural cell types and brain regions of AMI-affected DNMT-1 function are yet to be determined. It will be interesting to uncover the common and divergent sites affected by the global demethylating actions by these two drugs. It also remains to be seen whether other tricyclic and SNRI/SSRI drugs show similar effects on the DNA methylation machinery, and whether these effects might contribute to the recovery of disturbed neural processes.
Proposed model for VPA- and AMI- mediated epigenetic alterations in astrocytes. VPA and AMI display an input on global DNA demethylation, although through different mechanisms. As VPA inhibits HDACs, and AMI impairs DNMT function, the measurable difference in global epigenetic marks consists of hyperacetylation of histones in the case of VPA, which are not affected by AMI. Locally, downstream pathways further diverge as exemplified for the GLT-1 gene. The biochemical composition of the promoter is markedly altered by VPA exposure by means of histone hyperacetylation and DNA demethylation, resulting in an enhanced gene transcription. AMI does not possess similar features, and subsequently leaves gene activity unaffected. Round circles represent CpG sites; shading reflects the degree of methylation; oval circles depict core histones.
In conclusion, a promising new approach for the pharmacological treatment of psychiatric illnesses could be based on the reversal of aberrantly established epigenetic patterns. Two principle strategies could guide to the future objective: first, pharmacological intervention directly interferes with epigenetic machinery as shown in this study for VPA and AMI, resulting in overall changes in the epigenetic pattern of chromatin. In fact, HDAC-I NaB exerts antidepressant-like effects (Schroeder et al, 2007) and improves cognitive performance in mice (Fischer et al, 2007). Apparently, this strategy holds the risk of reactivating genes in a false spatial-temporal context. The parameter ensuring that only a subset of crucial genes are reactivated might rely on the composition of transcription factors governing the transcriptome characteristic for a specific cell type, such as an astrocyte or neuron. The second strategy comprises the indirect modulation of epigenetic marks either globally or locally by interfering with neurotransmitter and signal transduction pathways, ultimately resulting in epigenetically based and permanently changed target gene expression patterns. In general, identifying novel HDAC-I- or DNMT-1-regulated genes acting in support of neural circuits might provide a starting point for the development of new pharmacological treatment designs.
References
Allritz C, Bette S, Figiel M, Engele J (2009). Endothelin-1 reverses the histone deacetylase inhibitor-induced increase in glial glutamate transporter transcription without affecting histone acetylation levels. Neurochem Int 55: 22–27.
Azmitia EC (2001). Modern views on an ancient chemical: serotonin effects on cell proliferation, maturation, and apoptosis. Brain Res Bull 56: 413–424.
Barreto G, Schafer A, Marhold J, Stach D, Swaminathan SK, Handa V et al (2007). Gadd45a promotes epigenetic gene activation by repair-mediated DNA demethylation. Nature 445: 671–675.
Beutler AS, Li SD, Nicol R, Walsh MJ (2005). Carbamazepine is an inhibitor of histone deacetylases. Life Sci 76: 3107–3115.
Brunk BP, Goldhamer DJ, Emerson CP (1996). Regulated demethylation of the MyoD distal enhancer during skeletal myogenesis. Develop Biol 177: 490–503.
Cassel S, Carouge D, Gensburger C, Anglard P, Burgun C, Dietrich JB et al (2006). Fluoxetine and cocaine induce the epigenetic factors MeCP2 and MBD1 in adult rat brain. Mol Pharmacol 70: 487–492.
Chen PS, Peng GS, Li G, Yang S, Wu X, Wang CC et al (2006). Valproate protects dopaminergic neurons in midbrain neuron/glia cultures by stimulating the release of neurotrophic factors from astrocytes. Mol Psychiatr 11: 1116–1125.
Chen WG, Chang Q, Lin YX, Meissner A, West AE, Griffith EC et al (2003). Derepression of BDNF transcription involves calcium-dependent phosphorylation of MeCP2. Science 302: 885–889.
Chikhirzhina GI, Al-Shekhadat RI, Chikhirzhina EV (2008). Transcription factors of the NF1 family: role in chromatin remodeling. Mol Biol 42: 342–356.
Choi DW (1988). Glutamate neurotoxicity and diseases of the nervous-system. Neuron 1: 623–634.
Cotter D, MacKay D, Landau S, Kerwin R, Everall I (2001). Reduced glial cell density and neuronal size in the anterior cingulate cortex in major depressive disorder. Arch Gen Psychiatr 58: 545–553.
Czeh B, Simon M, Schmelting B, Hiemke C, Fuchs E (2006). Astroglial plasticity in the hippocampus is affected by chronic psychosocial stress and concomitant fluoxetine treatment. Neuropsychopharmacology 31: 1616–1626.
D'Alessio AC, Szyf M (2006). Epigenetic tete-a-tete: the bilateral relationship between chromatin modifications and dna methylation. Biochem Cell Biol-Biochim Biol Cell 84: 463–476.
Detich N, Bovenzi V, Szyf M (2003). Valproate induces replication-independent active DNA demethylation. J Biol Chem 278: 27586–27592.
Dong E, Guidotti A, Grayson DR, Costa E (2007). Histone hyperacetylation induces demethylation of reelin and 67-KDa glutamic acid decarboxylase promoters. Proc Natl Acad Sci USA 104: 4676–4681.
Eyal S, Yagen B, Sobol E, Altschuler Y, Shmuel M, Bialer M (2004). The activity of antiepileptic drugs as histone deacetylase inhibitors. Epilepsia 45: 737–744.
Fazzari MJ, Greally JM (2004). Epigenomics: beyond CpG islands. Nat Rev Genet 5: 446–455.
Fischer A, Sananbenesi F, Wang XY, Dobbin M, Tsai LH (2007). Recovery of learning and memory is associated with chromatin remodelling. Nature 447: 178–1U2.
Franke B, Figiel M, Engele J (1998). CNS glia are targets for GDNF and Neurturin. Histochem Cell Biol 110: 595–601.
Fuks F, Hurd PJ, Deplus R, Kouzarides T (2003). The DNA methyltransferases associate with HP1 and the SUV39H1 histone methyltransferase. Nucleic Acids Res 31: 2305–2312.
Gottlicher M, Minucci S, Zhu P, Kramer OH, Schimpf A, Giavara S et al (2001). Valproic acid defines a novel class of HDAC inhibitors inducing differentiation of transformed cells. EMBO J 20: 6969–6978.
Guardiola AR, Yao TP (2002). Molecular cloning and characterization of a novel histone deacetylase HDAC10. J Biol Chem 277: 3350–3356.
Gurvich N, Tsygankova OM, Meinkoth JL, Kein PS (2004). Histone deacetylase is a target of valproic acid-mediated cellular differentiation. Cancer Res 64: 1079–1086.
Hashimoto K, Sawa A, Iyo M (2007). Increased levels of glutamate in brains from patients with mood disorders. Biol Psychiatr 62: 1310–1316.
Hassel B, Iversen EG, Gjerstad L, Tauboll E (2001). Up-regulation of hippocampal glutamate transport during chronic treatment with sodium valproate. J Neurochem 77: 1285–1292.
Hauke J, Riessland M, Lunke S, Eyupoglu IY, Blumcke I, El-Osta A et al (2009). Survival motor neuron gene 2 silencing by DNA methylation correlates with spinal muscular atrophy disease severity and can be bypassed by histone deacetylase inhibition. Hum Mol Genet 18: 304–317.
Herman JG, Baylin SB (2003). Mechanisms of disease: gene silencing in cancer in association with promoter hypermethylation. N Engl J Med 349: 2042–2054.
Huang HS, Matevossian A, Whittle C, Kim SY, Schumacher A, Baker SP et al (2007). Prefrontal dysfunction in schizophrenia involves mixed-lineage leukemia 1-regulated histone methylation at GABAergic gene promoters. J Neurosci 27: 11254–11262.
Inazu M, Takeda H, Ikoshi H, Sugiswa M, Uchida Y, Matsumiya T (2001). Pharmacological characterization and visualization of the glial serotonin transporter. Neurochem Int 39: 39–49.
Inazu M, Takeda H, Matsumiya T (2003). Functional expression of the norepinephrine transporter in cultured Rat astrocytes. J Neurochem 84: 136–144.
Javitt DC (2004). Glutamate as a therapeutic target in psychiatric disorders. Mol Psychiatr 9: 984–997.
Karimi M, Johansson S, Stach D, Corcoran M, Grander D, Schalling M et al (2006). LUMA (LUminometric Methylation Assay) – a high throughput method to the analysis of genomic DNA methylation. Exp Cell Res 312: 1989–1995.
Ko ER, Ko D, Chen C, Lipsick JS (2008). A conserved acidic patch in the Myb domain is required for activation of an endogenous target gene and for chromatin binding. Mol Cancer 7: 77.
Kundakovic M, Chen Y, Guidotti A, Grayson DR (2009). The Reelin and GAD67 promoters are activated by epigenetic drugs that facilitate the disruption of local repressor complexes. Mol Pharmacol 75: 342–354.
Lachner M, Jenuwein T (2002). The many faces of histone lysine methylation. Curr Opin Cell Biol 14: 286–298.
Manev H, Uz T, Manev R (2003). Glia as a putative target for antidepressant treatments. J Affect Disord 75: 59–64.
McGowan PO, Sasaki A, D'Alessio AC, Dymov S, Labonte B, Szyf M et al (2009). Epigenetic regulation of the glucocorticoid receptor in human brain associates with childhood abuse. Nat Neurosci 12: 342–348.
McGowan PO, Sasaki A, Huang TC, Unterberger A, Suderman M, Ernst C et al (2008). Promoter-wide hypermethylation of the ribosomal RNA gene promoter in the suicide brain. PLoS ONE 3: e2085.
Mill J, Petronis A (2007). Molecular studies of major depressive disorder: the epigenetic perspective. Mol Psychiatr 12: 799–814.
Ng HH, Bird A (1999). DNA methylation and chromatin modification. Curr Opin Genet Develop 9: 158–163.
Ou JN, Torrisani J, Unterberger A, Provencal N, Shikimi K, Karimi M et al (2007). Histone deacetylase inhibitor trichostatin a induces global and gene-specific DNA demethylation in human cancer cell lines. Biochem Pharmacol 73: 1297–1307.
Pavone A, Cardile T (2003). An in vitro study of new antiepileptic drugs and astrocytes. Epilepsia 44: 34–39.
Petronis A (2003). Epigenetics and bipolar disorder: new opportunities and challenges. Am J Med Genet Part C-Semin Med Genet 123C: 65–75.
Phiel CJ, Zhang F, Huang EY, Guenther MG, Lazar MA, Klein PS (2001). Histone deacetylase is a direct target of valproic acid, a potent anticonvulsant, mood stabilizer, and teratogen. J Biol Chem 276: 36734–36741.
Poulter MO, Du LS, Weaver ICG, Palkovits M, Faludi G, Merali Z et al (2008). GABA(A) receptor promoter hypermethylation in suicide brain: implications for the involvement of epigenetic processes. Biol Psychiatr 64: 645–652.
Rajkowska G, Miguel-Hidalgo JJ, Wei JR, Dilley G, Pittman SD, Meltzer HY et al (1999). Morphometric evidence for neuronal and glial prefrontal cell pathology in major depression. Biol Psychiatr 45: 1085–1098.
Roopra A, Qazi R, Schoenike B, Daley TJ, Morrison JF (2004). Localized domains of G9a-mediated histone methylation are required for silencing of neuronal genes. Mol Cell 14: 727–738.
Rothstein JD, DykesHoberg M, Pardo CA, Bristol LA, Jin L, Kuncl RW et al (1996). Knockout of glutamate transporters reveals a major role for astroglial transport in excitotoxicity and clearance of glutamate. Neuron 16: 675–686.
Schipke CG, Kettenmann H (2004). Astrocyte responses to neuronal activity. Glia 47: 226–232.
Schousboe A, Waagepetersen HS (2006). Glial modulation of GABAergic and glutamat Ergic neurotransmission. Curr Topics Med Chem 6: 929–934.
Schroeder FA, Lin CL, Crusio WE, Akbarian S (2007). Antidepressant-like effects of the histone deacetylase inhibitor, sodium butyrate, in the mouse. Biol Psychiatr 62: 55–64.
Sharma RP, Tun N, Grayson DR (2008). Depolarization induces downregulation of DNMT1 and DNMT3a in primary cortical cultures. Epigenetics 3: 74–80.
Shimabukuro M, Jinno Y, Fuke C, Okazaki Y (2009). Haloperidol treatment induces tissue- and sex-specific changes in DNA methylation: a control study using rats. Behav Brain Funct 2: 2.
Szyf M, McGowan P, Meaney MJ (2008). The social environment and the epigenome. Environ Mol Mutagen 49: 46–60.
Tsankova NM, Berton O, Renthal W, Kumar A, Neve RL, Nestler EJ (2006). Sustained hippocampal chromatin regulation in a mouse model of depression and antidepressant action. Nat Neurosci 9: 519–525.
Vandenberg RJ (1998). Molecular pharmacology and physiology of glutamate transporters in the central nervous system. Clin Exp Pharmacol Physiol 25: 393–400.
Weaver ICG, Cervoni N, Champagne FA, D'Alessio AC, Sharma S, Seckl JR et al (2004). Epigenetic programming by maternal behavior. Nat Neurosci 7: 847–854.
Wilkinson MB, Xiao G, Kumar A, LaPlant Q, Renthal W, Sikder D et al (2009). Imipramine treatment and resiliency exhibit similar chromatin regulation in the mouse nucleus accumbens in depression models. J Neurosci 29: 7820–7832.
Wu JY, Niu FN, Huang R, Xu Y (2008a). Enhancement of glutamate uptake in 1-Methyl-4-Phenylpyridinium-treated astrocytes by Trichostatin A. Neuroreport 19: 1209–1212.
Wu LP, Wang X, Li L, Zhao Y, Lu SL, Yu Y et al (2008b). Histone deacetylase inhibitor depsipeptide activates silenced genes through decreasing both CpG and H3K9 methylation on the promoter. Mol Cell Biol 28: 3219–3235.
Wu XF, Chen PS, Dallas S, Wilson B, Block ML, Wang CC et al (2008c). Histone deacetylase inhibitors up-regulate astrocyte GDNF and bdnf gene transcription and protect dopaminergic neurons. Int J Neuropsychopharmacol 11: 1123–1134.
Yamauchi J, Miyamoto Y, Murabe M, Fujiwara Y, Sanbe A, Fujita Y et al (2007). Gadd45a, the gene induced by the mood stabilizer valproic acid, regulates neurite outgrowth through JNK and the substrate paxillin in N1E-115 Neuroblastoma cells. Exp Cell Res 313: 1886–1896.
Yang AS, Estecio MRH, Doshi K, Kondo Y, Tajara EH, Issa JPJ (2004). A simple method for estimating global DNA methylation using bisulfite PCR of repetitive DNA elements. Nucleic Acids Res 32: e38.
Zschocke J, Allritz C, Engele J, Rein T (2007). DNA methylation dependent silencing of the human glutamate transporter EAAT2 gene in glial cells. Glia 55: 663–674.
Zschocke J, Bayatti N, Clement AM, Witan H, Figiel M, Engele J et al (2005). Differential promotion of glutamate transporter expression and function by glucocorticoids in astrocytes from various brain regions. J Biol Chem 280: 34924–34932.
Acknowledgements
This study was in part supported by the NARSAD Young Investigator Award and by the Horst Kübler foundation. We are grateful to Barbara Berning for excellent technical support. We thank T. Jenuwein for generously providing methyl-histone antibodies.
Author information
Authors and Affiliations
Corresponding author
Additional information
DISCLOSURE
The authors declare no conflict of interest.
Supplementary Information accompanies the paper on the Neuropsychopharmacology website (http://www.nature.com/npp)
Supplementary information
Rights and permissions
About this article
Cite this article
Perisic, T., Zimmermann, N., Kirmeier, T. et al. Valproate and Amitriptyline Exert Common and Divergent Influences on Global and Gene Promoter-Specific Chromatin Modifications in Rat Primary Astrocytes. Neuropsychopharmacol 35, 792–805 (2010). https://doi.org/10.1038/npp.2009.188
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/npp.2009.188
Keywords
This article is cited by
-
Tricyclic antidepressants target FKBP51 SUMOylation to restore glucocorticoid receptor activity
Molecular Psychiatry (2022)
-
Differential and paradoxical roles of new-generation antidepressants in primary astrocytic inflammation
Journal of Neuroinflammation (2021)
-
Neurobiology, Functions, and Relevance of Excitatory Amino Acid Transporters (EAATs) to Treatment of Refractory Epilepsy
CNS Drugs (2020)
-
The Effect of Chronic Mild Stress and Venlafaxine on the Expression and Methylation Levels of Genes Involved in the Tryptophan Catabolites Pathway in the Blood and Brain Structures of Rats
Journal of Molecular Neuroscience (2020)
-
Drugging the pain epigenome
Nature Reviews Neurology (2017)