Abstract
The combination of synthetic and systems biology is a powerful framework to study fundamental questions in biology and produce chemicals of immediate practical application such as biofuels, polymers, or therapeutics. However, we cannot yet engineer biological systems as easily and precisely as we engineer physical systems. In this review, we describe the path from the choice of target molecule to scaling production up to commercial volumes. We present and explain some of the current challenges and gaps in our knowledge that must be overcome in order to bring our bioengineering capabilities to the level of other engineering disciplines. Challenges start at molecule selection, where a difficult balance between economic potential and biological feasibility must be struck. Pathway design and construction have recently been revolutionized by next-generation sequencing and exponentially improving DNA synthesis capabilities. Although pathway optimization can be significantly aided by enzyme expression characterization through proteomics, choosing optimal relative protein expression levels for maximum production is still the subject of heuristic, non-systematic approaches. Toxic metabolic intermediates and proteins can significantly affect production, and dynamic pathway regulation emerges as a powerful but yet immature tool to prevent it. Host engineering arises as a much needed complement to pathway engineering for high bioproduct yields; and systems biology approaches such as stoichiometric modeling or growth coupling strategies are required. A final, and often underestimated, challenge is the successful scale up of processes to commercial volumes. Sustained efforts in improving reproducibility and predictability are needed for further development of bioengineering.
Similar content being viewed by others
Introduction
Although the synthesis of urea by Wohler in 18281 established that biological entities are not radically distinct from purely physical ones, we are still unable to design and engineer biological systems with the same ease and precision with which we design physical ones (e.g., cell phones, automobiles or jet planes). Whether biological systems can be understood as a mechanistic composition of physical parts is a fundamental philosophical and scientific problem, epitomized by the understanding of the brain and the emergence of consciousness.2 Furthermore, engineering of biological systems has emerged as one of the most exciting recent technologies. Applications for human health include tumor-killing bacteria, in vivo diagnostics, engineered viruses and immune cells that target specific disease cells, synthetic drug delivery vectors, and even completely synthetic tissues.3–5 Environmental applications include microbes that sense, report, and degrade toxic chemicals,6,7 while synthetic biology has the capability to produce a variety of chemical products ranging from therapeutics to plastics and biofuels.8–10
This last application of microbial bioengineering to produce biofuels and other commodity products has attracted significant attention due to its potential to mitigate climate change, reduce society’s reliance on fossil fuels, and improve energy security. Renewable chemical production is critical to achieving these goals, particularly given the needs of the growing populations of emerging economies.11 In 2004, a US government report energized the metabolic engineering community by providing a list of targets that would have a transformative impact on biomanufacturing,12 if they could be made using microbial platforms from sustainable carbon sources. Efforts in the last decade, aided by systems and synthetic biology, have succeeded in producing a variety of these and other products, typically at titers of μg/l to mg/l. However, production of only a few compounds, such as 1,3-propanediol,13 1–4-butanediol,14 and artemisinin15 has reached commercial scale, attesting to the difficulty of this process.16 Improving the yield, titer, and productivity of microbial processes to enable commercialization requires detailed manipulation of microbial physiology, stress response, and metabolism, with particular emphasis on carbon and energy balance. In the case of bulk commodities, where commercial viability requires capturing every available carbon atom and eliminating every unneeded ATP sink, metabolic engineering requires a systems biology17 approach where the interaction of the exogenous pathway with host metabolism can be explicitly considered and understood.
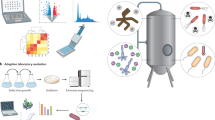
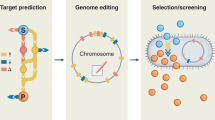
In this review, we present the difficulties of taking a microbial production process from conception to commercialization along with the tools that can be used to address some of the challenges and gaps in our knowledge and engineering capabilities (Figure 1). At a time when a significant acceleration of biological engineering is possible,18 but requires an influx of talent from a variety of engineering and physical sciences,19 this review presents the challenges that we think future metabolic engineers and synthetic biologists may want to address for fastest development of the field.
The process of bioengineering strains for commodity chemicals from initial concept (target molecule selection) to scale up (process engineering and implementation), along with a selection of tools applicable to each step and the grand challenges that need to be met. The lines, colored according to the type of tool/challenge, indicate which parts of the process the tool or challenge applies to (e.g., dynamic regulation can be used for pathway construction but also for control of toxic intermediates that affect host metabolism). Two of these tools and challenges are highlighted in the following figures (Figures 2 and 3). In spite of the linear diagram chosen to represent them, it must be understood that the process is rarely sequential: e.g., very often problems in engineering the process for scale up force researchers to go back to pathway construction and make significant changes.
Pathway design
Target molecule selection
The power of using microbial processes for chemical production is twofold: first, renewable carbon sources can serve as substrates, and second, the range and specificity of molecules that can be made biologically surpasses that of synthetic chemistry. The impressive selectivity of biological systems allows precise control over the chemical features of the final product, such as chirality and positioning of functional groups. This large range of molecules means that, while current efforts focus largely on well-characterized molecules, the most exciting applications will come when new molecules with unique chemical properties give rise to new materials, polymers, and fuels. Given the challenge in predicting chemical and material properties de novo, selecting promising future targets for biological production will require extensive collaboration between chemists, engineers and biologists, and the development of tools able to predict the bulk properties of materials composed of novel molecules.
Because of the difficulties of entering new chemical markets, most biological production has focused on molecules with large existing markets. Initial efforts focused on high-price, low-volume markets such as pharmaceuticals, where biological production could make an immediate and obvious impact.20 As interest has developed in high-volume, low-cost markets such as biofuels, economic considerations have become paramount, and development of a new project must begin with an analysis of the potential of process commercialization. This multi-disciplinary effort unites very disparate sets of knowledge. On one hand, the commercial potential of the molecule must be assessed. This potential depends not only on chemical properties and known performance but also on highly volatile market data: from the feedstock price (e.g., corn, sugar cane, or lignocellulose) to the current molecule price, both of which are tightly coupled to other market and political processes such as fluctuating oil prices, economic cycles, and uncertainties in government regulatory policies. On the other hand, the feasibility of producing the desired molecule biologically must be assessed, something which is still extremely difficult in the current state of bioengineering. Adding to these difficulties is the complexity of predicting the extra economic costs derived from scaling up production (described below) and downstream processing (e.g., molecule extraction and purification). All these factors make target molecule selection the least systematic part of the metabolic engineering process.
Due to the difficulty of assessing economic viability, metabolic engineers typically resort to a variety of available reports indicating high-value targets or intermediates as indicated by experts in the field,12,21–24 and rely on their own personal expertize to judge which of them can be biologically tractable. Currently, this approach works satisfactorily because troubleshooting the bioengineering effort takes significantly more time than choosing the target. However, as it becomes possible to biologically produce new types of molecules more efficiently,18 it would be desirable to create tools that combine technoeconomic analysis with market needs based on chemical properties, as well as systematic biological feasibility estimates and scale-up considerations.
Even in the cases where biomanufacturing has not achieved economic cost-competitiveness, it can help improve the sustainability of energy sources and other chemical commodity products. Despite the recent drop in oil and gas prices, 195 nations committed to reducing greenhouse gas emissions at the 2015 negotiations in Paris,25 and biomanufacturing provides a path toward a decarbonized, more sustainable economy. As countries across the world work to integrate the full social and environmental cost of fossil-derived fuels and products, bio-derived products will become increasingly attractive. The biomass-derived biofuel farnesene,26 e.g., is used to power part of the bus fleet in Sao Paulo, one of Brazil’s largest cities, even though the prize is higher than standard diesel: the city has pledged to reduce fossil diesel fuel use by 10% every year and are willing to subsidize higher prices to achieve this goal. Similarly, in the United States the establishment by congress of a renewable fuel standard27 that requires the use of 36 billion gallons of renewable fuel by 2022, provides the necessary incentives to pay for the offset in greenhouse gas emissions that renewable fuels provide as their main competitive advantage.
Gene discovery and pathway construction
Once a target is selected, the best production pathway needs to be identified. Historical methods for pathway discovery were limited to insights from experts in the field and manual gene selection that was informed by little functional data. More recently, computational methods based on biochemical reaction databases and genome mining have automated the process and implemented scoring methods to prioritize pathway selection. Developments in next-generation sequencing technology along with bioinformatic analyses have revolutionized gene discovery methods for metabolic engineering by providing a vast resource of genomic data to query for target genes. Many tools are available to use this data to identify biosynthetic gene clusters28,29 or select pathways based on homology-predicted enzyme function retro-synthesis of products.30–33 Yet, identification of gene function through homology comparison is quite poor for genes with unique functions, and functional screens are needed to identify specific steps in a desired pathway. Furthermore, experimental validation of predictions from these bioinformatic tools remains limited, so the success rate is unknown. Even when there is high confidence in the gene function, most retro-synthesis tools do not have the data needed to identify the organism containing the gene version that would yield optimal protein production and activity, so multiple gene variants must be tested to find the optimal enzyme for a given reaction step.
Pathway construction methods have been revolutionized by the advent of next-generation sequencing and affordable de novo DNA synthesis coupled with standardized expression vectors and genomic integration methods. Although traditional pathway construction is iterative and specific to the desired construct, these methods open the door to combinatorial pathway construction of vast libraries consisting of host genetic contexts, open reading frames, and/or protein expression variants. By producing a deep library of different pathway constructs a researcher can select a subset of strains either from pre-determined parameters or from design-of-experiment methods for subsequent testing and analysis. For systems with a well-defined high-throughput screen,34 selection, or biosensor system, large (107–1010) combinatorial libraries can be constructed, analyzed, and the identity of the best-performing strains can be determined by sequencing. This data driven workflow for strain construction and selection redistributes resources toward experimental design and data analysis efforts to reach production goals. Although these capabilities are ready to be deployed with traditional model systems such as Escherichia coli, they can also be used for many other microbes where synthetic biology and genetics are rapidly maturing, such as pseudomonads,35 autotrophs such as cyanobacteria,36 and Ralstonia eutropha.37 The challenges and opportunities to improve pathway construction, especially the automation of DNA design and construction, are described in several recent reviews38–40 and research efforts,41,42 and are not reviewed here.
Pathway optimization
Characterizing enzyme expression by proteomics
Merely introducing the genes into a host provides no guarantee that the synthetic pathway will function effectively. Often, the first troubleshooting steps involve ensuring expression of the introduced genes. Although reverse transcription–PCR or RNA sequencing can verify transcription, defects in translation or protein stability are more common, necessitating protein-level analysis. Consequently, development of rapid and accurate high-throughput assays to monitor protein expression can dramatically accelerate engineering efforts. Protein analysis is frequently achieved via immunoblot assays because they are selective and easily analyzed in parallel. However, assaying many different pathway proteins in the same strain can be challenging and quantification is often inaccurate and difficult to reproduce. Mass spectrometry-based proteomic methods have risen in popularity, as they can identify and quantify thousands of proteins. Relative quantification of proteins between different engineered strains is typically based on targeted proteomic methods via selected-reaction monitoring mass spectrometry, and takes advantage of isotopically labeled substrates or chemical tags (e.g., iTRAQ43 or TMT44) for accurate protein quantification.
The first reported targeted proteomics study of engineered E. coli identified protein-associated bottlenecks in the mevalonate pathway used to produce the sesquiterpene amorpha-4,11-diene.45 Here, levels of two proteins, mevalonate kinase (MK) and phosphomevalonate kinase (PMK), were particularly low. To overcome these bottlenecks the MK and PMK genes were codon-optimized for translation in E. coli and also expressed from a stronger promoter. These changes led to significant improvements in the protein levels and amorpha-4,11-diene production.
Aside from identifying pathway bottlenecks, comparative proteomics is also commonly used to quantify native host proteins, which can identify cellular stresses and perturbations to host metabolism. These may be based on targeted methods as above, or untargeted shotgun proteomic analyses via many “label-free” techniques.46 A number of recent studies have characterized cellular responses to potential biofuel products47–51 and identified metabolic sinks that impact carbon utilization.52 Such analysis will become even more critical as heterologous pathways increase in efficiency, demanding more cellular resources and imposing stresses and constraints on host metabolism. As current state-of-the-art methods still require relatively long chromatographic separation times (tens of minutes to hours) and are performed by using nano-flow chromatography, they are susceptible to variations in sample preparation and chromatography. As a result, these methods are not immediately suitable to high-throughput analysis and method development will need to address both throughput and data quality.
Optimizing expression levels
Product yield optimization can be achieved, in part, through fine-tuning of exogenous pathways in order to maximize the flux through the introduced pathway. Nonetheless, it is not straightforward to know in advance which pathway designs will produce the highest production. Ideally, the optimal expression level for each enzyme would be guided by accurate kinetic models of the pathway.53–57 However, this approach is hampered by a variety of challenges, such as lack of reliable data for enzyme activity and substrate affinity parameters, in vivo protein quantification, and the effects of activators and inhibitors. In spite of these hurdles, approaches that parameterize the kinetic model based on a subset of data via ensemble modeling58,59 have been successfully implemented, for instance to improve neurosporene productivity. In other studies, kinetic models have been used to pinpoint rate-limiting steps for hydrogen production.60 In the absence of accurate kinetic models, heuristic statistical approaches have been successfully used to guide product yield increases. Typically, predictions are extrapolated from data obtained from a relatively small (compared with the full combinatorial space) set of pathway designs. In this vein, Ajikumar et al.61 improved taxadiene production by dividing the pathway into different modules and performing a multivariate search, an approach that was expanded by the use of a linear regression model62 for amorphodiene production. A linear regression model was also used in order to relate protein expression levels to the production titers of the four primary products in the violacein pathway.63 Alonso-Gutierrez et al.64 used quantitative proteomics data and principal component analysis to increase limonene and bisabolene production, while George et al.65 used correlations between proteomic and metabolomic data to derive a qualitative model of the mevalonate pathway for isopentenol production. However, it must be recognized that for any of these methods, even if the desired expression level for maximum production is known, it is a non-trivial task to design the right pathway to obtain it.
In addition, optimizing expression levels is crucial if overexpression of key proteins imposes a large cellular burden due to toxicity. Although most observations of toxicity caused by protein expression relate to protein function, certain categories of proteins are more directly toxic to the cell. Examples include transport proteins that are important both for the import of carbon sources or nutrients, as well as exporters that are required to secrete final products or exclude harmful by-products left over from substrate processing.66,67 Overexpression of membrane proteins commonly leads to growth inhibition, likely due to altered membrane physical properties and limited expression and translocation of key proteins such as electron transport chain components, which has significant downstream impact on central carbon metabolism.68 These tradeoffs must be considered when expressing multiple membrane proteins during strain engineering.69 Protein expression burden can be addressed using directed evolution to obtain variants that perform efficiently within the range of expression that the strain accommodates70 or by controlling protein expression more dynamically in response to conditions under which the tolerance phenotype is required.71,72
Toxicity of pathway intermediates and the role of dynamic regulation
Natural metabolic pathways have evolved intricate mechanisms to avoid the formation of potentially toxic intermediates.73 As external pathways and enzymatic reactions are introduced into cells, it is not surprising that a number of the pathway intermediates with undesirable negative effects on cell growth and target production may accumulate. Even when the pathway is entirely native, metabolic engineering involves increasing flux by several orders of magnitude, meaning that imbalances in enzyme activity levels may lead to larger fluctuations in metabolite concentrations than would be found naturally. This has been observed in several pathways prominently used in metabolic engineering in the last decade. For example, the mevalonate-based isoprenoid pathway derived from Saccharomyces cerevisiae has been used for the production of anti-malarial drug artemisinin, as well as a number of biofuel molecules. Two intermediates in this pathway are known to cause cellular growth inhibition. One is 3-hydroxy-3-methylglutaryl-coenzyme A (HMG-CoA), which leads to a feedback regulation in the fatty acid biosynthesis pathway due to its similarity to malonyl-CoA.74 The second is downstream of the main monomer generated from this pathway, isopentenyl pyrophosphate. Isopentenyl pyrophosphate itself is toxic to the cell, but the subsequent prenyl diphosphates geranyl pyrophosphate (GPP) and farnesyl pyrophosphate (FPP), are progressively more toxic. The toxicity of these compounds was evaluated using strains that accumulate isopentenyl pyrophosphate, GPP, and FPP, and the accumulation of FPP was found to be highly growth inhibitory in E. coli.75 Accumulation of prenyl pyrophosphates is also inhibitory in other metabolic engineering strain platforms,76–78 and has required optimization.
Alleviation of intermediate toxicity can be achieved by rebalancing the expression of the pathway enzymes, in particular by increasing the expression of enzymes downstream of the intermediate.78 However, careful static manipulation of the expression of every pathway gene is difficult, and a more systems biology approach is to mimic the circuits built by nature to address this challenge. Circuits in which a metabolite regulates its own production or consumption can afford very tight control over the intracellular concentration. Such an approach was taken in Dahl et. al.,79 where a native E. coli promoter that was negatively regulated in response to FPP stress was used to regulate the upstream atoB gene, while a positively regulated promoter was used to drive the ads gene downstream.
The potential applications of dynamic regulation go beyond the relief of intermediate toxicity. Ideally, the activity of every enzyme would be regulated by the balance of its substrates and products, and overall pathway activity would be regulated by signals corresponding to sufficient carbon and energy availability, ensuring that production does not interfere with cell viability. The potential of this approach for improving production was demonstrated more than a decade ago80 by using promoters regulated by the cellular metabolic state to control the rate-limiting steps of the pathway. An analogous approach was also used to optimize fatty acid ethyl ester production in E. coli. Here the FabR transcriptional factor that interacts with a modified promoter only in its apo form was used to regulate expression of the pathway genes, resulting in optimal gene expression only in the presence of free fatty acids, again resulting in greater and more stable production levels.81 Since then a variety of regulatory motifs have been developed82 with one of the most advanced being a tiered control that could integrate five separate cues into one final output.83 Such regulatory motifs will be required as we develop strains that can withstand a range of perturbations and dynamically optimize production in the growth modes needed for industrial production.
Almost all instances of dynamic regulation developed to date rely on transcriptional regulatory circuits, since transcriptional control is highly modular and well characterized. However, transcriptional control circuits have serious limitations, mostly due to two closely related factors: first, metabolic fluctuations occur on time scales much faster than the transcriptional response, and second, transcriptional activation is largely irreversible, as accumulated protein must either slowly be diluted over a long generation time, or be degraded and resynthesized at a great metabolic cost. A significant challenge will be to engineer circuits based on allosteric regulation (Figure 2), or other fast and reversible protein–metabolite interactions. These circuits would mimic natural control and allow enhanced regulation of metabolic processes. We envision future cells that can be programmed to respond quickly to internal and external signals, and demonstrate phenotypes that are as robust as the metabolism of naturally evolved organisms.
Dynamic regulation for metabolic pathways displays significant advantages with respect to static regulation. The largest challenge in this area involves using allosteric mechanisms for this regulation. We envision future cells that can be programmed to respond quickly to internal and external signals, and demonstrate phenotypes that are as robust as the metabolism of naturally evolved organisms. (a) Shows the basic scheme of dynamic regulation as opposed to static regulation. (b) Explains two possible modes of dynamic regulation.
Engineering of host metabolism
Finding optimal flux distributions
In parallel to pathway optimization, manipulating host metabolism to direct as much flux as possible into the desired pathway is a critical aspect of metabolic engineering. An engineered pathway must operate in the context of the rest of cellular metabolism, which acts as both a blessing and a curse. On one hand, metabolic engineers can rely on innate cellular pathways for sugar catabolism and generation of not only the building blocks that form the starting point for the desired pathway, but also energy and redox cofactors such as ATP, NADH, or NADPH. Cells have machinery to sense an increased demand for these cofactors and adjust their metabolism accordingly,84 but this metabolic control is optimized for growth and survival, making host engineering necessary to optimize the cell for production. On the other hand, an overwhelming demand for carbon and energy from the engineered pathway may impose an insurmountable burden for the cell, which has to carry out a variety of other processes,85 necessitating significant systems-level approaches in host engineering that rely on large-scale models of cell metabolism.
Stoichiometric genome-scale metabolic models are now frequently used for considering the entire metabolic network and understanding how alterations in central pathways propagate to the rest of cellular metabolism.86 The first and simplest use of stoichiometric models (genome-scale or not) is finding the maximal theoretical yield of product and the distribution of fluxes in central metabolism that leads to the optimal yield.87 Algorithms such as OptStrain88 can search reaction databases and find additional reactions that could be added to the network to improve theoretical yield. Second, stoichiometric models can be used to predict the effects of gene deletions on the flux distribution of the considered strain. These algorithms rely on some heuristic assumptions, typically that the distribution will be as close as possible to the wild-type (e.g., MOMA,89 ROOM,90 RELATCH91) or that some aspect of the wild-type distribution, such as a specific flux ratio, will be conserved (PFF92). Despite the necessary heuristic assumptions, these methods have been quite successful and have found applications in metabolic engineering, such as for dihydroartemisinic acid production.93 When product yield optimization is the main goal, a third use involves identifying the fluxes that are required to change in order to achieve the desired product yield increase (e.g., OptForce94–96 and FSEOF97). The corresponding genes are attractive targets for overexpression or knockdown, even though there is no guarantee that manipulating their expression will lead to a corresponding change in flux. For the last two use cases, a critical input is the reference (wild-type) flux distribution, which is often obtained in silico by assuming maximal biomass yield (Flux Balance Analysis). However, since results may significantly depend on this reference flux profile and it is not clear that the growth maximization principle is universally applicable (particularly for bioengineered reference strains), it is desirable to obtain this reference flux distribution more accurately through 13C tracing experiments.98–100 The combination of these labeling experiments with genome-scale models101,102 provides a seamless route to use this information to improve predictions. A final use for stoichiometric models is designing “growth-coupling” strategies wherein maximization of biomass yield actually forces flux through the desired production pathway,103–105 as explained in detail below.
Next-generation genome-scale models expand current metabolic models by taking into account the metabolic investment in expressing proteins,106 or by coupling simulations for all the constituent cellular processes.107 It remains to be seen whether, for metabolic engineering purposes, the extra predictive power compensates for the added complexity in these modeling frameworks.
Balancing of required cofactors
Highly reduced products such as biofuels often rely on reductive anabolic pathways, which typically require NADPH as an electron donor. As most typical microbes generate NADH during glycolysis, the stoichiometric analysis described above often identifies the imbalance between these cofactors as a major problem. Alternative catabolic pathways may be available—a popular method includes deletion of the phosphoglucoisomerase (pgi) gene, forcing glucose catabolism through the NADPH-generating pentose phosphate pathway,108 while genome-scale stoichiometric models can identify other less intuitive genetic changes likely to increase NADPH production.109,110 A recent successful example focused on overexpression of Entner-Douderoff pathway enzymes from Zymomonas mobilis in E. coli.111
An alternative approach is to modify the specificity of an enzyme to accept the alternative redox cofactor, which can often be done by a rational, structure-guided approach. Javidpour et al.112 modified one step of the fatty acid biosynthesis pathway to accept NADH instead of NADPH, and found a concomitant increase in the production of either free fatty acids or long chain methyl ketones. Cofactor specificity could potentially also be modified in central metabolism, to produce NADPH instead of NADH, however, successful examples of this are relatively sparse. One key consideration is the thermodynamic constraints: since most organisms keep NADPH/NADP+ ratios significantly higher than NADH/NAD+ ratios, more energy is required to regenerate NADPH than NADH. Thus, algorithms based on stoichiometry to find optimal points in metabolism for altering cofactor specificity113 need to consider whether sufficient thermodynamic driving force can be maintained.114
Growth coupling
Although identifying the optimal flux distribution may guide some genetic changes that bring the metabolic network closer to the optimal state, there is generally no guarantee. Host cells are adept at finding new ways to reroute metabolism and avoid the fitness cost that underlies shifting resources away from growth to production of the targeted molecule. A powerful approach, which can also be guided by stoichiometric analysis, is to engineer a cell in which production is growth coupled, i.e., cellular growth by necessity produces the desired compound as a by-product. This is the situation in anaerobic fermentation, where excess reducing equivalents from sugar catabolism are forced to be secreted as products like ethanol and butanol. The requirement is that the pathway between sugar and product be redox balanced and produce other by-products necessary for cell growth, typically ATP from substrate-level phosphorylation.
This basic strategy was implemented in a number of studies. The first study in E. coli eliminated ethanol and lactate secretion pathways and imposed anaerobic growth conditions, leaving succinate production as the main electron sink.115 After several generations of evolution, the authors were able to obtain higher succinate titers than any previously reported. In a study by Shen et al.,116 the succinate sink was also eliminated and a heterologous pathway for n-butanol production was inserted as the sole electron sink, and a similar strategy was employed for n-hexanol117 and lactate production.118 When the pathway is not perfectly redox balanced, a more oxidized substrate can be used, such as gluconate instead of glucose.119 The net ATP gain is critical in these schemes, making the recent development of fatty acid biosynthesis by reverse β-oxidation120 very exciting. Fatty acid biosynthesis combined with chain termination enzymes can be used to produce a large variety of long-chain molecules with great potential for use as fuels and chemicals,121 but requires all of the ATP produced from glycolysis for each extension. However, in reverse β-oxidation, no ATP is required after the first extension step, making growth coupling possible. A careful investigation of the potential of this method is provided by Cintolesi et al.122
Many algorithms to identify other potential growth coupling schemes based on genome-scale stoichiometric models of metabolism have been developed,103,105,123,124 but success stories of growth coupling that are not redox based are sparse. Many examples exist of genetic screens for enzymes that complement auxotrophy and these have been used to select for functional enzymes in amino acid biosynthesis125 or novel pathways to isoprenoids.126 However, in these cases the selection pressure disappears after a minimum production level, and a stronger effect can be obtained by screening for resistance to an inhibitor.127 Although these assays allow for growth-based screening or short-term evolution, they do not retain the other major benefit of growth coupling, i.e., the maintenance of strain performance in a new environment, notably, the large-scale bioreactor.
Aside from the difficulty in generalizing the growth coupling strategy, the major drawback is that a substantial amount of carbon is diverted to biomass. The alternative is a process in which the growth and production stages are separated, typically by limiting a necessary nutrient.128,129 However, overall metabolic rates may be drastically lowered130 as there is little selection pressure for microbes to maintain active metabolism in the absence of growth. Understanding this regulation and engineering the decoupling of growth and metabolism is a major challenge in the field.
Scaling up
Physiology and metabolism
A grand challenge in metabolic engineering is scaling production from typical lab conditions (1–100 ml volumes and relatively low substrate and biomass concentrations) to commercial reactors (100–106 l volumes, high cell density and substrate loading) without losing performance.18 Average process development typically takes 5–10 years and is significantly more expensive than scaling up an equivalent chemical process.131 Typically, initial attempts lead to reduced yields, undesired side products, and ultimately a diminished batch-to-batch consistency and product quality, with important economic consequences.132
Many possible factors can account for this drop in performance. Although initial strain testing is done in batch cultures, the fed-batch process used at larger scale leads to different physiology and metabolic states, which may no longer provide the necessary flux distribution in central metabolism. A longer growth period may select for strains that avoid the fitness cost of diverting large amounts of carbon and energy to the product. The strategies outlined above, to either couple production to growth, or separate the growth and production stages, are critical to success. For strain improvement beyond the initial pathway characterization, it is critical to consider process parameters, and design small-scale testing procedures that mimic the large-scale process as much as possible.131 In this regard, a widely used approach is to keep certain dimensionless co-efficients (such as dimensionless mixing time, or power input number132) constant. Probably the most applied scale-up variable is the volumetric oxygen transfer coefficient (kLa), characterizing the oxygen availability.
The volume effect
One aspect of the scale-up process that is challenging to mimic at laboratory scale is the direct effect of reactor volume and associated inhomogeneity in the bioreactor. In typical bioreactor setups, inefficient mixing produces gradients of glucose and oxygen, which can create deviations as large as 400-fold and reduce biomass yield by as much as 20%.133 When similar inhomogeneities were reproduced in a smaller bioreactor for a S. cerevisiae fermentation, the aerobic/anaerobic shifts increased by-product formation, reduced biomass growth, and increased oxygen demand.134,135 Similarly, large-scale E. coli studies revealed increased acetate formation and reduced biomass growth as compared with lab-scale fermentations.131,133 More interestingly, Ying Lin et al.136 showed reduced heterologous protein formation for E. coli in an oscillating glucose supply, which also resulted in increased carbon dioxide production. Furthermore, transcriptional studies in E. coli under shifting aerobic/anaerobic conditions showed a 1.5- to 6-fold increase in transcript levels for mixed acid fermentation genes and several global regulators.137
Understanding and predicting the volume-related performance changes in large-scale fermentation usually consists of combining the information from down-scaling experiments with fluid dynamics simulations of the physical and chemical conditions in the bioreactor.138 This combination has been carried out by coupling kinetic models of metabolism with fluid dynamics simulations139,140 (Figure 3). However, while fluid dynamics simulations are rather sophisticated in predicting fluid flow, glucose, and dissolved oxygen profiles,140,141 the metabolic models coupled with them are not as sophisticated (e.g., six differential equations for the kinetic model in Lapin et al.141). This leads to a widespread opinion131,142 that more accurate modeling of microbial metabolism based on data gathered in scale-down experiments is required for effectively predicting performance at, e.g., 100 l from data obtained at a 35 ml shaking flask. This integration remains an open problem in the field that, if solved, would facilitate the commercialization of the products of metabolically engineered strains.
Integrated Fluid Dynamics (IFD) merges traditional fluid dynamics analysis describing fluid flow (blue lines in the bioreactor on top of the figure) and oxygen and substrate profiles, with models of bacterial metabolism. A significant challenge consists in coupling IFD with metabolic models that are as accurate and sophisticated as those describing the physicochemical characteristics of the fermentation.
Toxicity of final products
Another common failure mode of scale-up is the toxicity associated with the higher titer of the final product. Main categories of commodity compounds being targeted in metabolic engineering include fuels, fuel additives, precursors of polymers, plastics, materials, lubricants and adhesives, surfactants and solvents. C2–C12 compounds in these categories often have characteristics of hydrophobic solvents with octanol:water partition co-efficients143 that range from 1–4. Most microorganisms, even solventogentic microbes such as clostridium,144 are extremely sensitive to solvents in this range.145–147 A large number of systems biology studies have been used to explore the underlying causes of toxicity, spanning from transcript and proteome analysis in several different microbes (e.g., E. coli, clostridia, pseudomonads, and cyanobacteria)47,67,148 to MAGE, recombineering, and genome-wide fitness screens. Comprehensively reviewed elsewhere,47,149–153 these studies aim to identify candidates that can then be used to improve the tolerance characteristics.154,155 Screening heterologous gene libraries or fosmid libraries from other microbes with desirable functions is also an effective method to obtain genes that provide such phenotypes and has been used to discover novel efflux pumps from organisms that provided improved tolerance and increased production for monoterpenes in E. coli,156 and to mitigate toxicity from by-products of biomass pretreatments.72,157
Systems biology for synthetic biology: gaps and challenges
The combination of the unprecedented capabilities afforded by synthetic biology with the comprehensive description of biological entities provided by systems biology presents a unique range of opportunities in designing and understanding biology. The recent technical advances in next-generation sequencing, high sensitivity proteomic and metabolomic methods, and developments in fluxomic techniques make systems biology methods more powerful and accessible to the synthetic biology community. The opportunity to use these methods to inform both synthetic biology tool development and metabolic engineering efforts makes significant scientific discoveries possible. The successes described above form a foundation for systems-level development of a wide variety of synthetic biology approaches. Comprehensive genome engineering for tolerance or target molecule production directly benefits from sequencing methods for strain characterization leading to rapid progress toward the goal of developing robust microbial cell factories. However, despite many successes, exciting new techniques or systems that work beautifully in the laboratory often do not directly scale to industrial fermentation conditions, or are not robust to minor changes in system parameters. Efforts to standardize and report experimental conditions have greatly improved the power of transcriptomic data sets, yet the same efforts have not permeated the proteomic, metabolomic, or fluxomic worlds to a comparable degree. Furthermore, the low-sample throughput of these -omics methods severely limits the usefulness of the data beyond the hypotheses of direct interest. Consequently, both systems and synthetic biology would benefit from research efforts that emphasize reproducibility, method and data sharing, and increased throughput of proteomic, metabolomic, and fluxomic analyses.158
Limitations in reproducibility hamper predictability. A process that is not quantitatively reproducible or exhibits a large unknown variability can hardly be used to parameterize predictive models, and cannot be expected to perform as needed under the required conditions. This inability to predict the behavior of biological systems under conditions not experimentally studied lies at the base of our inability to predict the behavior of large-scale fermentations.159 Whereas we can model and predict the physical side of chemical engineering (e.g., fluid flows, dissolved oxygen profiles), our capability to predict biological systems lags behind, hindering our capability to assure proper functioning under non-studied conditions.
Greater reproducibility and predictability is enabled by efficient data and metadata collection and sharing. One cannot fully compare local experiments with similar ones performed in different laboratories without a detailed account of the conditions and reagents. Concurrently, computational biologists, modelers and data scientists worldwide are limited by experimental data with which to validate a growing number of computational approaches designed to predict biological behavior.107,160,161 However, few laboratories perform experimental and computational work simultaneously because of the difficulties of doing so. Hence, the collaboration between experimentalist and computational specialists could become much more fruitful and frequent through a more robust exchange of data. Several initiatives tackle this problem,162–164 but there is more work to be done to enable effective and fast data exchange. These data exchange are particularly important for the further development of synthetic biology and biological computer aided design:158 as has been shown by Davidsohn et al.,165 the rigorous characterization of pathway parts enables the accurate prediction of the behavior of full pathways.
Conclusion
The uses of synthetic biology showcase the extraordinary opportunities in what has been called the century of biology,166 from understanding fundamental scientific questions to unique practical applications. Once biological systems are pushed to their limits (in terms of, e.g., product yield improvement), systems biology becomes fundamental to understand and predict them. However, bioengineering takes significantly more time and effort and is much less precise than other types of engineering in more established fields (civil, mechanical, electrical, etc). Changing the status quo will require significant investments in the basic tools that will improve productivity and precision in biological design, just as, e.g., integrated circuits have for the computer sciences. In the information technologies field, implementation has been optimized to the point that a limited amount of money and a small team is sufficient to create enough value to launch a company. The situation is the opposite in biological engineering, where implementation of ideas takes much more effort than creating them. In this review, we have presented what we believe are some challenges that need to be overcome in order to reach that level of maturity.
References
Wöhler, F. Ueber künstliche Bildung des Harnstoffs. Ann. der Phys. 88, 253–256 (1828).
Edelman, G. M., Gally, J. A. & Baars, B. J. Biology of consciousness. Front. Psychol. 2, 4 (2011).
Lienert, F., Lohmueller, J. J., Garg, A. & Silver, P. A. Synthetic biology in mammalian cells: next generation research tools and therapeutics. Nat. Rev. Mol. Cell Biol. 15, 95–107 (2014).
Ruder, W. C., Lu, T. & Collins, J. J. Synthetic biology moving into the clinic. Science 333, 1248–1252 (2011).
Slomovic, S., Pardee, K. & Collins, J. J. Synthetic biology devices for in vitro and in vivo diagnostics. Proc. Natl Acad. Sci. USA 112, 14429–14435 (2015).
Hillson, N. J., Hu, P., Andersen, G. L. & Shapiro, L. Caulobacter crescentus as a whole-cell uranium biosensor. Appl. Environ. Microbiol. 73, 7615–7621 (2007).
Chen, J., Sun, S., Li, C. -Z., Zhu, Y. -G. & Rosen, B. P. Biosensor for organoarsenical herbicides and growth promoters. Environ. Sci. Technol. 48, 1141–1147 (2014).
Lee, J. W. et al. Systems metabolic engineering of microorganisms for natural and non-natural chemicals. Nat. Chem. Biol. 8, 536–546 (2012).
Beller, H. R., Lee, T. S. & Katz, L. Natural products as biofuels and bio-based chemicals: fatty acids and isoprenoids. Nat. Prod. Rep. 32, 1508–1526 (2015).
Fortman, J. L. et al. Biofuel alternatives to ethanol: pumping the microbial well. Trends Biotechnol. 26, 375–381 (2008).
Lorek, S. & Spangenberg, J. H. Sustainable consumption within a sustainable economy—beyond green growth and green economies. J. Clean. Prod. 63, 33–44 (2014).
Werpy, T. & Petersen, G. Top Value Added Chemicals from Biomass: Volume I — Results of Screening for Potential Candidates from Sugars and Synthesis Gas. Report No. DOE/GO-102004-1992 (National Renewable Energy Laboratory, 2004).
Nakamura, C. E. & Whited, G. M. Metabolic engineering for the microbial production of 1,3-propanediol. Curr. Opin. Biotechnol. 14, 454–459 (2003).
Yim, H. et al. Metabolic engineering of Escherichia coli for direct production of 1,4-butanediol. Nat. Chem. Biol. 7, 445–452 (2011).
Paddon, C. J. et al. High-level semi-synthetic production of the potent antimalarial artemisinin. Nature 496, 528–532 (2013).
Van Dien, S. From the first drop to the first truckload: commercialization of microbial processes for renewable chemicals. Curr. Opin. Biotechnol. 24, 1–8 (2013).
Kitano, H. Systems biology: a brief overview. Science 295, 1662–1664 (2002).
National Research Council. Industrialization of Biology: A Roadmap to Accelerate the Advanced Manufacturing of Chemicals (The National Academies Press, Washington, DC, USA, 2015).
American Academy of Arts & Sciences. Arise 2: Unleashing America’s Research and Innovation Enterprise (American Academy of Arts & Sciences, 2013).
Baeshen, N. A. et al. Cell factories for insulin production. Microb. Cell Fact. 13, 141 (2014).
Becker, J., Lange, A., Fabarius, J. & Wittmann, C. Top value platform chemicals: bio-based production of organic acids. Curr. Opin. Biotechnol. 36, 168–175 (2015).
Dusselier, M., Mascal, M. & Sels, B. F. Top chemical opportunities from carbohydrate biomass: a chemist’s view of the Biorefinery. Top. Curr. Chem. 353, 1–40 (2014).
Jong E. D. e. Bio-based chemicals value added products from biorefineries. (IEA Bioenergy, 2011).
Holladay, J. E., White, J. F., Bozell, J. J. & Johnson, D. Top Value-Added Chemicals from Biomass—Volume II?Results of Screening for Potential Candidates from Biorefinery Lignin. Evaluation II (2007).
Cornwall, W. CLIMATE. Inside the Paris climate deal. Science 350, 1451 (2015).
Leavell, M. D., McPhee, D. J. & Paddon, C. J. Developing fermentative terpenoid production for commercial usage. Curr. Opin. Biotechnol. 37, 114–119 (2015).
Posen, I. D., Griffin, W. M., Matthews, H. S. & Azevedo, I. L. Changing the renewable fuel standard to a renewable material standard: bioethylene case study. Environ. Sci. Technol. 49, 93–102 (2015).
Medema, M. H., Cimermancic, P., Sali, A., Takano, E. & Fischbach, M. A. A systematic computational analysis of biosynthetic gene cluster evolution: lessons for engineering biosynthesis. PLoS Comput. Biol. 10, e1004016 (2014).
Weber, T. et al. Metabolic engineering of antibiotic factories: new tools for antibiotic production in actinomycetes. Trends Biotechnol. 33, 15–26 (2014).
Hatzimanikatis, V. et al. Exploring the diversity of complex metabolic networks. Bioinformatics 21, 1603–1609 (2005).
Campodonico, M. A., Andrews, B. A., Asenjo, J. A., Palsson, B. O. & Feist, A. M. Generation of an atlas for commodity chemical production in Escherichia coli and a novel pathway prediction algorithm, GEM-Path. Metab. Eng. 25, 140–158 (2014).
Carbonell, P., Parutto, P., Herisson, J., Pandit, S. B. & Faulon, J. -L. XTMS: pathway design in an eXTended metabolic space. Nucleic Acids Res. 42, W389–W394 (2014).
Carbonell, P., Parutto, P., Baudier, C., Junot, C. & Faulon, J.-L. Retropath: automated pipeline for embedded metabolic circuits. ACS Synth. Biol. 3, 565–577 (2014).
Dietrich, J. A., McKee, A. E. & Keasling, J. D. High-Throughput Metabolic Engineering: Advances in Small-Molecule Screening and Selection. Annu. Rev. Biochem. 79, 563–590 (2010).
Nikel, P. I., Martínez-García, E. & de Lorenzo, V. Biotechnological domestication of pseudomonads using synthetic biology. Nat. Rev. Microbiol. 12, 368–379 (2014).
Berla, B. M. et al. Synthetic biology of cyanobacteria: unique challenges and opportunities. Front. Microbiol. 4, 246 (2013).
Bi, C. et al. Development of a broad-host synthetic biology toolbox for Ralstonia eutropha and its application to engineering hydrocarbon biofuel production. Microb. Cell Fact. 12, 107 (2013).
Casini, A., Storch, M., Baldwin, G. S. & Ellis, T. Bricks and blueprints: methods and standards for DNA assembly. Nat. Rev. Mol. Cell Biol. 16, 568–576 (2015).
Myers, C. J. Microbial Synthetic Biology. Methods in Microbiology 40. Elsevier, (2013).
Kelwick, R., MacDonald, J. T., Webb, A. J. & Freemont, P. Developments in the tools and methodologies of synthetic biology. Front. Bioeng. Biotechnol. 2, 60 (2014).
Casini, A. et al. One-pot DNA construction for synthetic biology: the Modular Overlap-Directed Assembly with Linkers (MODAL) strategy. Nucleic Acids Res. 42, e7 (2014).
Shih, S. C. C. et al. A Versatile Microfluidic Device for Automating Synthetic Biology. ACS Synth. Biol. 4, 1151–1164 (2015).
Wiese, S., Reidegeld, K. A., Meyer, H. E. & Warscheid, B. Protein labeling by iTRAQ: A new tool for quantitative mass spectrometry in proteome research. Proteomics 7, 340–350 (2007).
McAlister, G. C. et al. Increasing the multiplexing capacity of TMTs using reporter ion isotopologues with isobaric masses. Anal. Chem. 84, 7469–7478 (2012).
Redding-Johanson, A. M. et al. Targeted proteomics for metabolic pathway optimization Application to terpene production. Metab. Eng. 13: 1–10 (2011).
Arike, L. et al. Comparison and applications of label-free absolute proteome quantification methods on Escherichia coli. J. Proteomics 75, 5437–5448 (2012).
Rutherford, B. J. et al. Functional genomic study of exogenous n-butanol stress in Escherichia coli. Appl. Environ. Microbiol. 76, 1935–1945 (2010).
Qiao, J. et al. Quantitative iTRAQ LC-MS/MS proteomics reveals metabolic responses to biofuel ethanol in cyanobacterial Synechocystis sp. PCC 6803. J. Proteome Res. 11, 5286–5300 (2012).
Chen, L., Wu, L., Wang, J. & Zhang, W. Butanol tolerance regulated by a two-component response regulator Slr1037 in photosynthetic Synechocystis sp. PCC 6803. Biotechnol. Biofuels 7, 89 (2014).
Wang, H., Alvarez, S. & Hicks, L. M. Comprehensive comparison of iTRAQ and label-free LC-based quantitative proteomics approaches using two Chlamydomonas reinhardtii strains of interest for biofuels engineering. J. Proteome Res. 11, 487–501 (2012).
Liu, J., Chen, L., Wang, J., Qiao, J. & Zhang, W. Proteomic analysis reveals resistance mechanism against biofuel hexane in Synechocystis sp. PCC 6803. Biotechnol. Biofuels 5, 68 (2012).
McKee, A. E. et al. Manipulation of the carbon storage regulator system for metabolite remodeling and biofuel production in Escherichia coli. Microb. Cell Fact. 11, 79 (2012).
Matsuoka, Y. & Shimizu, K. Current status and future perspectives of kinetic modeling for the cell metabolism with incorporation of the metabolic regulation mechanism. Bioresour. Bioprocess. 2, 4 (2015).
Khodayari, A., Zomorrodi, A. R., Liao, J. C. & Maranas, C. D. A kinetic model of Escherichia coli core metabolism satisfying multiple sets of mutant flux data. Metab. Eng. 25, 50–62 (2014).
Garcia, H. G. & Phillips, R. Quantitative dissection of the simple repression input-output function. Proc. Natl Acad. Sci. USA 108, 12173–12178 (2011).
Weaver, L. J. et al. A kinetic-based approach to understanding heterologous mevalonate pathway function in E. coli. Biotechnol. Bioeng. 112, 111–119 (2015).
Chowdhury, A., Khodayari, A. & Maranas, C. D. Improving prediction fidelity of cellular metabolism with kinetic descriptions. Curr. Opin. Biotechnol. 36, 57–64 (2015).
Farasat, I. et al. Efficient search, mapping, and optimization of multi-protein genetic systems in diverse bacteria. Mol. Syst. Biol. 10, 731 (2014).
Contador, C. A., Rizk, M. L., Asenjo, J. A. & Liao, J. C. Ensemble modeling for strain development of L-lysine-producing Escherichia coli. Metab. Eng. 11, 221–233 (2009).
Rollin, J. A. et al. High-yield hydrogen production from biomass by in vitro metabolic engineering: Mixed sugars coutilization and kinetic modeling. Proc. Natl Acad. Sci. USA 112, 4964–4969 (2015).
Ajikumar, P. K. et al. Isoprenoid pathway optimization for Taxol precursor overproduction in Escherichia coli. Science 330, 70–74 (2010).
Zhang, C., Zou, R., Chen, X., Stephanopoulos, G. & Too, H.-P. Experimental design-aided systematic pathway optimization of glucose uptake and deoxyxylulose phosphate pathway for improved amorphadiene production. Appl. Microbiol. Biotechnol. 99, 3825–3837 (2015).
Lee, M. E., Aswani, A., Han, A. S., Tomlin, C. J. & Dueber, J. E. Expression-level optimization of a multi-enzyme pathway in the absence of a high-throughput assay. Nucleic Acids Res. 41, 10668–10678 (2013).
Alonso-Gutierrez, J. et al. Principal component analysis of proteomics (PCAP) as a tool to direct metabolic engineering. Metab. Eng. 28, 123–133 (2014).
George, K. W. et al. Correlation analysis of targeted proteins and metabolites to assess and engineer microbial isopentenol production. Biotechnol. Bioeng. 111, 1648–1658 (2014).
Kell, D. B., Swainston, N., Pir, P. & Oliver, S. G. Membrane transporter engineering in industrial biotechnology and whole cell biocatalysis. Trends Biotechnol. 33, 237–246 (2015).
Mukhopadhyay, A. Tolerance engineering in bacteria for the production of advanced biofuels and chemicals. Trends Microbiol. 23, 498–508 (2015).
Wagner, S. et al. Consequences of membrane protein overexpression in Escherichia coli. Mol. Cell. Proteomics 6, 1527–1550 (2007).
Turner, W. J. & Dunlop, M. J. Trade-Offs in Improving Biofuel Tolerance Using Combinations of Efflux Pumps. ACS Synth. Biol. 4, 1056–1063 (2015).
Mingardon, F. et al. Improving olefin tolerance and production in E. coli using native and evolved AcrB. Biotechnol. Bioeng. 112, 879–888 (2015).
Frederix, M. et al. Development of a native Escherichia coli induction system for ionic liquid tolerance. PLoS ONE 9, e101115 (2014).
Ruegg, T. L. et al. An auto-inducible mechanism for ionic liquid resistance in microbial biofuel production. Nat. Commun. 5, 3490 (2014).
Bar-Even, A., Flamholz, A., Noor, E. & Milo, R. Rethinking glycolysis: on the biochemical logic of metabolic pathways. Nat. Chem. Biol. 8, 509–517 (2012).
Kizer, L., Pitera, D. J., Pfleger, B. F. & Keasling, J. D. Application of Functional Genomics to Pathway Optimization for Increased Isoprenoid Production. Appl. Environ. Microbiol. 74, 3229–3241 (2008).
Martin, V. J. J., Pitera, D. J., Withers, S. T., Newman, J. D. & Keasling, J. D. Engineering a mevalonate pathway in Escherichia coli for production of terpenoids. Nat. Biotechnol. 21, 796–802 (2003).
Heider, S. A. E., Wolf, N., Hofemeier, A., Peters-Wendisch, P. & Wendisch, V. F. Optimization of the IPP precursor supply for the production of lycopene, decaprenoxanthin and astaxanthin by Corynebacterium glutamicum. Front. Bioeng. Biotechnol. 2, 28 (2014).
Sivy, T. L., Fall, R. & Rosenstiel, T. N. Evidence of isoprenoid precursor toxicity in Bacillus subtilis . Biosci. Biotechnol. Biochem. 75, 2376–2383 (2014).
Peralta-Yahya, P. P. et al. Identification and microbial production of a terpene-based advanced biofuel. Nat. Commun 2, 483 (2011).
Dahl, R. H. et al. Engineering dynamic pathway regulation using stress-response promoters. Nat. Biotechnol. 31, 1039–1046 (2013).
Farmer, W. R. & Liao, J. C. Improving lycopene production in Escherichia coli by engineering metabolic control. Nat. Biotechnol. 18, 533–537 (2000).
Zhang, F., Carothers, J. M. & Keasling, J. D. Design of a dynamic sensor-regulator system for production of chemicals and fuels derived from fatty acids. Nat. Biotechnol. 30, 354–359 (2012).
Slusarczyk, A. L., Lin, A. & Weiss, R. Foundations for the design and implementation of synthetic genetic circuits. Nat. Rev. Genet. 13, 406–420 (2012).
Moon, T. S., Lou, C., Tamsir, A., Stanton, B. C. & Voigt, C. A. Genetic programs constructed from layered logic gates in single cells. Nature 491, 249–253 (2012).
Chubukov, V., Gerosa, L., Kochanowski, K. & Sauer, U. Coordination of microbial metabolism. Nat. Rev. Microbiol. 12, 327–340 (2014).
Wu, S. G., He, L., Wang, Q. & Tang, Y. J. An ancient Chinese wisdom for metabolic engineering: Yin-Yang. Microb. Cell Fact. 14, 39 (2015).
King, Z. A., Lloyd, C. J., Feist, A. M. & Palsson, B. O. Next-generation genome-scale models for metabolic engineering. Curr. Opin. Biotechnol. 35, 23–29 (2015).
Shen, C. R. & Liao, J. C. Synergy as design principle for metabolic engineering of 1-propanol production in Escherichia coli. Metab. Eng. 17, 12–22 (2013).
Pharkya, P., Burgard, A. P. & Maranas, C. D. OptStrain: a computational framework for redesign of microbial production systems. Genome Res. 14, 2367–2376 (2004).
Segrè, D., Vitkup, D. & Church, G. M. Analysis of optimality in natural and perturbed metabolic networks. Proc. Natl Acad. Sci. USA 99, 15112–15117 (2002).
Shlomi, T., Berkman, O. & Ruppin, E. Regulatory on/off minimization of metabolic flux changes after genetic perturbations. Proc. Natl Acad. Sci. USA 102, 7695–7700 (2005).
Kim, J. & Reed, J. L. RELATCH: relative optimality in metabolic networks explains robust metabolic and regulatory responses to perturbations. Genome Biol. 13, R78 (2012).
Ip, K., Donoghue, N., Kim, M. K. & Lun, D. S. Constraint-based modeling of heterologous pathways: Application and experimental demonstration for overproduction of fatty acids in Escherichia coli. Biotechnol. Bioeng. 111, 2056–2066 (2014).
Misra, A. et al. Metabolic analyses elucidate non-trivial gene targets for amplifying dihydroartemisinic acid production in yeast. Front. Microbiol 4, 200 (2013).
Ranganathan, S., Suthers, P. F. & Maranas, C. D. OptForce: an optimization procedure for identifying all genetic manipulations leading to targeted overproductions. PLoS Comput. Biol. 6, e1000744 (2010).
Xu, P., Ranganathan, S., Fowler, Z. L., Maranas, C. D. & Koffas, M. a. G. Genome-scale metabolic network modeling results in minimal interventions that cooperatively force carbon flux towards malonyl-CoA. Metab. Eng. 13, 578–587 (2011).
Ranganathan, S. et al. An integrated computational and experimental study for overproducing fatty acids in Escherichia coli. Metab. Eng. 1–18 (2012).
Choi, H. S., Lee, S. Y., Kim, T. Y. & Woo, H. M. In silico identification of gene amplification targets for improvement of lycopene production. Appl. Environ. Microbiol. 76, 3097–3105 (2010).
Wiechert, W. 13C metabolic flux analysis. Metab. Eng. 3, 195–206 (2001).
Stephanopoulos, G. N., Aristidiou, A. A. & Nielsen, J. Metabolic Engineering, Princi ples and Methodologies (1998).
Sauer, U. Metabolic networks in motion: 13C-based flux analysis. Mol. Syst. Biol. 2, 62 (2006).
Gopalakrishnan, S. & Maranas, C. D. 13C metabolic flux analysis at a genome-scale. Metab. Eng. 32, 12–22 (2015).
García Martín, H. et al. A method to constrain genome-scale models with 13C labeling data. PLOS Comput. Biol. 11, e1004363 (2015).
Burgard, A. P., Pharkya, P. & Maranas, C. D. Optknock: a bilevel programming framework for identifying gene knockout strategies for microbial strain optimization. Biotechnol. Bioeng. 84, 647–657 (2003).
Patil, K. R., Rocha, I., Förster, J. & Nielsen, J. Evolutionary programming as a platform for in silico metabolic engineering. BMC Bioinformatics 6, 308 (2005).
Tepper, N. & Shlomi, T. Predicting metabolic engineering knockout strategies for chemical production: accounting for competing pathways. Bioinformatics 26, 536–543 (2010).
O’Brien, E. J., Lerman, J. A., Chang, R. L., Hyduke, D. R. & Palsson, B. Ø . Genome-scale models of metabolism and gene expression extend and refine growth phenotype prediction. Mol. Syst. Biol. 9, 693 (2013).
Karr, J. R. et al. A whole-cell computational model predicts phenotype from genotype. Cell 150, 389–401 (2012).
Bartek, T. et al. Importance of NADPH supply for improved L-valine formation in Corynebacterium glutamicum. Biotechnol. Prog. 26, 361–371 (2010).
Chemler, J. A., Fowler, Z. L., McHugh, K. P. & Koffas, M. A. G. Improving NADPH availability for natural product biosynthesis in Escherichia coli by metabolic engineering. Metab. Eng. 12, 96–104 (2010).
Moreira dos Santos, M., Raghevendran, V., Kötter, P., Olsson, L. & Nielsen, J. Manipulation of malic enzyme in Saccharomyces cerevisiae for increasing NADPH production capacity aerobically in different cellular compartments. Metab. Eng. 6, 352–363 (2004).
Ng, C. Y., Farasat, I., Maranas, C. D. & Salis, H. M. Rational design of a synthetic entner-doudoroff pathway for improved and controllable NADPH regeneration. Metab. Eng. 29, 86–96 (2015).
Javidpour, P. et al. Biochemical and structural studies of NADH-dependent FabG used to increase the bacterial production of fatty acids under anaerobic conditions. Appl. Environ. Microbiol. 80, 497–505 (2014).
King, Z. A. & Feist, A. M. Optimal cofactor swapping can increase the theoretical yield for chemical production in Escherichia coli and Saccharomyces cerevisiae . Metab. Eng. 24, 117–128 (2014).
Noor, E. et al. Pathway thermodynamics highlights kinetic obstacles in central metabolism. PLoS Comput. Biol. 10, e1003483 (2014).
Jantama, K. et al. Combining metabolic engineering and metabolic evolution to develop nonrecombinant strains of Escherichia coli C that produce succinate and malate. Biotechnol. Bioeng. 99, 1140–1153 (2008).
Shen, C. R. et al. Driving forces enable high-titer anaerobic 1-butanol synthesis in Escherichia coli . Appl. Environ. Microbiol. 77, 2905 (2011).
Machado, H. B., Dekishima, Y., Luo, H., Lan, E. I. & Liao, J. C. A selection platform for carbon chain elongation using the CoA-dependent pathway to produce linear higher alcohols. Metab. Eng. 14, 504–511 (2012).
Fong, S. S. et al. In silico design and adaptive evolution of Escherichia coli for production of lactic acid. Biotechnol. Bioeng. 91, 643–648 (2005).
Sekar, K. & Tyo, K. E. J. Regulatory effects on central carbon metabolism from poly-3-hydroxybutryate synthesis. Metab. Eng. 28, 180–189 (2015).
Dellomonaco, C., Clomburg, J. M., Miller, E. N. & Gonzalez, R. Engineered reversal of the β-oxidation cycle for the synthesis of fuels and chemicals. Nature 476, 355–359 (2011).
Lennen, R. M. & Pfleger, B. F. Microbial production of fatty acid-derived fuels and chemicals. Curr. Opin. Biotechnol. 24, 1044–1053 (2013).
Cintolesi, A., Clomburg, J. M. & Gonzalez, R. In silico assessment of the metabolic capabilities of an engineered functional reversal of the β-oxidation cycle for the synthesis of longer-chain (C≥4) products. Metab. Eng. 23, 100–115 (2014).
Feist, A. M. et al. Model-driven evaluation of the production potential for growth-coupled products of Escherichia coli. Metab. Eng. 12, 173–186 (2010).
Tervo, C. J. & Reed, J. L. Expanding Metabolic Engineering Algorithms Using Feasible Space and Shadow Price Constraint Modules. Metab. Eng. Commun. 1, 1–11 (2014).
Yano, T., Oue, S. & Kagamiyama, H. Directed evolution of an aspartate aminotransferase with new substrate specificities. Proc. Natl. Acad. Sci 95, 5511–5515 (1998).
Kirby, J. et al. Enhancing Terpene yield from sugars via novel routes to 1-deoxy-d-xylulose 5-phosphate. Appl. Environ. Microbiol. 81, 130–138 (2015).
Stahmann, K. -P., Revuelta, J. L. & Seulberger, H. Three biotechnical processes using Ashbya gossypii, Candida famata, or Bacillus subtilis compete with chemical riboflavin production. Appl. Microbiol. Biotechnol. 53, 509–516 (2000).
Youngquist, J. T., Rose, J. P. & Pfleger, B. F. Free fatty acid production in Escherichia coli under phosphate-limited conditions. Appl. Microbiol. Biotechnol. 97, 5149–5159 (2013).
Shiio, I., Otsuka, S. I. & Takahashi, M. Effect of biotin on the bacterial formation of glutamic acid. I. Glutamate formation and cellular premeability of amino acids. J. Biochem. 51, 56–62 (1962).
Chubukov, V. & Sauer, U. Environmental dependence of stationary-phase metabolism in Bacillus subtilis and Escherichia coli. Appl. Environ. Microbiol. 80, 2901–2909 (2014).
Neubauer, P. et al. Consistent development of bioprocesses from microliter cultures to the industrial scale. Eng. Life Sci. 13, 224–238 (2013).
Schmidt, F. R. Optimization and scale up of industrial fermentation processes. Appl. Microbiol. Biotechnol. 68, 425–435 (2005).
Bylund, F., Collet, E., Enfors, S.-O. & Larsson, G. Substrate gradient formation in the large-scale bioreactor lowers cell yield and increases by-product formation. Bioprocess Eng. 18, 171 (1998).
Sweere, A. P., Mesters, J. R., Janse, L., Luyben, K. C. & Kossen, N. W. Experimental simulation of oxygen profiles and their influence on baker’s yeast production: I. One-fermentor system. Biotechnol. Bioeng. 31, 567–578 (1988).
Sweere, A. P., Janse, L., Luyben, K. C. & Kossen, N. W. Experimental simulation of oxygen profiles and their influence on baker’s yeast production: II. Two-fermentor system. Biotechnol. Bioeng. 31, 579–586 (1988).
Ying Lin, H. & Neubauer, P. Influence of controlled glucose oscillations on a fed-batch process of recombinant Escherichia coli. J. Biotechnol. 79, 27–37 (2000).
Lara, A. R. et al. Transcriptional and metabolic response of recombinant Escherichia coli to spatial dissolved oxygen tension gradients simulated in a scale-down system. Biotechnol. Bioeng. 93, 372–385 (2006).
Noorman, H. An industrial perspective on bioreactor scale-down: what we can learn from combined large-scale bioprocess and model fluid studies. Biotechnol. J 6, 934–943 (2011).
Xia, J. et al. Advances and practices of bioprocess scale-up. Adv. Biochem. Eng. Biotechnol. 152, 137–151 (2015).
Schmalzriedt, S., Jenne, M., Mauch, K. & Reuss, M. Process Integration in Biochemical Engineering 80. Springer Berlin Heidelberg, (2003).
Lapin, A., Müller, D. & Reuss, M. Dynamic behavior of microbial populations in stirred bioreactors simulated with euler-lagrange methods: traveling along the lifelines of single cells†. Ind. Eng. Chem. Res. 43, 4647–4656 (2004).
Lidén, G. Understanding the bioreactor. Bioprocess Biosyst. Eng. 24, 273–279 (2001).
de Bont, J. Solvent-tolerant bacteria in biocatalysis. Trends Biotechnol. 16, 493–499 (1998).
Zingaro, K. A. & Papoutsakis, E. T. Toward a semisynthetic stress response system to engineer microbial solvent tolerance. MBio 3, e00308–e00312 (2012).
Inoue, A. & Horikoshi, K. Estimation of solvent-tolerance of bacteria by the solvent parameter log P. J. Ferment. Bioeng. 71, 194–196 (1991).
Sikkema, J., de Bont, J. A. & Poolman, B. Mechanisms of membrane toxicity of hydrocarbons. Microbiol. Rev. 59, 201–222 (1995).
Zaldivar, J., Martinez, A. & Ingram, L. O. Effect of alcohol compounds found in hemicellulose hydrolysate on the growth and fermentation of ethanologenic Escherichia coli. Biotechnol. Bioeng. 68, 524–530 (2000).
Brynildsen, M. P. & Liao, J. C. An integrated network approach identifies the isobutanol response network of Escherichia coli. Mol. Syst. Biol. 5, 1–13 (2009).
Woodruff, L. B. A., Boyle, N. R. & Gill, R. T. Engineering improved ethanol production in Escherichia coli with a genome-wide approach. Metab. Eng. 17, 1–11 (2013).
Reyes, L. H., Almario, M. P. & Kao, K. C. Genomic Library Screens for Genes Involved in n-Butanol Tolerance in Escherichia coli. PLoS ONE 6, e17678 (2011).
Brynildsen, M. P. & Liao, J. C. An integrated network approach identifies the isobutanol response network of Escherichia coli. Mol. Syst. Biol. 5, 277 (2009).
Deutschbauer, A. et al. Towards an informative mutant phenotype for every bacterial gene. J. Bacteriol. 196, 3643–3655 (2014).
Gallagher, R. R., Li, Z., Lewis, A. O. & Isaacs, F. J. Rapid editing and evolution of bacterial genomes using libraries of synthetic DNA. Nat. Protoc. 9, 2301–2316 (2014).
Tomas, C. A., Welker, N. E. & Papoutsakis, E. T. Overexpression of groESL in Clostridium acetobutylicum results in increased solvent production and tolerance, prolonged metabolism, and changes in the cell’s transcriptional program. Appl. Environ. Microbiol. 69, 4951–4965 (2003).
Foo, J. L. et al. Improving microbial biogasoline production in Escherichia coli using tolerance engineering. MBio 5, e01932 (2014).
Dunlop, M. J. et al. Engineering microbial biofuel tolerance and export using efflux pumps. Mol. Syst. Biol. 7, 487 (2011).
Sommer, M. O. A., Church, G. M. & Dantas, G. A functional metagenomic approach for expanding the synthetic biology toolbox for biomass conversion. Mol. Syst. Biol. 6, 360 (2010).
Gardner, T. S. Synthetic biology: from hype to impact. Trends Biotechnol. 31, 123–125 (2013).
Takors, R. Scale-up of microbial processes: impacts, tools and open questions. J. Biotechnol. 160, 3–9 (2012).
Hyduke, D. R., Lewis, N. E. & Palsson, B. Ø . Analysis of omics data with genome-scale models of metabolism. Mol. Biosyst. 9: 167–174 (2013).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Rocca-Serra, P. et al. ISA software suite: supporting standards-compliant experimental annotation and enabling curation at the community level. Bioinformatics 26, 2354–2356 (2010).
Haug, K. et al. MetaboLights—an open-access general-purpose repository for metabolomics studies and associated meta-data. Nucleic Acids Res. 41, D781–D786 (2013).
Jones, P. et al. PRIDE: a public repository of protein and peptide identifications for the proteomics community. Nucleic Acids Res. 34, D659–D663 (2006).
Davidsohn, N. et al. Accurate Predictions of Genetic Circuit Behavior from Part Characterization and Modular Composition. ACS Synth. Biol 4, 673–681 (2015).
Venter, C. & Cohen, D. The Century of Biology. New Perspect. Q 21, 73–77 (2004).
Acknowledgements
We thank Anthe George, Blake A Simmons, Steven W Singer, Nathan J Hillson, Argentina Lario Lago and Corinne D Scown for valuable feedback on this manuscript. This work was part of the DOE Joint BioEnergy Institute (http://www.jbei.org) supported by the US Department of Energy, Office of Science, Office of Biological and Environmental Research, through contract DE-AC02-05CH11231 between Lawrence Berkeley National Laboratory and the US Department of Energy. The United States Government retains and the publisher, by accepting the article for publication, acknowledges that the United States Government retains a non-exclusive, paid-up, irrevocable, worldwide license to publish or reproduce the published form of this manuscript, or allow others to do so, for United States Government purposes.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
JK has financial interests in Amyris and Lygos. The funders had no role in the study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Chubukov, V., Mukhopadhyay, A., Petzold, C. et al. Synthetic and systems biology for microbial production of commodity chemicals. npj Syst Biol Appl 2, 16009 (2016). https://doi.org/10.1038/npjsba.2016.9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/npjsba.2016.9
This article is cited by
-
A comprehensive review on methane’s dual role: effects in climate change and potential as a carbon–neutral energy source
Environmental Science and Pollution Research (2023)
-
Bioprospecting of microbial strains for biofuel production: metabolic engineering, applications, and challenges
Biotechnology for Biofuels (2021)
-
Microbial production of advanced biofuels
Nature Reviews Microbiology (2021)
-
Cell-free synthetic biochemistry upgrading of ethanol to 1,3 butanediol
Scientific Reports (2021)
-
Distributed biological computation: from oscillators, logic gates and switches to a multicellular processor and neural computing applications
Neural Computing and Applications (2021)