Abstract
What is cause and what is consequence of aging and whether reactive oxygen species (ROS) contribute to this phenomenon is debated since more than 50 years. Notwithstanding, little is known about the cellular buffer and redox systems in aging Saccharomyces cerevisiae, which is a model for aging stem cells. Using genetically encoded fluorescent sensors, we measured pH, H2O2 levels and the glutathione redox potential compartment-specific in the cytosol of living, replicatively aging yeast cells, growing under fermenting and respiratory conditions until the end of their lifespan. We found that the pH decreases under both conditions at later stages of the replicative lifespan. H2O2 levels increase in fermenting cells in the post-replicative stage, but increase continuously with age in respiring cells. The glutathione redox couple becomes also more oxidizing in respiring cells but surprisingly more reducing under fermenting conditions. In strains deleted for the gene encoding glutathione reductase Glr1, such a reduction of the glutathione redox couple with age is not observed. We demonstrate that in vivo Glr1 is activated at lower pH explaining the reduced glutathione potential. The deletion of glr1 dramatically increases the glutathione redox potential especially under respiratory conditions but does not reduce lifespan. Our data demonstrate that pH and the glutathione redox couple is linked through Glr1 and that yeast cells can cope with a high glutathione redox potential without impact on longevity. Our data further suggest that a breakdown of cellular energy metabolism marks the end of replicative lifespan in yeast.
Similar content being viewed by others
Introduction
Aging is one of the fundamental processes of life, observed not only in multicellular organisms but also in unicellular eukaryotes and even in bacteria.1 Despite many years of research, the cause and consequence of this process is still heavily debated. Saccharomyces cerevisiae has long been used as model organism for aging owing to short generation times and genetic tractability.2–4 Two types of aging are distinguished in yeast, chronological aging and replicative aging.5 Chronological aging is the survival of a (stationary) culture under nutrient-limiting conditions, which is believed to be a model for aging of post-replicative, non-dividing cells in multicellular organisms. Replicative aging is defined as the number of divisions an individual cell can accomplish before it enters a senescent, post-replicative state, modeling the aging of proliferating cells and possibly stem cells.
Different factors may contribute to replicative aging, including the production of reactive oxygen species (ROS), accumulation of aggregated or carbonylated proteins in the mother cell, telomere shortening, loss of heterochromatin, loss of mitochondrial fitness, deregulation in protein levels and subunit stoichiometries, metabolic dysfunctions and pH homeostasis.6–13
The production of ROS has been studied extensively,14 and there is indirect evidence that oxidation products accumulate with age.10,15,16 However, it is not clear how old cells adjust their redox systems to increasing oxidative challenges. All cells use the glutathione disulfide/glutathione (GSSG/2GSH) redox system to maintain a reducing environment in the cytosol, nucleus and the mitochondrial matrix. How the glutathione potential changes in old cells has not been addressed reliably in a compartment-specific manner in living cells.
The pH also seems to change when cells age. It was shown previously that the pH in the vacuole and close to the plasma membrane increases early during replicative aging and that measures aimed to counteract such an increase have an impact on longevity.12
Here, we determined the changes in cytosolic pH, the relative levels of H2O2 and the glutathione redox potential during replicative aging under fermentative and respiratory conditions. There are different approaches to study replicative aging in yeast. Traditionally, manual dissection by micromanipulation is used to follow a small number of individual cells throughout their lifespan. Although, still being the gold standard to determine the lifespan of a strain, number, duration of divisions and cell morphology are the only readout of this assay. In a liquid culture, however, it is much more difficult to selectively observe the fraction of old cells during logarithmic growth, as common lab strains have an average median lifespan of 20–35 divisions and daughter cells readily rejuvenate, outgrowing the old fraction. In recent years, microfluidic chambers that trap mother cells and drain away their daughters opened the field also to fluorescence microscopy.17 To also study proteomics and other physiological parameters, the cells were labeled with biotin and subsequently enriched using paramagnetic beads.13,18,19 By elaborated genetic engineering, a system called the ‘mother enrichment program’ was generated where the division of daughter cells was selectively suppressed, enhancing the yield of the fraction of old cells.20 However, sorting by paramagnetic beads cannot distinguish between living and dead cells and enrichment may cause stress to the cells and impact the observed parameters. In addition, there may also be contaminating daughter cells, which have to be excluded by other observations.
To accurately measure the conditions in the cytosol, we used GFP-based sensor proteins for pH (ratiometric pHluorin),21 H2O2 (roGFP2-Orp1)22 and glutathione redox potential (roGFP2-Grx1).23 We found that the pH decreases late in life and that this is most likely a consequence of decreasing ATP concentrations in the cell. H2O2 levels increase with age in both fermenting and respiring cells, albeit at different stages of aging. Although the GSSG/2GSH redox system becomes more oxidizing rather early in aging cells grown under respiratory conditions, the GSSG/2GSH redox couple, unexpectedly, becomes more reducing in fermenting cells. Furthermore, we demonstrate that the oxidation level of the GSSG/2GSH redox couple has no impact on replicative longevity in liquid culture.
Results
To follow a large number of cells during the aging process and to avoid potential artifacts of current enrichment protocols for aged cells, we established a flow cytometry-based approach to selectively measure young and old cells in parallel from logarithmic growing cultures directly within the growth medium. Briefly, we used a fluorescent dye cross-linked to the yeast cell wall to distinguish the population of old mother cells from their unlabeled offspring. By using flow cytometry and sorting, we could both measure and retrieve old and young cells and follow them over their replicative lifespan.
Commonly used laboratory strains, e.g., S288c and derivatives, including the mother enrichment and the BY strains exhibit population heterogeneity due to sporadic loss of mitochondrial DNA (petite or ρ0 phenotype), which leads to respiration deficiency.24–26 For this reason, very few aging studies have been performed under respiratory conditions. For our study, we chose a modified W303-1A strain, which already shows a lower petite frequency, and replaced the mutated mitochondrial DNA polymerase encoding allele (mip1-661A) by the wild-type allele of Σ1278b and introduced the ADE2 wild-type gene, further referred to as WΣ (Supplementary Figure S1).
The cells were constantly grown in synthetic medium and enriched by flow cytometry before depletion of nutrients. At each time point, aliquots were taken for measurements and to determine the age of the population by budscar counting (Figure 1a, Supplementary Figures S1 and S2). The budscar distribution at successive sorts and the population-average of budscars for one experiment is shown in Figures 1b and c. Of note, the budscar distribution becomes very wide at late stages of the aging culture (after 60 h) indicating that some cells stop dividing early but remain viable for a rather long time, whereas other cells continue to divide until they reach the end of their lifespan. As we only evaluated viable cells as judged by GFP fluorescence, which is lost as soon as plasma membrane integrity is compromised, and propidium iodide staining, which is excluded by viable cells, the average number of budscars sometime even decreased by the last day of the experiments, because cells with the highest number of budscars already died, whereas cells that arrested growth earlier are still alive.
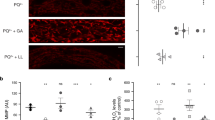
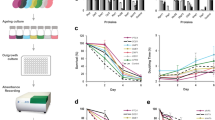
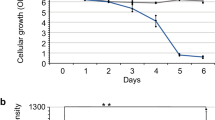
The cytosolic pH decreases in aging yeast cells. (a) Images of logarithmic WΣ cells (left), which are labeled and then initially sorted to result mostly cells without budscars; WΣ cells after the 24 h of growth; and cells at a final stage of aging (see also Supplementary Figures S1 and S2). Budscars colored with solophenylflavin. Scale bars, 2 μm. (b) Budscar distribution of aging WΣ yeast cells at successive FACS (fluorescence-activated cell sorting) enrichment steps at the indicated time points. For every time point, the budscars of 56 to 395 (average 130) single cells were counted. (c) Representative aging curve of WΣ grown under fermenting conditions. Budscars were counted and averaged for every time point. (d) Cellular distribution of pHluorin, roGFP2-Orp1 and roGFP2-Grx1. Wide-field fluorescence image in maximal projection. Scale bar, 2 μm. Z-stacks of confocal images are provided as Supplementary Figures S3, S4, S6–S9. (e) Average cytosolic pH of WΣ cells logarithmically growing on fermentable (2% glucose, SC) and non-fermentable (3% glycerol, 2% ethanol, SG) as indicated. (wt SC, n=27; wt SG, n=31; Error bars represent the standard deviation). (f and g) Development of the population-averaged pH of aging cells grown under fermentative (f, SC-medium with 2% glucose) or respiratory (g, SG-medium with 3% glycerol and 2% ethanol) conditions. The young cells (blue) are the rejuvenated offspring of the old population (red) within the same culture and used as internal control. Standard error of the mean is smaller than the symbols. Numbers below the red symbols indicate the average number of budscars at each time point. Asterisks above the symbols indicate the probability for the statistical significance of the difference between old cells and their rejuvenated offspring within the same culture using Dunn’s multiple comparison test (****P<0.0001). (h) Difference of the population-averaged pH of aging cells and their rejuvenated offspring grown under fermentative (green, SC: 2% glucose) or respiratory conditions (black, SG: 3% glycerol and 2% ethanol) plotted versus the average number of budscars of the old cells. (i) Comparison of budscars of aging fermenting (green) and respiring (black) cells. To clarify the trend, the data were fitted by a Gompertz function (plateau of the fit 23.7±0.08 (SC) and 27.1±0.12 (SG)). The data of several independent aging experiments are shown. The error bars indicate the standard error of the mean (n=56–395). The dotted lines indicate the maximum of the Gompertz function. Inset, average of the maximal budscar average measured in different experiments of WΣ cells grown in SC (green, n=8) or SG (black, n=2) medium. The error bars indicate standard deviation (P=0.0008; Student’s t-test). (j) Replicative competence of aged fermenting (green) and respiring (black) cells at different replicative age (colony-forming units were calculated from the sort of 384 single cells from each time point on YPD-agar plates). The data of 10 (fermentation) and two (respiration) independent aging experiments are shown. (k) Replicative lifespan analysis using the dissecting assay. Two independent assays are shown for cells growing on 2% glucose (Glc)- or 3% glycerol/2% ethanol-containing plates (Gly/EtOH). The difference is highly significant (log-rank Mantel–Cox test P<0.0001; Gehan–Breslow–Wilcoxon test P<0.0001).
Cytosolic pH decreases in old yeast cells
The cytosolic pH of logarithmically growing young and old cells was determined under fermentative and respiratory conditions using genetically encoded ratiometric pHluorin, which was expressed in the cytosol (Figure 1d, Supplementary Figures S3 and S4). The average cytosolic pH under fermentative and respiratory conditions was 7.65±0.08 and 7.36±0.08 (Figure 1e; see Supplementary Discussion). A consistent acidification of the cytosol was observed during the aging process, whereas the rejuvenated offspring all retained a high pH (Figures 1f and g; Supplementary Figure S5). When plotting the difference of pH between the old and young cells versus the number of budscars (Figure 1h), it becomes apparent that most changes occur when the average number of budscars does not increase anymore, indicating that the change in pH most likely happens in post-replicative cells. Interestingly, the respiring cells have an about 0.3 units more acidic pH during logarithmic growth than the fermenting cells, but pH also decreased with age.
While measuring the pH-reporter, we noticed that cells growing under respiratory conditions became older and were more vital over their entire lifespan (Figures 1i and j). This observation was validated using a standard dissecting aging assay (Figure 1k). This result is surprising, assuming that respiration comes with an elevated production of ROS.
Growth medium impacts on the GSSG/2GSH redox couple during aging
To see whether cytosolic peroxide levels differ between the growth conditions, we used the H2O2-sensitive redox-sensor roGFP2-Orp1 (Figure 1d, Supplementary Figures S6 and S7). The fermenting cells started to show an increased probe oxidation only at later stages of their lifetime (Figure 2a), whereas probe oxidation increased linearly in respiring cells up to 20 divisions on average and then stayed constant until the end of the replicative lifespan (Figure 2b). The differences observed were highly statistically significant (P<0.0001).
Change of H2O2 levels and glutathione-reducing potential in aging yeast cells. (a and b) Development of the population-averaged H2O2 production (measured as % probe oxidation of the roGFP2-Orp1 reporter) in aging fermenting (a) and respiring (b) cells. Red, old cells; blue, rejuvenated offspring in the same culture. Standard error of the mean is smaller than the symbols. Asterisks below or above the symbols indicate the probability for the statistical significance of the difference between the old cells and their rejuvenated offspring within the same culture using Dunn’s multiple comparison test (****P<0.0001). The inset shows the difference Δ %OxD between the old and young cells from two independent experiments plotted against the age in average number of budscars within the population. (c and d) Redox potential of the GSSG/2GSH redox couple in terms of roGFP2-Grx1 probe oxidation in aging fermenting (c) and respiring (d) cells. Red, old cells; blue, rejuvenated offspring in the same culture. Standard error of the mean is smaller than the symbols. Asterisks below the symbols indicate the probability for the statistical significance of the difference between the old cells and their rejuvenated offspring within the same culture using Dunn’s multiple comparison test (****P<0.0001). The inset shows the difference Δ %OxD between the old and young cells from two independent experiments plotted against the age in average number of budscars within the population.
As the cytosolic glutathione system provides a reducing milieu, we anticipated that the increased H2O2 levels would also affect the GSSG/2GSH redox couple during the aging process. We therefore measured the cytosolic glutathione redox potential using the roGFP2-Grx1 probe (Figure 1d, Supplementary Figures S8 and S9). The respiring cells indeed became more oxidizing during the aging process paralleling the H2O2 increase (Figure 2d) but, unexpectedly, the probe indicated more-reducing conditions in aged fermenting cells (Figure 2c) that prevailed even in cells at the end of their lifespan, which exhibited increased H2O2 levels. These differences were highly statistically significant (P<0.0001).
WΣ as a derivative of W303 contains the hypomorphic ybp1-1 allele.27 Ybp1 (Yap1 binding protein 1) is one of the components that are important for H2O2 activation of the transcription factor Yap1, the main regulator of the peroxide stress response.28 Although in strains with non-functional Ybp1, Yap1 is activated in a Ybp1-independent way,29 we wondered whether the ybp1-1 allele is responsible for the reducing phenotype despite increased H2O2 levels in aged cells. However, a more-reducing glutathione potential in old cells as compared with the young cells was also observed when the mutant ybp1-1 allele of WΣ was replaced by the wild-type allele (Supplementary Figure S10A) and in a strain with different genetic background (Supplementary Figures S10B and S11).
Glr1 causes the decreased glutathione potential in fermenting cells
There are three ways of shifting the GSSG/2GSH equilibrium to a more-reducing state: removal of oxidized GSSG from the cytosol, reduction of GSSG to GSH and de novo synthesis of GSH. The low affinity transporter Ycf1 sequesters GSSG to the vacuole.30 Therefore, we deleted ycf1 in WΣ and determined the oxidation state of the GSSG/2GSH redox couple during aging. However, the ycf1 deletion had no influence on the age-associated more-reducing glutathione potential (Supplementary Figure S10C).
Regeneration of GSH in S. cerevisiae is mediated by glutathione reductase Glr1, which is the major glutathione reductase in yeast and the most effective protein to maintain the cytosolic redox state.30,31 Although the thioredoxins Trx1 and Trx2 have been shown to be able to reduce GSSG to 2GSH, they only correspond to about 2% of the GSSG-dependent NADPH oxidizing activity in yeast.32 Indeed, the Δglr1 strain in contrast to the Δycf1 strain exhibited a high glutathione redox potential in the cytosol33 (Supplementary Figure S10D). During replicative aging, the decrease in glutathione potential in fermenting Δglr1 cells was diminished (Figure 3a), demonstrating that Glr1 is responsible for the more-reducing glutathione potential in aging cells. This observation also suggests that de novo synthesis of GSH does not contribute to a large extent to the more-reducing glutathione potential in old fermenting cells as such a mechanism should also work in Δglr1 cells. In contrast, under respiratory conditions, the early increase in glutathione potential persisted (Figures 3b and c).
Δglr1 cells have a highly oxidized cytosol but no reduced lifespan under fermentative growth conditions. (a) Glutathione potential in aging WΣ-Δglr1 cells growing under fermentative conditions; red, old cells; blue, rejuvenated young cells in the same culture. Standard error of the mean is smaller than the symbols. Asterisks above the symbols indicate the probability for the statistical significance of the difference between old cells and their rejuvenated offspring within the same culture using Dunn’s multiple comparison test (****P<0.0001). The inset shows the steady-state probe oxidation for wild-type (n=44) and Δglr1 (n=7) cells. Error bars are the standard error of the mean of n independent cultures. (b) Glutathione potential in aging WΣ-Δglr1 cells growing under respiratory conditions. (Error bars and asterisks as in a). (c) Difference Δ %OxD of old and young WΣ-Δglr1 cells growing under fermenting (green) or respiratory (turquois) conditions plotted versus the average number of budscars of the populations. (d) Comparison of the population-averaged number of budscars of wild-type (SC: gray triangles, SG: black inverted triangles) and Δglr (SC: magenta circles; SG: purple squares) cells, enriched from the flow-cytometric aging assay. A Gompertz function was fit to the data to visualize the trend. The data for wild-type cells are the same as in Figure 1i shown for comparison. (e) Replicative competence of aged wild-type and Δglr1 cells grown under fermenting (SC) and respiring (SG) conditions at different replicative age. Colony-forming units were calculated from the sort of 384 single viable cells (as judged by GFP fluorescence) from each time point on YPD-agar plates. The data of 10 (wild type, SC), two (wild type, SG), two (Δglr1, SC) and two (Δglr1, SG) independent aging experiments are shown. The data for wild-type cells are the same as in Figure 1j shown for comparison. (f) Classical lifespan determination of WΣ wild-type (gray and black) and Δglr1 (magenta and purple) by micromanipulation on 2% glucose (Glc)- or 3% glycerol- and 2% ethanol (GlyEtOH)-containing plates. For wt (Glc: median=23; n=71; GlyEtOH: median=30.5; n=92; same data as in Figure 1k shown for comparison). For Δglr1 (Glc: median=22; n=72; GlyEtOH: median=24; n=94). Data from two independent experiments each shown in one survival curve for clarity.
Taken together, it can be concluded that the cytosolic GSSG/2GSH redox system becomes more reduced in a Glr1-dependent manner when cells age under fermentative growth conditions.
Lifespan is not impaired in Δglr1 cells
As Glr1 is also responsible for GSSG reduction in mitochondria and peroxisomes, highly oxidizing conditions prevail throughout Δglr1 cells. Considering these highly oxidizing conditions and referring to the free-radical theory of aging,34 it would be expected that the lifespan of Δglr1 cells is affected. We therefore determined the lifespan by classical micro-dissection and by budscar counting of the enriched fractions from the flow cytometry-based aging assay. We also determined the fraction of cells that were able to form colonies during aging. With either method, we did not detect any remarkable differences in lifespan or replicative potential between wild-type and Δglr1 cells grown under fermentative conditions (Figures 3d–f). Surprisingly, when grown under respiratory conditions, Δglr1 cells did not reach the same median lifespan as wild-type cells in the dissecting assay but in liquid culture they became as old as wild-type cells and retained the ability to form colonies like wild type. Thus, the oxidation level of the GSSG/2GSH redox system does not limit replicative lifespan in yeast—at least in unstressed cells in liquid culture.
In contrast, Δglr1 cells are more sensitive to H2O2 stress. We treated wild-type WΣ, WΣ-YBP1, WΣ Δglr1 and WΣ overexpressing glr1 for 15 min with different concentrations of H2O2 before washing the cells and spotting dilutions onto YPD plates (Supplementary Figure S12). The wild-type WΣ shows no loss in plating efficiency up to 8 mmol/l H2O2, whereas the Δglr1 cells were already compromised between 1 and 2 mmol/l H2O2. The WΣ-YBP1 cells were as resistant as wild-type cells and the glr1-overexpressing cells were about 2-fold more resistant.
Taken together, these data clearly show that although Glr1 is important for dealing with an acute H2O2 shock, it has no direct impact on longevity in yeast.
The GSSG/2GSH redox system is more potent in aged cells
We next asked whether the GSSG/2GSH redox couple of fermenting old cells would respond differently when challenged with severe acute peroxide stress. We measured the response of the glutathione system after treatment with high but sublethal concentrations of H2O2. Young wild-type cells responded to peroxide with an oxidation followed by a fast adaptation phase, reducing the oxidation level of the GSSG/2GSH redox couple to pre-shock levels, though the H2O2 concentration remains constant in our highly dilute cultures throughout the measurements (Supplementary Figure S13). Surprisingly, aged wild-type cells not only started with a more-reducing cytosol but also lacked the initial oxidation peak and equilibrated to a final average oxidation level similar to young cells (Figure 4a; Supplementary Figure S14A).
Aged yeast cells buffer an H2O2 shock more quickly than young cells through a Glr1-dependent pathway. (a) Response kinetics of glutathione potential after addition of 2 mmol/l H2O2 to aging wild-type cells growing under fermentative conditions at the indicated time points. Moving average over 25 cells (raw data are shown in Supplementary Figure S4A). (b) Same kinetics as in a for WΣ-Δglr1 but with 0.5 mmol/l H2O2. Moving average over 25 cells (raw data are shown in Supplementary Figure S4B). (c) Steady-state oxidation of the GSSG/2GSH redox couple in wild-type and Δglr1 strains and upon glr1 overexpression. GSSG/2GSH redox couple measured as % oxidation of the roGFP2-Grx1 probe in fermenting cells. Ectopic expression of glr1 can complement the knockout, but does not lead to a more reduced glutathione potential in wild-type cells (wt, n=1; wt+Glr1, n=4; Δglr1, n=2; Δglr1+Glr1, n=4. Error bars are the s.d.). (d and e) Glutathione potential of fermenting young wild-type (d) and glr1-overexpressing (e) cells in response to 2 mmol/l H2O2. The dashed green line indicates steady-state oxidation. (f and g) roGFP2-Orp1 probe oxidation in fermenting young wild-type (f) and glr1-overexpressing (g) cells in response to 2 mmol/l H2O2.
The fast adaptation of the GSSG/2GSH redox system seems to be accomplished by the action of Glr1, as knockout cells are severely defective in this adaptation (Figure 4b, Supplementary Figure S14B).30
Increased Glr1 levels do not lead to a more-reducing cytosol
As deletion of glr1 leads to a strong glutathione oxidation, we reasoned that the more-reducing cytosol observed in old fermenting cells might be caused by increased Glr1 levels. To test this hypothesis, we attempted to phenocopy the aging phenotype by overexpressing Glr1 in young cells. Surprisingly, overexpression did not lead to a more-reducing glutathione buffer, though the plasmid could completely rescue the knockout phenotype (Figure 4c).
Unexpectedly, when glr1-overexpressing cells were challenged with peroxide, the initial oxidation peak was barely detectable, but instead the reduction phase was much more prominent and even led to hyper-reduction of the glutathione reporter (Figures 4d and e) despite the fact that the cells showed similar kinetics in the peroxide reporter as compared with the wild-type cells (Figures 4f and g).
Glr1 activity is pH dependent
From previous measurements, we noticed that addition of 2 mmol/l H2O2 impacts the cytosolic pH of fermenting but much less respiring yeast cells. This difference in sensitivity of respiring cells as compared with fermenting cells is observed over a wide range of H2O2 concentrations (Supplementary Figures S12B and S15). Interestingly, the pH changed with similar initial kinetics as the adaptation of the GSSG/2GSH redox system. We therefore hypothesized that the activity of Glr1 might be regulated by pH. To test this hypothesis, we artificially altered the cytosolic pH of the cells using the protonophor 2,4-dinitrophenol (DNP) while measuring the glutathione oxidation levels. Because of similar spectral properties, ratiometric pHluorin and roGFPs cannot be measured within the same cell. As a compromise, we co-cultured two yeast strains containing either the pHluorin reporter or the roGFP2-Grx1 probe. By differentially tagging the two strains, we could determine both the pH and the oxidation status of the GSSG/2GSH redox system within the same culture. As the cells were growing in synthetic complete medium (SC) with a pH of 3.9, addition of DNP leads to an acidification of the cytosol.
Concomitant with acidification of the cytosol (Figure 5a) the glutathione potential became strongly reducing (Figure 5b). This cannot be a direct effect of pH on the electrochemical redox potential of the glutathione system. According to the Nernst equation, the potential of the glutathione system as a two-electron/two-proton reaction is affected in the same direction as the redox potential of roGFP, which like all 2-cysteine to cystine transitions is also a two-electron/two-proton reaction. With the roGFP2-Grx1 probe, we measure the ability of the GSSG/2GSH redox couple to regulate the 2-cysteine/cystine equilibrium of cytoplasmic proteins.
The activity of Glr1 is pH dependent in vivo. (a and b) Change of cytosolic pH (a) and GSSG/2GSH redox couple oxidation (b) in response to addition of 25 μmol/l DNP (2,4-dinitrophenol) in wild-type yeast growing in synthetic medium with glucose. Both pHluorin and roGFP2-Grx1 were measured within the same culture but in different cells, one marked with TagBFP. (c and d) Gradually lowering the cytosolic pH of a fermenting (c) and respiring (d) yeast culture by different concentrations of DNP (0–100 μmol/l). The relative oxidation of the cytosolic GSSG/2GSH redox couple of differentially tagged co-cultured wild-type (green), Δglr1 (magenta) and glr1-overexpressing (↑glr1, blue) cells was determined simultaneously 30 min after DNP addition. The gray area is outside the unambiguous range of the pHluorin reporter and pH measurements in this area are imprecise. The open circles in d indicate a pH upshift by the addition of Tris base. Standard error of the mean is smaller then the symbols. (e) Cytosolic pH of wild-type WΣ and Δglr1 strains under fermenting (2% glucose) and respiring (3% glycerol+2% ethanol) conditions (data for wild-type WΣ cells are the same as shown in Figure 1g and shown for comparison); wt glucose, n=27; wt glycerol, n=31, Δglr1 glucose, n=13; Δglr1 glycerol, n=13. Error bars are the s.d. (f) Gradual uncoupling of cells by DNP titration. The cytosolic pH, measured 30 min after DNP addition, for fermenting cells (blue colors) and respiring cells (red colors) is shown. The pH upshift was achieved by increasing amounts of Tris base as indicated by the arrow. Shown are three independent experiments.
We next gradually changed the pH by adding different amounts of DNP to wild-type, Δglr1- and Glr1-overexpressing cells (Figure 5c). Our data clearly show that Glr1 is responsible for the glutathione reduction in a pH-dependent manner, demonstrating that the Glr1 activity is pH regulated and Glr1 is more active at lower pH with a broad optimum between pH 6 and pH 7.
Effect of growth conditions on maintaining the cytosolic pH
We noticed that the oxidation state of the GSSG/2GSH redox couple of respiring cells, which have already a lower pH at steady state (Figures 5d and e), was not influenced much by further acidification but became more oxidizing by pH upshifting using Tris base (Figure 5d). Plotting the pH against the concentration of DNP (positive values) or the amount of added Tris revealed a striking difference between the fermenting and respiring cells (Figure 5f).
Maintenance of the cytosolic pH in yeast cells is mainly mediated by the vacuolar V-ATPase and the essential plasma membrane proton pump Pma1.35,36 These transporters are driven by the hydrolysis of ATP. When glucose is available, yeast cells produce ATP mainly by glycolysis. When growing on non-fermentable carbon sources, oxidative phosphorylation in the mitochondria is the predominant source of ATP through the proton gradient-driven FoF1-complex. DNP uncouples not only the cytosol with the medium but also the mitochondrial matrix. Therefore, respiring cells are much stronger affected by the addition of protonophors than fermenting cells. The ATP production of fermenting cells does not depend on a membrane potential and they are still able to generate enough ATP through glycolysis to power their proton pumps in the presence of DNP.
For mammalian cells, it was shown that addition of H2O2 depletes the ATP levels by inhibiting the glycolytic pathway.37,38 For yeast, it was reported that H2O2 treatment depletes adenine nucleotide levels with a sharp transition between 0.6 and 1 mmol/l H2O2.39 Using the pHluorin reporter, we detected a transition between cells that maintain their cytosolic pH and cells that are unable to maintain their pH at exactly this range (Supplementary Figure S15). Contrarily, the pH of respiring cells is much less affected by H2O2 in this range.
Taken together, our data provide strong evidence that the cellular pH reports on ATP availability.
Discussion
In this study, we established a flow cytometry-based assay for following replicative aging in S. cerevisiae (see Supplementary Discussion) and gained four novel insights into replicative aging. (1) The pH decreases at the end of the replicative lifespan in cells growing under fermentative and respiratory conditions. (2) Despite a general increase in H2O2 levels in aged cells, the glutathione potential becomes more reducing with progressing age in a Glr1 and pH-dependent manner in fermenting but not respiring cells. (3) Glr1 activity is pH dependent in vivo. (4) A high cytosolic glutathione redox potential does not shorten lifespan in yeast grown under fermentative conditions.
Our pH measurements appear to be in conflict with recently published work12 (see Supplementary Discussion). Contradictory to our data, these authors report a slightly increasing pH in aging cells growing under fermentative conditions and propose that this increase is caused by an accumulation of Pma1 in the plasma membrane. Interestingly, Henderson et al.12 observed an increased pH already after three divisions with no significant further increase in the observed time frame and no analysis was given for cells beyond half of the median lifespan of their strain. In contrast, we see the most dramatic changes at a time when cells reach their median lifespan and stationary cells accumulate. Therefore, our work rather describes the situation of cells at the end of their lifespan.
Why is pH decreasing in aging cells? Respiring cells have a lower cytosolic pH than fermenting cells and it could be that at late stages of their life yeast switches to respiration instead of fermentation. However, the decrease in pH observed during respiration is only 0.3 pH units while we observed an age-related decrease of 0.6–0.8. In addition, we also observed an age-related decrease in pH in the respiring cells.
As the cytosolic pH in yeast is maintained by the ATP-dependent plasma membrane and vacuolar proton pumps Pma1 and V-ATPase, there are three possible explanations: either the ATPases become defective or reduced in numbers during aging or ATP supply is limiting. Our DNP titration experiment strongly suggests that cytosolic pH is tightly coupled to ATP levels in the cell (Figure 5f). Thus, impairment of glycolysis, the main source for ATP in fermenting cells, could also be responsible for the decreasing pH. Glucose starvation has been shown to lead to a reduction of cytosolic pH.40 However, in our system glucose never becomes limiting as is demonstrated by the young cells of the same culture, which do not show any decrease in cytosolic pH. Glycolysis is known to be inhibited by peroxides.37,38 However, in fermenting cells, pH started to decrease after around 15 divisions while H2O2 levels only increased at the very end of the lifespan at around 25 divisions, excluding the H2O2 increase that we measured as cause of the decreasing pH. Other age-related processes may affect glycolysis and make it less efficient.
In contrast, inhibition of glycolysis could not be the reason for the decreasing pH in cells grown under respiratory conditions on ethanol and glycerol. Consistently, H2O2 had little effect on cytosolic pH in cells grown on ethanol (Supplementary Figure S15B). There are several possible explanations for the decreasing pH in respiring cells. It was reported that mitochondrial functions break down in aging cells owing to an overload with neutral amino acids caused by an impaired vacuolar storage function.11 Although this was only measured for fermenting cells, it might also apply to respiring cells. In addition, defective and aggregate-associated mitochondria might be retained in mother cells, as has been reported for fermenting cells.7,41 All these changes would also cause a decline of ATP in respiratory growing cells. Our data suggest that a breakdown of cellular energy metabolism marks the end of replicative lifespan in yeast.
What are the consequences of the decreased pH? Depending on the individual pKa values of the amino-acid side chains, protonation can influence the surface charge of a protein and therefore also affects protein folding35 (see also Supplementary Discussion). Another consequence of the decreased pH is the increased activity of Glr1 in fermenting cells as we deduced from our data, consistent with previous biochemical experiments,42 leading to a decreased cytosolic GSSG/2GSH redox state. Of note, Glr1 uses NADPH to reduce GSSG to 2GSH. The GSSG–2GSH transition is a two-electron/two-proton reaction, whereas the NADPH–NADP+ transition is a two-electron/one-proton reaction and thus, according to the Nernst equation, less affected by pH than the GSSG/2GSH redox couple.43 The consequence is that the difference in electrochemical redox potential between the two redox couples increases with decreasing pH, promoting the reduction of GSSG.
The changes of pH, GSSG/2GSH redox state and H2O2 levels in fermenting cells occur between 40 and 50 h of logarithmic growth. At the same time, the budscar distribution becomes wider and the increase of the mean of budscars slows down significantly, indicating that more and more cells cease to divide (Figures 1b, i and j). This suggests that the increasing H2O2 level, the decreasing pH and the Glr1-mediated more-reducing glutathione potential are a consequence not a cause of senescence.
We also found that old fermenting cells seem to cope better with an acute H2O2 challenge than young cells (Figure 4a), suggesting a higher buffering capacity against oxidative stress. These findings are consistent with an earlier report that the transcription factor Yap1, which becomes oxidized by peroxides and subsequently translocates to the nucleus to induce the appropriate stress response, migrates in old cells upon peroxide stress twofold slower to the nucleus than in young cells.44 The more-reducing GSSG/2GSH redox couple and the more-active Glr1 could buffer incoming peroxides and thus slow down Yap1 oxidation.
In contrast, an increase in cytosolic glutathione redox potential is observed in cells aging under respiratory conditions, despite the fact that their pH, which is already lower in young cells as compared with fermenting cells, further decreases with age. We conclude that Glr1 is already at its activity optimum in young respiring cells, which was demonstrated experimentally (Figure 5d; see Supplementary Discussion). Parallel to the increase of glutathione redox potential, the H2O2 level rises. Electron leakage from the mitochondrial respiratory complexes is considered to be a source of ROS,45 which are rapidly interconvert to subspecies like •OH and H2O2.46 This would imply that the number or the metabolic activity of mitochondria correlate with the H2O2 level. As the cells age, they increase in size, suggesting that their total content of mitochondria increases as well. When cells cease to increase in size and presumably enter the stationary phase, their mitochondrial content should not increase further and thus it is expected that H2O2 level also stay constant as observed in our experiments. Whether the endoplasmic reticulum resident NADPH oxidase Yno1, which is stabilized by signaling events from dysfunctional mitochondria and seems to be a major source for ROS in yeast,47 has a role in the H2O2 accumulation observed by us, is unclear. However, it would be expected that dysfunctional mitochondria accumulate at later stages of replicative life and thus stabilization of Yno1 and its ROS production should increase in parallel, which was not observed by us.
Taken together, our data are consistent with the following hypothetical scenario for age-related changes in pH and glutathione-reducing potential. In young fermenting cells, glycolysis sustains a high concentration of cytosolic ATP, allowing Pma1 to maintain the high cytosolic pH. Glr1 reduces GSSG to 2GSH at intermediate activity (Figure 6a). In old/post-replicative cells, the glycolysis rate declines owing to diverse aging factors. This leads to limiting ATP concentrations and reduced Pma1 activity, resulting in a lower cytosolic pH, an increased Glr1 activity and consequently a lower glutathione potential. Respiring cells have already a lower pH and thus Glr1 is at its maximal activity. The glutathione redox potential in young respiring cells is more reducing than in young fermenting cells. As respiring cells age, they increase in size and presumably also their mitochondrial content and therefore the capacity for oxidative phosphorylation. Aging processes may promote mitochondrial dysfunction and electron leakage and consequently decreasing availability of ATP (Figure 6b). This results in a gradual decrease of the pH but not in a more-reducing glutathione potential. Instead, the glutathione system becomes, already at an early stage of aging, more oxidizing, possibly due to continuous ROS production by the mitochondria or endoplasmic reticulum resident NADPH oxidase Yno147 consistent with increasing H2O2 levels measured with the roGFP2-Orp1 sensor.
Model for age-dependent changes in pH, H2O2 levels and glutathione potential. (a) In young fermenting cells, glycolysis runs at an optimal rate and ATP levels are sufficiently high to power the plasma membrane proton pump Pma1, which is able to maintain a high cytosolic pH. Glr1 operates at suboptimal rates keeping the GSSG/2GSH redox couple in a slightly oxidized steady state. In old cells, ATP levels are reduced possibly due to the impairment of glycolysis by elevated H2O2 levels and/or other age-associated factors. The proton pumps (like Pma1) work at limiting rates and cannot maintain the high cytosolic pH anymore. Glr1 is activated by the lower pH and works at its maximal rate, leading to a more-reducing glutathione potential. Top panels indicate idealized curves for the changes in pH, H2O2 (roGFP2-Orp1) and glutathione potential (roGFP2-Grx1). (b) In young respiring cells, ATP is produced by oxidative phosphorylation (OxPhos). The respiring cells already have a lower cytosolic pH in which Glr1 shows its highest activity, leading to a more-reducing steady-state glutathione potential as compared with the fermenting cells. In old respiring cells, mitochondrial energy production may be less efficient, which possibly leads to generation of more ROS including H2O2. This constitutive burden may also affect the steady state of the GSSG/2GSH redox system and leads to a more -oxidizing glutathione potential. Top panels indicate idealized curves for the changes in pH, H2O2 (roGFP2-Orp1) and oxidation state of the GSSG/2GSH redox couple (roGFP2-Grx1).
Despite the fact that H2O2 levels increase and glutathione potential becomes more oxidizing in respiring cells, they have a significantly longer lifespan, divide more often and remain longer proliferation competent (Figures 1i–k). Similarly, Δglr1 cells, which have a much more oxidizing GSSG/2GSH redox potential, do not live shorter than wild-type yeast cells when grown in liquid culture and under fermentative conditions on plates (Figures 3d–f). Under respiratory conditions on plates, Δglr1 cells still get older than under fermentative conditions, despite the fact that probe oxidation is even higher (>70%), but not as old as wild-type cells. These differences may be explained with the sensitivity of Δglr1 cells to acute oxidative stress (Supplementary Figure S5). Growth on plates or the dissecting process may involve more oxidative stress than growth and sorting in liquid culture, resulting in a shorter lifespan of Δglr1 cells on plates. Although the thioredoxin system might still maintain a suitably reduced cytosol,48 cells missing Glr1 are much more sensitive to acute oxidative stress than wild-type cells (Supplementary Figure S12A), and it seems reasonable to assume that in such cells any ROS that emerges from metabolic processes is more likely to escape the redox buffers than in wild-type cells. Whether Δglr1 cells accumulate more oxidative damage, needs to be addressed in future studies. Notwithstanding, our observations clearly demonstrate that the oxidation state of the GSSG/2GSH redox system at least up to 60% OxD in fermenting cells and up to 75% in respiring cells does not correlate with lifespan in yeast.
Materials and methods
Aging assay
A yeast culture was grown logarithmically for at least seven doublings at 25 °C. The cells were then collected by centrifugation and washed twice with phosphate-buffered saline, resuspended in 200 mmol/l NaHCO3 (pH 8.4) and labeled for 15 min with Alexa633-NHS (Life Technologies, Darmstadt, Germany) at a final concentration of 2 μmol/l. Labeling reaction was quenched by washing the cells with Tris/HCl (500 mmol/l, pH 8.4). About 4–8×106 single cells of small size (unbudded) were sorted by FACS (fluorescence-activated cell sorting) with highest purity and inoculated into 100 ml medium. Before reaching an OD600 of 0.4 aliquots were taken for measurements. The remaining cells were collected by centrifugation at 4 °C and quantitatively resuspended in about 3 ml of supernatant. The culture was then shortly sonicated to separate cell aggregates. Enrichment was performed with the Aria III cell sorter (BD, Heidelberg, Germany; filter configuration Ex407 nm/Em502 nm–520 nm, Ex488 nm/Em515 nm–545 nm, Ex633 nm/Em650 nm–670 nm), selecting for Alexa-positive cells with reporter expression (and for propidium iodide negative cells, when this staining was used). The recovered old cells were then inoculated into fresh medium and cultured for the next round. A few cells from the single-cell gate were sorted separately and were either stained with Solophenylflavin (Bruno Ludewig GmbH, Emsbüren, Germany)49 for budscar analysis or sorted on YPD-Agar-plates to determine the replicative competence. The data for the budscars were fitted with a Gompertz function:
Measurements and data processing
Aliquots of the aged/logarithmic culture were shortly sonicated and directly measured with a Canto II (BD; filter configuration: Ex405 nm/Em502 nm–535 nm, Ex488 nm/Em515 nm–545 nm and Ex633 nm/Em650 nm–670 nm; see Supplementary Methods). For all the measurements, a single-cell gate was defined using the FSC-W and SSC-W parameters. Only cells with reporter expression were considered. The data were analyzed either by the Diva (BD) or FlowJo software (FlowJo LLC, Ashland, OR, USA) and then exported to Microsoft Excel or Prism (GraphPad, La Jolla, CA, USA) for further calculations. Autofluorescence was determined in independent experiments and subtracted from all the data.
For kinetic measurements, H2O2 was added and cultures immediately transferred to the analyzer, resulting in a delay of approximately 10 s between H2O2 addition and the first events counted.
References
Nyström, T. A bacterial kind of aging. PLoS Genet. 3, e224 (2007).
Steinkraus, K. A., Kaeberlein, M. & Kennedy, B. K. Replicative aging in yeast: the means to the end. Annu. Rev. Cell Dev. Biol. 24, 29–54 (2008).
Nyström, T. & Liu, B. The mystery of aging and rejuvenation—a budding topic. Curr. Opin. Microbiol. 18, 61–67 (2014).
Kaeberlein, M. Lessons on longevity from budding yeast. Nature 464, 513–519 (2010).
Longo, V. D., Shadel, G. S., Kaeberlein, M. & Kennedy, B. Replicative and chronological aging in Saccharomyces cerevisiae. Cell Metab. 16, 18–31 (2012).
Hu, Z. et al. Nucleosome loss leads to global transcriptional up-regulation and genomic instability during yeast aging. Genes Dev. 28, 396–408 (2014).
Higuchi, R. et al. Actin dynamics affect mitochondrial quality control and aging in budding yeast. Curr. Biol. 23, 2417–2422 (2013).
Erjavec, N., Larsson, L., Grantham, J. & Nyström, T. Accelerated aging and failure to segregate damaged proteins in Sir2 mutants can be suppressed by overproducing the protein aggregation-remodeling factor Hsp104p. Genes Dev. 21, 2410–2421 (2007).
Xie, Z. et al. Early telomerase inactivation accelerates aging independently of telomere length. Cell 160, 928–939 (2015).
Lam, Y. T., Aung-Htut, M. T., Lim, Y. L., Yang, H. & Dawes, I. W. Changes in reactive oxygen species begin early during replicative aging of Saccharomyces cerevisiae cells. Free Radic. Biol. Med. 50, 963–970 (2011).
Hughes, A. L. & Gottschling, D. E. An early age increase in vacuolar pH limits mitochondrial function and lifespan in yeast. Nature 492, 261–265 (2012).
Henderson, K. A., Hughes, A. L. & Gottschling, D. E. Mother-daughter asymmetry of pH underlies aging and rejuvenation in yeast. Elife 3, e03504 (2014).
Janssens, G. E. et al. Protein biogenesis machinery is a driver of replicative aging in yeast. Elife 4, 1–43 (2015).
Afanas'ev, I. Signaling and damaging functions of free radicals in aging-free radical theory, hormesis, and TOR. Aging Dis. 1, 75–88 (2010).
Laun, P. et al. Aged mother cells of Saccharomyces cerevisiae show markers of oxidative stress and apoptosis. Mol. Microbiol. 39, 1166–1173 (2001).
Xie, Z. et al. Molecular phenotyping of aging in single yeast cells using a novel microfluidic device. Aging Cell 11, 599–606 (2012).
Jo, M. C., Liu, W., Gu, L., Dang, W. & Qin, L. High-throughput analysis of yeast replicative aging using a microfluidic system. Proc. Natl Acad. Sci. USA 112, 9364–9369 (2015).
Chen, C. et al. A high-throughput screening system for genes extending life-span. Exp. Gerontol. 38, 1051–1063 (2003).
Smeal, T., Claus, J., Kennedy, B., Cole, F. & Guarente, L. Loss of transcriptional silencing causes sterility in old mother cells of S. cerevisiae. Cell 84, 633–642 (1996).
Lindstrom, D. L. & Gottschling, D. E. The Mother Enrichment Program: a genetic system for facile replicative life span analysis in Saccharomyces cerevisiae. Genetics 183, 413–422 (2009).
Miesenböck, G., De Angelis, D. A. & Rothman, J. E. Visualizing secretion and synaptic transmission with pH-sensitive green fluorescent proteins. Nature 394, 192–195 (1998).
Gutscher, M. et al. Proximity-based protein thiol oxidation by H2O2-scavenging peroxidases. J. Biol. Chem. 284, 31532–31540 (2009).
Gutscher, M. et al. Real-time imaging of the intracellular glutathione redox potential. Nat. Methods 5, 553–559 (2008).
Brachmann, C. B. et al. Designer deletion strains derived from Saccharomyces cerevisiae S288C: a useful set of strains and plasmids for PCR-mediated gene disruption and other applications. Yeast 14, 115–132 (1998).
Dimitrov, L. N., Brem, R. B., Kruglyak, L. & Gottschling, D. E. Polymorphisms in multiple genes contribute to the spontaneous mitochondrial genome instability of Saccharomyces cerevisiae S288C strains. Genetics 183, 365–383 (2009).
Veatch, J. R., McMurray, M. A., Nelson, Z. W. & Gottschling, D. E. Mitochondrial dysfunction leads to nuclear genome instability via an iron-sulfur cluster defect. Cell 137, 1247–1258 (2009).
Veal, E. A., Ross, S. J., Malakasi, P., Peacock, E. & Morgan, B. A. Ybp1 is required for the hydrogen peroxide-induced oxidation of the Yap1 transcription factor. J. Biol. Chem. 278, 30896–30904 (2003).
Herrero, E., Ros, J., Bellí, G. & Cabiscol, E. Redox control and oxidative stress in yeast cells. Biochim. Biophys. Acta 1780, 1217–1235 (2008).
Okazaki, S., Naganuma, A. & Kuge, S. Peroxiredoxin-mediated redox regulation of the nuclear localization of Yap1, a transcription factor in budding yeast. Antioxid. Redox Signal. 7, 327–334 (2005).
Morgan, B. et al. Multiple glutathione disulfide removal pathways mediate cytosolic redox homeostasis. Nat. Chem. Biol. 9, 119–125 (2013).
Grant, C. M., Collinson, L. P., Roe, J. H. & Dawes, I. W. Yeast glutathione reductase is required for protection against oxidative stress and is a target gene for yAP-1 transcriptional regulation. Mol. Microbiol. 21, 171–179 (1996).
Tan, S.-X. et al. The thioredoxin-thioredoxin reductase system can function in vivo as an alternative system to reduce oxidized glutathione in Saccharomyces cerevisiae. J. Biol. Chem. 285, 6118–6126 (2010).
Ayer, A. et al. A genome-wide screen in yeast identifies specific oxidative stress genes required for the maintenance of sub-cellular redox homeostasis. PLoS ONE 7, e44278 (2012).
Harman, D. Aging: a theory based on free radical and radiation chemistry. J. Gerontol. 11, 298–300 (1956).
Orij, R., Brul, S. & Smits, G. J. Intracellular pH is a tightly controlled signal in yeast. Biochim. Biophys. Acta 1810, 933–944 (2011).
Giaever, G. et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature 418, 387–391 (2002).
Spragg, R. G., Hinshaw, D. B., Hyslop, P. A., Schraufstätter, I. U. & Cochrane, C. G. Alterations in adenosine triphosphate and energy charge in cultured endothelial and P388D1 cells after oxidant injury. J. Clin. Invest. 76, 1471–1476 (1985).
Colussi, C. et al. H2O2-induced block of glycolysis as an active ADP-ribosylation reaction protecting cells from apoptosis. FASEB J. 14, 2266–2276 (2000).
Osorio, H. et al. H2O2, but not menadione, provokes a decrease in the ATP and an increase in the inosine levels in Saccharomyces cerevisiae. An experimental and theoretical approach. Eur. J. Biochem. 270, 1578–1589 (2003).
Dechant, R. et al. Cytosolic pH is a second messenger for glucose and regulates the PKA pathway through V-ATPase. EMBO J. 29, 2515–2526 (2010).
Zhou, C. et al. Organelle-based aggregation and retention of damaged proteins in asymmetrically dividing cells. Cell 159, 530–542 (2014).
Veine, D. M., Arscott, L. D. & Williams, C. H. Redox potentials for yeast, Escherichia coli and human glutathione reductase relative to the NAD +/NADH redox couple: enzyme forms active in catalysis †. Biochemistry 37, 15575–15582 (1998).
Schafer, F. Q. & Buettner, G. R. Redox environment of the cell as viewed through the redox state of the glutathione disulfide/glutathione couple. Free Radic. Biol. Med. 30, 1191–1212 (2001).
Erjavec, N. & Nyström, T. Sir2p-dependent protein segregation gives rise to a superior reactive oxygen species management in the progeny of Saccharomyces cerevisiae. Proc. Natl Acad. Sci. USA 104, 10877–10881 (2007).
Brand, M. D. & Nicholls, D. G. Assessing mitochondrial dysfunction in cells. Biochem. J. 435, 297–312 (2011).
Halliwell, B. & Gutteridge, J. Free Radicals in Biology and Medicine (Oxford Univ. Press, 2007).
Leadsham, J. E. et al. Loss of cytochrome c oxidase promotes RAS-dependent ROS production from the ER resident NADPH oxidase, Yno1p, in yeast. Cell Metab. 18, 279–286 (2013).
Toledano, M. B., Delaunay-Moisan, A., Outten, C. E. & Igbaria, A. Functions and cellular compartmentation of the thioredoxin and glutathione pathways in yeast. Antioxid. Redox Signal. 18, 1699–1711 (2013).
Hoch, H. C., Galvani, C. D., Szarowski, D. H. & Turner, J. N. Two new fluorescent dyes applicable for visualization of fungal cell walls. Mycologia 97, 580–588 (2005).
Acknowledgements
The authors thank Dr T Dick, Dr B Morgan and Dr M Seedorf for plasmids and helpful discussions; Dr E Deuerling and Dr T Dick for critical reading of the manuscript; Dr M Langlotz for help in the FACS facility of the ZMBH; Dr H Lorenz in the imaging facility of the ZMBH; and S Hennes for excellent technical assistance. This work was supported by the Baden-Württemberg Stiftung (Network Aging Research) and the Deutsche Forschungsgemeinschaft (SFB1036 TP9 to MPM).
Author information
Authors and Affiliations
Contributions
RAK designed the study, performed all the experiments and was responsible for the construction of strains, analysis and interpretation of data, preparation of figures and also wrote the manuscript. MPM was responsible for the design of the study, analysis and interpretation of data, preparation of figures and also wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the npj Aging and Mechanisms of Disease website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Knieß, R., Mayer, M. The oxidation state of the cytoplasmic glutathione redox system does not correlate with replicative lifespan in yeast. npj Aging Mech Dis 2, 16028 (2016). https://doi.org/10.1038/npjamd.2016.28
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/npjamd.2016.28