Abstract
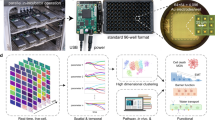
Screening methods that use traditional genomic1,2,3, transcriptional4, proteomic5,6 and metabonomic7 signatures to characterize drug mechanisms are known. However, they are time consuming and require specialized equipment. Here, we present a high-throughput multichannel sensor platform that can profile the mechanisms of various chemotherapeutic drugs in minutes. The sensor consists of a gold nanoparticle complexed with three different fluorescent proteins that can sense drug-induced physicochemical changes on cell surfaces8,9,10. In the presence of cells, fluorescent proteins are rapidly displaced from the gold nanoparticle surface and fluorescence is restored. Fluorescence ‘turn on’ of the fluorescent proteins depends on the drug-induced cell surface changes, generating patterns that identify specific mechanisms of cell death induced by drugs. The nanosensor is generalizable to different cell types and does not require processing steps before analysis, offering an effective way to expedite research in drug discovery, toxicology and cell-based sensing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lamb, J. et al. The connectivity map: using gene-expression signatures to connect small molecules, genes, and disease. Science 313, 1929–1935 (2006).
Jiang, H., Pritchard, J. R., Williams, R. T., Lauffenburger, D. A. & Hemann, M. T. A mammalian functional-genetic approach to characterizing cancer therapeutics. Nature Chem. Biol. 7, 92–100 (2011).
Parsons, A. B. et al. Exploring the mode-of-action of bioactive compounds by chemical-genetic profiling in yeast. Cell 126, 611–625 (2006).
Butcher, R. A. & Schreiber, S. L. Using genome-wide transcriptional profiling to elucidate small-molecule mechanism. Curr. Opin. Chem. Biol. 9, 25–30 (2005).
Schirle, M., Bantscheff, M. & Kuster, B. Mass spectrometry-based proteomics in preclinical drug discovery. Chem. Biol. 19, 72–84 (2012).
Krutzik, P. O., Crane, J. M., Clutter, M. R. & Nolan, G. P. High-content single-cell drug screening with phosphospecific flow cytometry. Nature Chem. Biol. 4, 132–142 (2008).
Nicholson, J. K., Connelly, J., Lindon, J. C. & Holmes, E. Metabonomics: a platform for studying drug toxicity and gene function. Nature Rev. Drug Discov. 1, 153–161 (2002).
Chan, D. & Lee, S-C. in Oncogene and Cancer—From Bench to Clinic (ed. Siregar, Y.) Ch. 1 (InTech, 2013).
Obeid, M. et al. Calreticulin exposure dictates the immunogenicity of cancer cell death. Nature Med. 13, 54–61 (2007).
Azuma, Y., Taniguchi, A. & Matsumoto, K. Decrease in cell surface sialic acid in etoposide-treated Jurkat cells and the role of cell surface sialidase. Glycoconj. J. 17, 301–306 (2000).
Editorial. Mechanism matters. Nature Med. 16, 347 (2010).
De Castro, D. G., Clarke, P. A., Al-Lazikani, B. & Workman, P. Personalized cancer medicine: molecular diagnostics, predictive biomarkers, and drug resistance. Clin. Pharmacol. Ther. 93, 252–259 (2013).
Feng, Y., Mitchison, T. J., Bender, A., Young, D. W. & Tallarico, J. A. Multi-parameter phenotypic profiling: using cellular effects to characterize small-molecule compounds. Nature Rev. Drug Discov. 8, 567–578 (2009).
Perlman, Z. E. et al. Multidimensional drug profiling by automated microscopy. Science 306, 1194–1198 (2004).
Rihel, J. et al. Zebrafish behavioral profiling links drugs to biological targets and rest/wake regulation. Science 327, 348–351 (2010).
Young, D. W. et al. Integrating high-content screening and ligand–target prediction to identify mechanism of action. Nature Chem. Biol. 4, 59–68 (2008).
Bajaj, A. et al. Detection and differentiation of normal, cancerous, and metastatic cells using nanoparticle–polymer sensor arrays. Proc. Natl Acad. Sci. USA 106, 10912–10916 (2009).
El-Boubbou, K. et al. Magnetic glyco-nanoparticles: a tool to detect, differentiate, and unlock the glyco-codes of cancer via magnetic resonance imaging. J. Am. Chem. Soc. 132, 4490–4499 (2010).
Wright, A. T. & Anslyn, E. V. Differential receptor arrays and assays for solution-based molecular recognition. Chem. Soc. Rev. 35, 14–28 (2006).
Peng, G. et al. Diagnosing lung cancer in exhaled breath using gold nanoparticles. Nature Nanotech. 4, 669–673 (2009).
Tanaka, M. et al. An unbiased cell morphology-based screen for new, biologically active small molecules. PLoS Biol. 3, e128 (2005).
Rana, S. et al. Array-based sensing of metastatic cells and tissues using nanoparticle–fluorescent protein conjugates. ACS Nano 9, 8233–8240 (2012).
Shaner, N. C., Steinbach, P. A. & Tsien, R. Y. A guide to choosing fluorescent proteins. Nature Methods 2, 905–909 (2005).
Vermes, I., Haanen, C. & Reutelingsperger, C. Flow cytometry of apoptotic cell death. J. Immunol. Methods 243, 167–190 (2000).
Reis-Filho, J. S. & Tutt, A. N. J. Triple negative tumours: a critical review. Histopathology 52, 108–118 (2008).
Saha, K. et al. Surface functionality of nanoparticles determines cellular uptake mechanisms in mammalian cells. Small 9, 300–305 (2013).
Pritchard, J. R., Bruno, P. M., Hemanna, M. T. & Lauffenburger, D. A. Predicting cancer drug mechanisms of action using molecular network signatures. Mol. Biosyst. 9, 1604–1619 (2013).
Dunphy, K. A. et al. Oncogenic transformation of mammary epithelial cells by transforming growth factor beta independent of mammary stem cell regulation. Cancer Cell Int. 13, 74 (2013).
Dancey, J. E. & Chen, H. X. Strategies for optimizing combinations of molecularly targeted anticancer agents. Nature Rev. Drug Discov. 5, 649–659 (2006).
Gottesman, M. M. Mechanisms of cancer drug resistance. Annu. Rev. Med. 53, 615–627 (2002).
Jia, J. et al. Mechanisms of drug combinations: interaction and network perspectives. Nature Rev. Drug Discov. 8, 111–128 (2009).
Pritchard, J. R. et al. Defining principles of combination drug mechanisms of action. Proc. Natl Acad. Sci. USA 110, E170–E179 (2013).
Geva-Zatorsky, N. et al. Protein dynamics in drug combinations: a linear superposition of individual-drug responses. Cell 140, 643–651 (2010).
Nandakumar, D. N., Nagaraj, V. A., Vathsala, P. G., Rangarajan, P. & Padmanaban, G. Curcumin–artemisinin combination therapy for malaria. Antimicrob. Agents Chemother. 50, 1859–1860 (2006).
Chan, L-P. et al. Apigenin induces apoptosis via tumor necrosis factor receptor- and Bcl-2-mediated pathway and enhances susceptibility of head and neck squamous cell carcinoma to 5-fluorouracil and cisplatin. Biochim. Biophys. Acta Gen. Subjects 1820, 1081–1091 (2012).
Acknowledgements
The authors thank D. Joseph Jerry for providing the pTD cell line and C. Stanier for reading the manuscript and providing useful suggestions. This work was supported by National Institutes of Health grant GM077173 and National Science Foundation Center for Hierarchical Manufacturing grant CMMI-1025020.
Author information
Authors and Affiliations
Contributions
S.R. and V.M.R. conceived the concepts. S.R. designed the experiments. O.R.M. and G.Y. synthesized and characterized the nanoparticle. R.M. and S.R. expressed the fluorescent proteins. S.R., N.L., R.M. and K.S. performed the drug screening studies. S.R., R.M. and N.L. analysed the data, with statistical inputs from C.R. and R.B. S.R. and V.M.R. wrote most of the paper, with contribution from the other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary Information (PDF 3295 kb)
Rights and permissions
About this article
Cite this article
Rana, S., Le, N., Mout, R. et al. A multichannel nanosensor for instantaneous readout of cancer drug mechanisms. Nature Nanotech 10, 65–69 (2015). https://doi.org/10.1038/nnano.2014.285
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2014.285
This article is cited by
-
VIBRANT: spectral profiling for single-cell drug responses
Nature Methods (2024)
-
Immobilization of azide-functionalized proteins to micro- and nanoparticles directly from cell lysate
Microchimica Acta (2024)
-
Accurate identification of kidney injury progression via a fluorescent biosensor array
Microchimica Acta (2022)
-
Anti-cancer adjuvant drug screening via epithelial-mesenchymal transition-related aptamer probe
Analytical and Bioanalytical Chemistry (2021)
-
Biomolecule-templated photochemical synthesis of silver nanoparticles: Multiple readouts of localized surface plasmon resonance for pattern recognition
Nano Research (2018)