Abstract
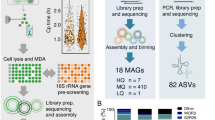
Our view of microbial diversity has expanded greatly over the past 40 years, primarily through the wide application of PCR-based surveys of the small-subunit ribosomal RNA (SSU rRNA) gene. Yet significant gaps in knowledge remain due to well-recognized limitations of this method. Here, we systematically survey primer fidelity in SSU rRNA gene sequences recovered from over 6,000 assembled metagenomes sampled globally. Our findings show that approximately 10% of environmental microbial sequences might be missed from classical PCR-based SSU rRNA gene surveys, mostly members of the Candidate Phyla Radiation (CPR) and as yet uncharacterized Archaea. These results underscore the extent of uncharacterized microbial diversity and provide fruitful avenues for describing additional phylogenetic lineages.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Pace, N. R. Mapping the tree of life: Progress and prospects. Microbiol. Mol. Biol. Rev. 73, 565–576 (2009).
Caporaso, J. G. et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 6, 1621–1624 (2012).
Yarza, P. et al. Uniting the classification of cultured and uncultured bacteria and archaea using 16S rRNA gene sequences. Nature Rev. Microbiol. 12, 635–645 (2014).
Brown, C. T. et al. Unusual biology across a group comprising more than 15% of domain Bacteria. Nature 523, 208–211 (2015).
Castelle, C. J. et al. Genomic expansion of domain archaea highlights roles for organisms from new phyla in anaerobic carbon cycling. Curr. Biol. 25, 690–701 (2015).
Rinke, C. et al. Insights into the phylogeny and coding potential of microbial dark matter. Nature 499, 431–437 (2013).
Lynch, M. D. J. & Neufeld, J. D. Ecology and exploration of the rare biosphere. Nature Rev. Microbiol. 13, 217–229 (2015).
Engelbrektson, A. et al. Experimental factors affecting PCR-based estimates of microbial species richness and evenness. ISME J. 4, 642–647 (2010).
Woyke, T. & Rubin, E. M. Searching for new branches on the tree of life. Science 346, 698–699 (2014).
Markowitz, V. M. et al. IMG/M 4 version of the integrated metagenome comparative analysis system. Nucleic Acids Res. 42, D568–D573 (2014).
Gilbert, J. A. et al. The Earth Microbiome Project: Meeting report of the ‘1st EMP meeting on sample selection and acquisition’ at Argonne National Laboratory October 6th 2010. Stand. Genomic Sci. 3, 249–253 (2010).
Parada, A., Needham, D. M. & Fuhrman, J. A. Every base matters: assessing small subunit rRNA primers for marine microbiomes with mock communities, time-series and global field samples. Environ. Microbiol. http://dx.doi.org/10.1111/1462-2920.13023 (2015).
Klindworth, A. et al. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 41, e1 (2013).
The Human Microbiome Consortium. Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214 (2012).
Spang, A. et al. Complex archaea that bridge the gap between prokaryotes and eukaryotes. Nature 521, 173–179 (2015).
Albertsen, M. et al. Genome sequences of rare, uncultured bacteria obtained by differential coverage binning of multiple metagenomes. Nature Biotechnol. 31, 533–538 (2013).
Walters, W. A. et al. PrimerProspector: de novo design and taxonomic analysis of barcoded polymerase chain reaction primers. Bioinformatics 27, 1159–1161 (2011).
Kent, W. BLAT—The BLAST-Like Alignment Tool. Genome Res. 12, 656–664 (2002).
Nawrocki, E. P. & Eddy, S. R. Infernal 1.1: 100-fold faster RNA homology searches. Bioinformatics 29, 2933–2935 (2013).
Stamatakis, A. RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22, 2688–2690 (2006).
Crooks, G. E. et al. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Acknowledgements
This work was conducted by the US Department of Energy Joint Genome Institute, a DOE Office of Science User Facility (contract no. DE-AC02-05CH11231), and used resources of the National Energy Research Scientific Computing Center, which is supported by the Office of Science of the US Department of Energy (contract no. DE-AC02-05CH11231).
Author information
Authors and Affiliations
Contributions
E.A.E.-F., N.N.I., T.W. and N.C.K. designed the project, analysed the data and wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1–3, Supplementary Tables 1,2. (PDF 273 kb)
Rights and permissions
About this article
Cite this article
Eloe-Fadrosh, E., Ivanova, N., Woyke, T. et al. Metagenomics uncovers gaps in amplicon-based detection of microbial diversity. Nat Microbiol 1, 15032 (2016). https://doi.org/10.1038/nmicrobiol.2015.32
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nmicrobiol.2015.32
This article is cited by
-
Alteration of taste perception, food neophobia and oral microbiota composition in children with food allergy
Scientific Reports (2023)
-
Gut dysbiosis in Thai intrahepatic cholangiocarcinoma and hepatocellular carcinoma
Scientific Reports (2023)
-
ASAP 2: a pipeline and web server to analyze marker gene amplicon sequencing data automatically and consistently
BMC Bioinformatics (2022)
-
Dissecting the dominant hot spring microbial populations based on community-wide sampling at single-cell genomic resolution
The ISME Journal (2022)
-
Longitudinal analysis of the Five Sisters hot springs in Yellowstone National Park reveals a dynamic thermoalkaline environment
Scientific Reports (2022)