Abstract
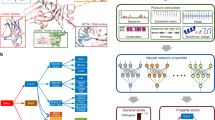
The baker's yeast mutation collections are extensively used genetic resources that are the basis for many genome-wide screens and new technologies. Anecdotal evidence has previously pointed to the putative existence of a neighboring gene effect (NGE) in these collections. NGE occurs when the phenotype of a strain carrying a particular perturbed gene is due to the lack of proper function of its adjacent gene. Here we performed a large-scale study of NGEs, presenting a network-based algorithm for detecting NGEs and validating software predictions using complementation experiments. We applied our approach to four datasets uncovering a similar magnitude of NGE in each (7–15%). These results have important consequences for systems biology, as the mutation collections are extensively used in almost every aspect of the field, from genetic network analysis to functional gene annotation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hillenmeyer, M.E. et al. The chemical genomic portrait of yeast: uncovering a phenotype for all genes. Science 320, 362–365 (2008).
Hughes, T.R. et al. Functional discovery via a compendium of expression profiles. Cell 102, 109–126 (2000).
Costanzo, M. et al. The genetic landscape of a cell. Science 327, 425–431 (2010).
Winzeler, E.A. et al. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 285, 901–906 (1999).
Tong, A.H. et al. Systematic genetic analysis with ordered arrays of yeast deletion mutants. Science 294, 2364–2368 (2001).
Pan, X. et al. A DNA integrity network in the yeast Saccharomyces cerevisiae. Cell 124, 1069–1081 (2006).
Addinall, S.G. et al. A genomewide suppressor and enhancer analysis of cdc13–1 reveals varied cellular processes influencing telomere capping in Saccharomyces cerevisiae. Genetics 180, 2251–2266 (2008).
Tian, W. et al. Combining guilt-by-association and guilt-by-profiling to predict Saccharomyces cerevisiae gene function. Genome Biol. 9 (suppl. 1), S7 (2008).
Kelley, R. & Ideker, T. Systematic interpretation of genetic interactions using protein networks. Nat. Biotechnol. 23, 561–566 (2005).
van Wageningen, S. et al. Functional overlap and regulatory links shape genetic interactions between signaling pathways. Cell 143, 991–1004 (2010).
Breslow, D.K. et al. A comprehensive strategy enabling high-resolution functional analysis of the yeast genome. Nat. Methods 5, 711–718 (2008).
Ben-Aroya, S. et al. Toward a comprehensive temperature-sensitive mutant repository of the essential genes of Saccharomyces cerevisiae. Mol. Cell 30, 248–258 (2008).
Li, Z. et al. Systematic exploration of essential yeast gene function with temperature-sensitive mutants. Nat. Biotechnol. 29, 361–367 (2011).
Shachar, R., Ungar, L., Kupiec, M., Ruppin, E. & Sharan, R. A systems-level approach to mapping the telomere length maintenance gene circuitry. Mol. Syst. Biol. 4, 172 (2008).
Ungar, L. et al. A genome-wide screen for essential yeast genes that affect telomere length maintenance. Nucleic Acids Res. 37, 3840–3849 (2009).
Yosef, N. et al. Toward accurate reconstruction of functional protein networks. Mol. Syst. Biol. 5, 248 (2009).
Dittrich, M., Klau, G., Rosenwald, A., Dandekar, T. & Müller, T. Identifying functional modules in protein-protein interaction networks: an integrated exact approach. Bioinformatics 24, i223–i231 (2008).
Huang, S.S. & Fraenkel, E. Integrating proteomic, transcriptional, and interactome data reveals hidden components of signaling and regulatory networks. Sci. Signal. 2, ra40 (2009).
Yosef, N. et al. ANAT–a software tool for reconstructing and analyzing functional networks of proteins. Sci. Signal. 4, l1 (2011).
Askree, S.H. et al. A genome-wide screen for Saccharomyces cerevisiae deletion mutants that affect telomere length. Proc. Natl. Acad. Sci. USA 101, 8658–8663 (2004).
Gatbonton, T. et al. Telomere length as a quantitative trait: genome-wide survey and genetic mapping of telomere length-control genes in yeast. PLoS Genet. 2, e35 (2006).
Parsons, A.B. et al. Integration of chemical-genetic and genetic interaction data links bioactive compounds to cellular target pathways. Nat. Biotechnol. 22, 62–69 (2004).
Chan, T.F., Carvalho, J., Riles, L. & Zheng, X.F. A chemical genomics approach toward understanding the global functions of the target of rapamycin protein (TOR). Proc. Natl. Acad. Sci. USA 97, 13227–13232 (2000).
Crespo, J.L. & Hall, M.N. Elucidating TOR signaling and rapamycin action: lessons from Saccharomyces cerevisiae. Microbiol. Mol. Biol. Rev. 66, 579–591 (2002).
Reid, R.J. et al. Selective ploidy ablation, a high-throughput plasmid transfer protocol, identifies new genes affecting topoisomerase I-induced DNA damage. Genome Res. 21, 477–486 (2011).
Gustavsson, M. & Ronne, H. Evidence that tRNA modifying enzymes are important in vivo targets for 5-fluorouracil in yeast. RNA 14, 666–674 (2008).
Tuller, T. et al. Higher-order genomic organization of cellular functions in yeast. J. Comput. Biol. 16, 303–316 (2009).
Xu, Z. et al. Bidirectional promoters generate pervasive transcription in yeast. Nature 457, 1033–1037 (2009).
Snyder, M. & Gallagher, J.E. Systems biology from a yeast omics perspective. FEBS Lett. 583, 3895–3899 (2009).
Sharan, R. et al. Conserved patterns of protein interaction in multiple species. Proc. Natl. Acad. Sci. USA 102, 1974–1979 (2005).
Gavin, A.C. et al. Functional organization of the yeast proteome by systematic analysis of protein complexes. Nature 415, 141–147 (2002).
Krogan, N.J. et al. Global landscape of protein complexes in the yeast Saccharomyces cerevisiae. Nature 440, 637–643 (2006).
Ito, T. et al. A comprehensive two-hybrid analysis to explore the yeast protein interactome. Proc. Natl. Acad. Sci. USA 98, 4569–4574 (2001).
Uetz, P. et al. A comprehensive analysis of protein-protein interactions in Saccharomyces cerevisiae. Nature 403, 623–627 (2000).
Reguly, T. et al. Comprehensive curation and analysis of global interaction networks in Saccharomyces cerevisiae. J. Biol. 5, 11 (2006).
Breitkreutz, B.J. et al. The BioGRID Interaction Database: 2008 update. Nucleic Acids Res. 36 Database issue, D637–D640 (2008).
Yosef, N. et al. ANAT: a tool for constructing and analyzing functional protein networks. Sci. Signal. 4, pl1 (2011).
Charikar, M. et al. Approximation algorithms for directed Steiner problems. J. Algorithms 33, 73–91 (1999).
Acknowledgements
We thank current and past members of the Sharan, Ruppin and Kupiec laboratories for helpful discussions, and D. Shore, R. Rolfes, B. Cairns, A. Johnson, S. Harashima, A.S. Bystrom, S. Moye-Rowley, M. Johnston, M.A. Clayton, G.H. Braus, B. Polevoda, R. Movva, E. Alani, J.P. Gélugne, V. Measday, D. Ramotar, A. Munn, E. Miller, B.C. Laurent, C. Brenner, M. Polymenis, J. Gerst, L. Aragon, D. Spatt, J. Boeke, G. Brown, C. Burd, M. Tamas, D.H. Wolf, Y. Hannun, Y. Ohsumi, Z. Ciesla, M. Choder, E. Bi, M. Bard, R. Schaffrath, H.U. Mosch, A. Amon, R. Parker, Y. Saeki, J. Kim, H.O. Park, E. Etsuchi, H. Nakatogana, B. Daignan-Fornier for providing plasmids. This work was supported by grants from the Israeli Ministry of Science and Technology and the Israel Cancer Foundation to M.K., and by a grant from the James McDonnel Fund to M.K., R.S. and E.R.; E.R. and R.S. were also supported by a Bikura grant from the Israel Science Foundation.
Author information
Authors and Affiliations
Contributions
T.B.-S., N.Y., R.S., E.R. and M.K. conceived and designed the experiments and wrote the paper. T.B.-S. and K.S. carried out the wet laboratory experiments. N.Y. analyzed the data obtained. R.S. and E.R. contributed equally to this project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 2, 5 and 7–9 (PDF 874 kb)
Supplementary Table 1
NGE analysis of the TLM set. (XLS 107 kb)
Supplementary Table 3
Complexes that participate in the TLM and RR sets. (XLS 44 kb)
Supplementary Table 4
NGE analysis of the RR set. (XLS 102 kb)
Supplementary Table 6
Predicted and verified NGE in the TLM set. (XLS 37 kb)
Supplementary Software
NIRVANA software. (ZIP 2239 kb)
Rights and permissions
About this article
Cite this article
Ben-Shitrit, T., Yosef, N., Shemesh, K. et al. Systematic identification of gene annotation errors in the widely used yeast mutation collections. Nat Methods 9, 373–378 (2012). https://doi.org/10.1038/nmeth.1890
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1890