Abstract
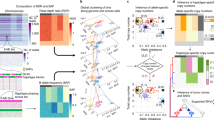
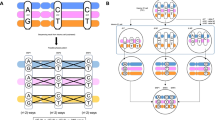
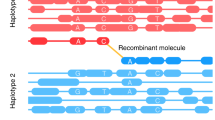
Direct observation of haplotypes is still technically challenging. Here we report a method for the determination of haplotypes through chromosome microdissection. We determined human long-range chromosomal haplotypes with more than 98.85% accuracy at 24,245 genome-wide heterozygous single-nucleotide polymorphism (SNP) loci.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Levenstien, M.A., Ott, J. & Gordon, D. PLoS Genet. 2, e127 (2006).
Marchini, J. et al. Am. J. Hum. Genet. 78, 437–450 (2006).
Stephens, M. & Donnelly, P. Am. J. Hum. Genet. 73, 1162–1169 (2003).
Ragoussis, J. Annu. Rev. Genomics Hum. Genet. 10, 117–133 (2009).
Xiao, M. et al. Nat. Methods 6, 199–201 (2009).
Yan, H. et al. Nature 403, 723–724 (2000).
Douglas, J.A., Boehnke, M., Gillanders, E., Trent, J.M. & Gruber, S.B. Nat. Genet. 28, 361–364 (2001).
Mitra, R.D. et al. Proc. Natl. Acad. Sci. USA 100, 5926–5931 (2003).
Zhang, K. et al. Nat. Genet. 38, 382–387 (2006).
Liu, N., Zhang, K. & Zhao, H. Adv. Genet. 60, 335–405 (2008).
The International HapMap Consortium. Nature 426, 789–796 (2003).
Hodge, S.E., Boehnke, M. & Spence, M.A. Nat. Genet. 21, 360–361 (1999).
Highsmith, W.E. Jr., Meyer, K.J., Marley, V.M. & Jenkins, R.B. In Curr. Protoc. Hum. Genet. 3.6 (John Wiley and Sons, Inc., 2007).
Tjio, J.H. & Levan, A. Hereditas 42, 1–6 (1956).
Jurka, J. Trends Genet. 16, 418–420 (2000).
Acknowledgements
This work was supported by US National Institutes of Health grants (HL003676, RR014758, RR003034, NS45012 and HG004436), an American Heart Association grant (09GRNT2300003) and in part by the Medical Research Service of the Department of Veterans Affairs. K.Z. was supported by National Institutes of Health (GM074913). We thank S.J. Kittner, R.A. Kittles, G. Newman and G.H. Gibbons for reading and discussing the manuscript, and the people who donated blood samples and members of the International HapMap project.
Author information
Authors and Affiliations
Contributions
L.M. performed experiments and data analysis. Y.X. and Q.W. assisted with microdissection experiments. H.H. assisted with cytogenetic experiments. L.M. and W.R. performed computational data analysis. Y.F. contributed to data analysis and manuscript preparation. K.Z. conducted statistical analysis. Q.S. designed the experiments, performed data analysis and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4, Supplementary Tables 1, 3–7 and Supplementary Notes 1–2 (PDF 227 kb)
Supplementary Table 2
The 7DDNA haplotype report. (XLS 5608 kb)
Rights and permissions
About this article
Cite this article
Ma, L., Xiao, Y., Huang, H. et al. Direct determination of molecular haplotypes by chromosome microdissection. Nat Methods 7, 299–301 (2010). https://doi.org/10.1038/nmeth.1443
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1443
This article is cited by
-
Island-specific evolution of a sex-primed autosome in a sexual planarian
Nature (2022)
-
Experimental method for haplotype phasing across the entire length of chromosome 21 in trisomy 21 cells using a chromosome elimination technique
Journal of Human Genetics (2022)
-
Isolation and sequencing of a single copy of an introgressed chromosome from a complex genome for gene and SNP identification
Theoretical and Applied Genetics (2022)
-
Determination of complete chromosomal haplotypes by bulk DNA sequencing
Genome Biology (2021)
-
Cataloguing experimentally confirmed 80.7 kb-long ACKR1 haplotypes from the 1000 Genomes Project database
BMC Bioinformatics (2021)