Abstract
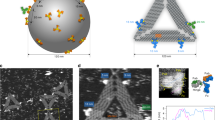
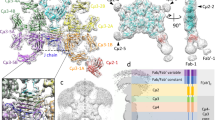
The conformational flexibility of antibodies in solution directly affects their immune function. Namely, the flexible hinge regions of immunoglobulin G (IgG) antibodies are essential in epitope-specific antigen recognition and biological effector function. The antibody structure, which is strongly related to its functions, has been partially revealed by electron microscopy1,2,3,4 and X-ray crystallography5,6, but only under non-physiological conditions. Here we observed monoclonal IgG antibodies in aqueous solution by high-resolution frequency modulation atomic force microscopy7,8 (FM-AFM). We found that monoclonal antibodies self-assemble into hexamers, which form two-dimensional crystals in aqueous solution. Furthermore, by directly observing antibody–antigen interactions using FM-AFM, we revealed that IgG molecules in the crystal retain immunoactivity. As the self-assembled monolayer crystal of antibodies retains immunoactivity at a neutral pH and is functionally stable at a wide range of pH and temperature, the antibody crystal is applicable to new biotechnological platforms for biosensors or bioassays.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Feinstein, A. & Rowe, A. J. Molecular mechanism of formation of an antigen–antibody complex. Nature 205, 147–149 (1965).
Valentine, R. C. & Green, N. M. Electron microscopy of an antibody–hapten complex. J. Mol. Biol. 27, 615–617 (1967).
Lansdorp, P. M., Aalberse, R. C., Bos, R., Schutter, W. G. & Van Bruggen, E. F. J. Cyclic tetramolecular complexes of monoclonal antibodies: A new type of cross-linking reagent. Eur. J. Immunol. 16, 679–683 (1986).
Roux, K. H. Immunoglobulin structure and function as revealed by electron microscopy. Int. Arch. Allergy Immunol. 120, 85–99 (1999).
Harris, L. J. et al. The three-dimensional structure of an intact monoclonal antibody for canine lymphoma. Nature 360, 369–372 (1992).
Harris, L. J., Skaletsky, E. & McPherson, A. Crystallographic structure of an intact IgG1 monoclonal antibody. J. Mol. Biol. 275, 861–872 (1998).
Albrecht, T. R., Grütter, P., Horne, D. & Rugar, D. Frequency modulation detection using high- Q cantilevers for enhanced force microscope sensitivity. J. Appl. Phys. 69, 668–673 (1991).
Giessibl, F. J. Advances in atomic force microscopy. Rev. Mod. Phys. 75, 949–983 (2003).
Binnig, G., Quate, C. F. & Gerber, C. Atomic force microscope. Phys. Rev. Lett. 56, 930–933 (1986).
Müller, D. J. & Engel, A. The height of biomolecules measured with the atomic force microscope depends on electrostatic interactions. Biophys. J. 73, 1633–1644 (1997).
Müller, D. J., Fotiadis, D. & Engel, A. Mapping flexible protein domains at subnanometre resolution with the atomic force microscope. FEBS Lett. 430, 105–111 (1998).
Müller, D. J., Fotiadis, D., Scheuring, S., Müller, S. A. & Engel, A. Electrostatically balanced subnanometre imaging of biological specimens by atomic force microscope. Biophys. J. 76, 1101–1111 (1999).
Müller, D. J. & Dufrene, Y. F. Atomic force microscopy as a multifunctional molecular toolbox in nanobiotechnology. Nature Nanotech. 3, 261–269 (2008).
Allison, D. P., Mortensen, N. P., Sullivan, C. J. & Doktycz, M. J. Atomic force microscopy of biological samples. WIREs Nanomed. Nanobiotechnol. 2, 618–634 (2010).
Zhang, Y., Sheng, S. & Shao, Z. Imaging biological structures with the cryo atomic force microscope. Biophys. J. 71, 2168–2176 (1996).
Hansma, H. G. Varieties of imaging with scanning probe microscopes. Proc. Natl Acad. Sci. USA 96, 14678–14680 (1999).
Cheung, C. L., Hafner, J. H. & Lieber, C. M. Carbon nanotube atomic force microscopy tips: Direct growth by chemical vapour deposition and application to high-resolution imaging. Proc. Natl Acad. Sci. USA 97, 3809–3813 (2000).
Kienberger, F., Mueller, H., Pastushenko, V. & Hinterdorfer, P. Following single antibody binding to purple membranes in real time. EMBO Rep. 5, 579–583 (2004).
Thomson, N. H. The substructure of immunoglobulin G resolved to 25 kDa using amplitude modulation AFM in air. Ultramicroscopy 105, 103–110 (2005).
Preiner, J. et al. Imaging and detection of single molecule recognition events on organic semiconductor surfaces. Nano Lett. 9, 571–575 (2008).
Martinez, N. F. et al. Bimodal atomic force microscopy imaging of isolated antibodies in air and liquids. Nanotechnology 19, 384011 (2008).
Makky, A. et al. Substructures high resolution imaging of individual IgG and IgM antibodies with piezoelectric tuning fork atomic force microscopy. Sens. Actuat. B 162, 269–277 (2011).
Fukuma, T., Kobayashi, K., Matsushige, K. & Yamada, H. True atomic resolution in liquid by frequency-modulation atomic force microscopy. Appl. Phys. Lett. 87, 034101 (2005).
Ido, S. et al. Beyond the helix pitch: Direct visualization of native DNA in aqueous solution. ACS Nano 7, 1817–1822 (2013).
Hansma, H. G. & Laney, D. E. DNA binding to mica correlates with cationic radius: Assay by atomic force microscopy. Biophys. J. 70, 1933–1939 (1996).
Padlan, E. A. Anatomy of the antibody molecule. Mol. Immunol. 31, 169–217 (1994).
Pinteric, L., Painter, R. H. & Connell, G. E. Ultrastructure of the Fc fragment of human immunonoglobulin G. Immunochemistry 8, 1041–1045 (1971).
Burton, D. R. Immunoglobulin G: Functional sites. Mol. Immunol. 22, 161–206 (1985).
Yagi, H., Takahashi, N., Yamaguchi, Y. & Kato, K. Temperature-dependent isologous Fab–Fab interaction that mediates cryocrystallization of a monoclonal immunoglobulin G. Mol. Immunol. 41, 1211–1215 (2004).
Kanai, S., Liu, J., Patapoff, T. W. & Shire, S. J. Reversible self-association of a concentrated monoclonal antibody solution mediated by Fab–Fab interaction that impacts solution viscosity. J. Pharm. Sci. 97, 4219–4227 (2008).
Reidler, J., Uzgiris, E. E. & Kornberg, R. D. in Handbook of Experimental Immunology Vol. 1 (eds Ouchterlony, O., Nilsson, L. & Weir, D. M.) Ch. 17, 17.11–17.15 (Blackwell, 1967).
Uzgiris, E. E. & Kornberg, R. D. Two-dimensional crystallization technique for imaging macromolecules, with application to antigen–antibody–complement complexes. Nature 301, 125–129 (1983).
Kuznetsov, Y. G., Day, J., Newman, R. & McPherson, A. Chimeric human–simian anti-CD4 antibodies form crystalline high symmetry particles. J. Struct. Biol. 131, 108–115 (2000).
Horcas, I. et al. WSXM: A software for scanning probe microscopy and a tool for nanotechnology. Rev. Sci. Instrum. 78, 013705 (2007).
Acknowledgements
This work was supported by Grants-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science and Technology of Japan, the SENTAN Program of the Japan Science and Technology Agency, and the Global COE Program of the Japanese Society for the Promotion of Science. S.I. was supported by a Japan Society for the Promotion of Science Fellowship.
Author information
Authors and Affiliations
Contributions
S.I. performed AFM imaging, analysed data and wrote the paper. H. Kimiya prepared the biological sample and wrote the paper. K.K. developed AFM instruments and electronics and wrote the paper. H. Kominami performed AFM imaging. K.M. designed the study. H.Y. designed the study, developed AFM instruments, analysed the data and wrote the paper. All authors have discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1461 kb)
Rights and permissions
About this article
Cite this article
Ido, S., Kimiya, H., Kobayashi, K. et al. Immunoactive two-dimensional self-assembly of monoclonal antibodies in aqueous solution revealed by atomic force microscopy. Nature Mater 13, 264–270 (2014). https://doi.org/10.1038/nmat3847
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmat3847
This article is cited by
-
Isolation of exosomes from whole blood by a new microfluidic device: proof of concept application in the diagnosis and monitoring of pancreatic cancer
Journal of Nanobiotechnology (2020)
-
Assembly of a patchy protein into variable 2D lattices via tunable multiscale interactions
Nature Communications (2020)
-
Capturing transient antibody conformations with DNA origami epitopes
Nature Communications (2020)
-
Molecular-scale visualization and surface charge density measurement of Z-DNA in aqueous solution
Scientific Reports (2019)
-
Enhancement of Capturing Efficacy for Circulating Tumor Cells by Centrifugation
BioChip Journal (2018)