Abstract
Genome-wide association studies (GWAS) have identified more than 80 susceptibility loci for type 2 diabetes (T2D), but most of its heritability still remains to be elucidated. In this study, we conducted a meta-analysis of GWAS for T2D in the Japanese population. Combined data from discovery and subsequent validation analyses (23,399 T2D cases and 31,722 controls) identify 7 new loci with genome-wide significance (P<5 × 10−8), rs1116357 near CCDC85A, rs147538848 in FAM60A, rs1575972 near DMRTA1, rs9309245 near ASB3, rs67156297 near ATP8B2, rs7107784 near MIR4686 and rs67839313 near INAFM2. Of these, the association of 4 loci with T2D is replicated in multi-ethnic populations other than Japanese (up to 65,936 T2Ds and 158,030 controls, P<0.007). These results indicate that expansion of single ethnic GWAS is still useful to identify novel susceptibility loci to complex traits not only for ethnicity-specific loci but also for common loci across different ethnicities.
Similar content being viewed by others
Introduction
To date, more than 80 susceptibility loci for type 2 diabetes (T2D) have been identified through genome-wide association studies (GWAS)1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18. However, the joint effects of these variants account for <10% of the heritability for T2D10,19. GWAS for T2D have been extensively conducted in populations of European descent and, accordingly, the majority of established T2D susceptibility genetic loci were originally identified by European GWAS1,2,8,9,10,11,16. Cumulative evidence suggests that Asian populations may be more genetically susceptible to T2D than populations with European ancestry20. In addition, there are significant interethnic differences in the risk allele frequency or in effect sizes at several loci, which may affect the power to detect associations in these populations2. On the other hand, both overlap in T2D susceptibility loci among different ancestry groups and coincident risk alleles at lead single-nucleotide polymorphisms (SNPs) across diverse populations have been reported, suggesting that causal variants at many of these loci are shared across different ancestry groups12. Moreover, a recently published transethnic GWAS has successfully identified seven novel T2D susceptibility loci by combining the association data from European, South Asian, East Asian and Mexican/Latinos GWAS12. Therefore, it is valuable to perform GWAS for T2D using non-European and European populations, to facilitate identification of both ethnicity-specific and common-susceptibility loci among different ethnic groups.
Four T2D GWAS loci discovered in a Japanese population earlier have been shown to be significantly associated with T2D in the largest European GWAS meta-analysis10: KCNQ1 (refs 3, 4), UBE2E2 (ref. 5), C2CD4A–C2CD4B5 and ANK1 (ref. 6), highlighting that there are common loci conferring susceptibility to T2D among the different ethnic groups studied. Three additional loci (MIR129-LEP, GPSM1 and SLC16A11-SLC16A13) have been identified by a large-scale Japanese GWAS (n=∼25,000) based on the imputation of genotypes using the 1000 Genomes Project data as a reference7. One of the findings, the association in the SLC16A11–SLC16A13 was also confirmed in the Mexican GWAS study15.
To identify novel loci for susceptibility to T2D, we have expanded the Japanese GWAS data set by incorporating new Japanese GWAS data (9,817 T2D cases and 6,763 controls) with GWAS data in previously reported case–control individuals (5,646 T2D cases and 19,420 controls)7 followed by a validation study using independent Japanese case–control individuals (7,936 T2D and 5,539 controls) and multi-ethnic replication studies (East Asians: 12,554 T2D and 17,383 controls; Europeans: 38,947 T2D and 121,903 controls; South Asians: 10,587 T2D and 14,378 controls; and Mexian/Latinos: 3,848 T2D and 4,366 controls). As a result, we identify seven novel loci for T2D and the result indicates that expansion of single ethnic GWAS is still useful to identify novel susceptibility loci to complex traits.
Results
GWAS meta-analysis and validation in the Japanese population
Imputed genotype dosage data for 9,817 T2D cases and 6,763 controls for 7,521,072 autosomal SNPs (Stage-1, set-1) were obtained and combined with an independent GWAS data of previously reported case–control individuals7 (Stage-1, set-2: 5,646 T2D cases and 19,420 controls; 7,521,072 autosomal SNPs), as shown in Fig. 1a. There was no obvious inflation in the quantile–quantile plots for each study (Stage-1: λGC=1.13 and λGC adjusted for 1,000 cases and controls (λGC-1000)21=1.012; Stage-2: λGC=1.082 and λGC-1000=1.009), as shown in Supplementary Fig. 1A,B. SNPs with a low imputation quality (r2<0.7 either in set-1 or set-2) or with an inconsistent direction of effect between the studies were excluded from the analysis. We obtained 42 loci exhibiting a suggestive association with T2D (P<1 × 10−6). The most significant association in this meta-analysis was rs2237896 located at intron 15 of KCNQ1 (P=2.81 × 10−70), which was previously identified in Japanese GWAS3,4 (Supplementary Fig. 1C). Out of the 42 loci, 25 were previously established T2D susceptibility loci (Supplementary Table 1) and the remaining 17 were further evaluated using an independent Japanese case–control study (Stage-2: 7,936 T2D cases and 5,539 controls, multi-centre) and de novo genotyping (Supplementary Tables 2 and 3).
To explore the novel T2D susceptibility loci, we combined the association data for the candidate SNPs from all the Japanese case–control samples (Stage-1 set-1, Stage-1 set-2 and Stage-2) using a meta-analysis (Fig. 1b and Supplementary Table 3) and identified seven T2D susceptibility loci with a significant association (P<5 × 10−8; Table 1, Fig. 2 and Supplementary Table 3): rs1116357 near CCDC85A (P=6.97 × 10−10, odds ratio (OR)=1.09, 95% confidence interval (CI)=1.06–1.12), rs147538848 in FAM60A (P=7.83 × 10−10, OR=1.11, 95% CI=1.07–1.15), rs1575972 near DMRTA1 (P=1.50 × 10−9, OR=1.19, 95% CI=1.13–1.26), rs9309245 near ASB3 (P=1.25 × 10−8, OR=1.10, 95% CI=1.07–1.14), rs67156297 near ATP8B2 (P=1.95 × 10−8, OR=1.14, 95% CI=1.09–1.19), rs7107784 near MIR4686 (P=2.07 × 10−8, OR=1.14, 95% CI=1.09–1.20) and rs67839313 near INAFM2 (P=2.42 × 10−8, OR=1.09, 95% CI=1.06–1.12). The effect sizes (OR) of these seven SNPs were similar before and after adjusting for age, sex and body mass index (BMI) (Supplementary Table 4). The rs1575972 locus in DMRTA1 was located 170 kbp away from the T2D locus CDKN2A/B at 9p21 (refs 10, 22, 23, 24) and the rs7107784 locus near MIR4686 was located ∼620 kbp upstream of KCNQ1 (refs 3, 4). The linkage disequilibrium (LD) between rs1575972 and rs10811661, which was a lead SNP within the CDKN2A/B locus10, was weak (JPT: r2=0.01, CEU: r2=0.02). The association of rs1575972 was significant even after conditioning on rs10811661 (P for meta-analysis=2.45 × 10−9, OR=1.19, 95% CI=1.12–1.26; Supplementary Table 5); therefore, we considered the rs1575972 locus as a novel T2D susceptibility locus, independent of the CDKN2A/B locus. We also observed that the rs7107784 locus near MIR4686 was not in LD with rs2237897 (JPT: r2<0.01, CEU: r2<0.01), which was a lead SNP within the KCNQ1 locus3, and the association was similar after conditioning on rs2237897 (P for meta-analysis=2.75 × 10−8, OR=1.16, 95% CI=1.10–1.22; Supplementary Table 6).
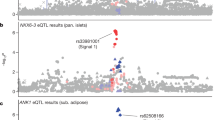
Each plot shows −log10 P-values against the chromosomal positions of SNPs in the specific region. The SNP with the strongest association signal (lead SNP) in each locus is represented as a purple diamond; the other SNPs are coloured according to the extent of LD with the lead SNP. Estimated recombination rates from the hg19/1000 Genomes Project March 2012 East Asian references are shown as light-blue lines. (a) CCDC85A, (b) FAM60A, (c) DMRTA1, (d) ASB3, (e) ATP8B2, (f) MIR4686 and (g) INAFM2. *Gene position for INAFM2 was added manually.
We also examined the association of these seven SNPs with glycaemic traits in Stage-2 control individuals, including fasting plasma glucose, homeostasis model assessment (HOMA) of β-cell function (HOMA-β) and HOMA of insulin resistance (IR). However, we did not detect any significant associations between the T2D risk alleles and these glycaemic traits (P≥0.0024 Supplementary Table 7). We also searched the publicly available European GWAS data11,25,26 (MAGIC, http://www.magicinvestigators.org) and found that the T2D risk allele at the DMRTA1 locus (rs11791293-C; proxy for rs1575972-T, CEU r2=1) and at the MIR4686 locus (rs7111341-T; proxy for rs7107784-G, CEU r2=0.95) were associated with a decrease in fasting plasma insulin (FPI) (P=0.0039; Supplementary Table 8) and with an increase in FPI (P=0.0066; Supplementary Table 8), respectively, although these associations were not statistically significant (P≥0.0008=0.05/63 (7 SNPs × 9 traits)).
Examination of seven novel loci in diverse ethnic groups
We analysed the association of these seven variants with disease susceptibility in populations other than Japanese. We obtained association data for four ethnic groups using de novo genotyping, in silico replication and by examining publicly available GWAS data10 including East Asian (n=up to 29,937: 12,554 T2Ds and 17,383 controls), South Asian (n=up to 24,965: 10,587 T2Ds and 14,378 controls), European (n=up to 160,850: 38,947 T2Ds and 121,903 controls) and Mexican (n=up to 8,214: 3,848 T2Ds and 4,366 controls) populations (Supplementary Table 9). Meta-analyses of the combined data from the four non-Japanese ethnicities indicated that four SNP loci, namely rs147538848 in FAM60A, rs1575972 near DMRTA1, rs7107784 near MIR4686 and rs67839313 near INAFM2 were associated with the disease after Bonferroni’s correction (P<0.00714=0.05/7; Table 2). The disease association of these four SNPs was further corroborated by combining the Japanese data with the multi-ethnic replication data sets (Supplementary Table 10). The rs67156297 locus in ATP8B2 was nominally associated with T2D in the combined meta-analysis for multi-ethnic groups other than the Japanese populations. We did not detect any disease association for the remaining two SNP loci other than in the Japanese population; however, the effect direction for each of the seven loci was consistent with that in the Japanese population.
Sex- and BMI-stratified analyses in the Japanese population
We performed BMI-stratified (BMI<25 or≥25) and sex-stratified analyses in the novel and established GWAS loci, to determine whether significant heterogeneity in allelic effects existed between non-obese and obese individuals or males and females in the Japanese population. BMI-stratified analysis for 83 previously established loci revealed evidence of significant heterogeneity in the effect size between non-obese and obese individuals at KCNQ1 (P for heterogeneity=8.89 × 10−5; Supplementary Table 11). The effect size of KCNQ1 was greater in the non-obese group than in the obese group (Supplementary Table 11). In sex-stratified analyses, individual established loci did not show significant heterogeneity in effect sizes between men and women (P>6 × 10−4; Supplementary Table 12).
For the seven novel T2D-associated loci identified in this study, no significant heterogeneity was detected in BMI-stratified or sex-stratified analyses (Supplementary Tables 13 and 14).
Fine mapping analyses for established T2D loci
We examined the association data of 83 previously identified T2D susceptibility loci in the Japanese GWAS meta-analysis data (Supplementary Data 1 and Supplementary Fig. 2). Variants at 19 loci were found associated with T2D at a genome-wide level of significance and additional 30 loci were determined to be significantly associated with T2D (P<6.02 × 10−4=0.05/83). Of the above 49 significant associations, ADCY5, HNF1A and PRC1 were not previously evaluated in the Japanese population, because the lead SNPs within these loci in the European GWAS (rs11708067 and rs11717195 at the ADCY5 locus9,10, rs12427353 and rs7957197 at the HNF1A locus8,10 and rs8042680 at the PRC1 locus8) were monoallelic in the Japanese population. In this study, rs79223353 at the ADCY5 locus, rs55783344 at the HNF1A locus and rs79548680 at the PRC1 locus were determined to be significantly associated with T2D (P<6.02 × 10−4; Supplementary Data 1 and Supplementary Fig. 3). Meta-analysis combining the GWAS data with de novo genotyping data for Stage-2 individuals revealed that the association of rs79223353 within the ADCY5 locus and rs79548680 within the PRC1 locus reached genome-wide significance in the Japanese population (Supplementary Table 15). We did not detect any disease-associated SNPs within the 16 loci (9 derived from European GWAS, 3 from East Asian, 2 from trans-ethnic, 1 from South Asian and 1 from African American, P≥0.05) using fine mapping analyses (Supplementary Data 1 and Supplementary Fig. 2). We also identified a secondary association signal located at EXOC6 near the IDE-HHEX locus10. The associations of rs78627331 and rs34773007 within the EXOC6 locus were significant after conditioning on rs1111875 (r2=0.01 for rs78627331 and r2=0.04 for rs34773007 in JPT), which was a previously reported lead SNP within the IDE-HHEX locus (P=1.49 × 10−8 for rs78627331, P=2.20 × 10−8 for rs34773007; Supplementary Table 16).
Drug targets search by a bioinformatics approach
We applied the genetic information from previously reported and the present GWAS, to investigate potential drug targets for the treatment of T2D. First, we defined 286 T2D potential risk genes located in any of the 90 T2D risk loci (7 novel T2D loci that were identified in the present study and 83 previously identified T2D loci; see Supplementary Note). Among the 286 genes, by using a previously described scoring system27, we selected 40 genes with a score of 2 or higher (Supplementary Note, Supplementary Tables 17–20, Supplementary Data 2 and 3, and Supplementary Figs 4 and 5) as ‘biological T2D risk genes’ (Fig. 3 and Supplementary Data 4). In brief, we scored each of the 286 biological candidate genes by adopting the following six selection criteria and calculating the number of satisfied criteria as follows: (1) genes for which T2D risk SNPs or any of the SNPs in LD (r2≥0.80) with them were annotated as missense variants; (2) genes for which cis-eQTL genes of any of lymphoblastoid cell lines, adipose tissue or liver tissues were observed for T2D risk SNPs (P<0.05 for lymphoblastoid cell lines and adipose tissues, and P<0.004 for liver tissues); (3) monogenic diabetes genes; (4) genes for which at least three out of six associated phenotype labels (homeostasis/metabolism, liver/biliary system, endocrine/exocrine gland, growth/size/body, mortality/ageing and embryogenesis; P<9.2 × 10−5) were observed in knockout mouse28; (5) genes prioritized by PubMed text mining genes using GRAIL29 with gene-based P<0.05; and (6) genes prioritized by protein–protein interaction (PPI) network using DAPPLE30 with gene based P<0.05.
The gene score for each gene was calculated by summing up the number of criteria satisfied (filled red box indicates criterion satisfied; 40 genes with a score ≥2 out of 286 genes included in the T2D risk loci were defined as ‘biological candidate genes’; see Supplementary Data 4). Filled blue boxes indicate the nearest gene to the T2D risk SNP. Filled green boxes indicate overlap with drug target genes.
As these criteria exhibited weak correlations with each other (r2<0.34; Supplementary Fig. 5), each gene was given a score based on the number of criteria that were met (scores ranged from 0 to 6). Genes with a score of 2 or higher were defined as biological T2D risk genes.
We searched for overlapping genes between the 871 drug target genes corresponding to approved, in clinical trials or experimental drugs for various human diseases described in the previous report27, and the 40 biological T2D risk genes plus 712 genes that are known to have products that have direct PPI30 with the biological T2D risk gene products. We identified a total of 83 overlapping genes (Supplementary Fig. 6 and Supplementary Table 21). Fourteen drug target genes with approved T2D treatments demonstrated significant overlap with the 40 biological T2D risk genes and 712 genes with direct PPI (4 genes overlapped with 5.6-fold enrichment as determined using permutation analysis, P=0.0042; Supplementary Table 22 and Supplementary Fig. 6). The 871 drug target genes had overlap with the identified 83 genes, which is 1.8-fold more enrichment than would be expected by chance, but this is 3.1-fold less enrichment compared with overlap of the targets of T2D drugs (Supplementary Fig. 6).
Of the 83 overlapping genes, 5 were biological T2D risk genes (PPARG, KCNJ11, ABCC8, GCK and KIF11; Fig. 4). Three of these are targets of approved T2D drug treatments: PPARG, thiazolidinediones; KCNJ11, sulfonylurea; ABCC8, sulfonylureas and glinide. GCK is a target gene of a GCK activator that was in clinical trials as of August 2014 (Supplementary Table 23). Of the remaining 78 genes, 2 genes exhibit PPI with 3 biological T2D risk gene products. GSK3B interacts with NOTCH1, NOTCH2 and CCND2, whereas JUN interacts with FBXW7, HHEX and CCND2. Eight genes interact with 2 biological T2D risk gene products and 68 genes interact with a single biological T2D risk gene product (Supplementary Table 21). A list of therapeutic drugs that are currently under clinical trials targeting GCK, KIF11, GSK3B and JUN is shown in Supplementary Table 23.
Representative connections between T2D risk SNPs (blue), T2D biological genes (green), drug target genes (purple) and targeted drugs are shown (see Supplementary Table 21). *Compounds under clinical trial; **see Supplementary Table 23.
Discussion
In this study, we performed a GWAS meta-analysis in the Japanese population followed by validation using an independent Japanese sample. Integration of the results for ∼55,000 Japanese individuals identified 7 novel loci associated with T2D that reached genome-wide significance. In a subsequent transethnic meta-analysis, four loci were confirmed and one locus was suggested as common susceptibility loci for T2D in populations other than the Japanese population.
GWAS have been extensively performed in diverse ethnic groups, including populations of European, East Asian, South Asian and Mexican decent1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18. To this point, the sample size of GWAS for European populations has grown to over 100,000 (ref. 10) and these studies have identified nearly 50 loci associated with T2D. GWAS on populations of non-European origin and transethnic GWAS meta-analysis have identified more than 30 loci associated with T2D, which were not detected in earlier European GWAS3,4,5,6,7,12,13,14,15,18. Among these, several loci have been shown to be associated with T2D in larger European populations10, which suggests that further expansion of GWAS for non-European populations could prove useful in identifying additional susceptibility loci associated with T2D.
Among the seven novel loci identified in this study, rs147538848 in FAM60A, rs1575972 near DMRTA1, rs7107784 near MIR4686 and rs67839313 near INAFM2 were shown to be common susceptibility loci for T2D across different ethnicities, although the significance of the association differed among individual ethnic groups for several loci.
rs147538848 is located in the intron of FAM60A, which encodes a subunit of the Sin3 deacetylase complex (Sin3/HDAC1) that has been shown to be important for the repression of genes encoding components of the transforming growth factor-β signalling pathway31. Studies using a rat intrauterine growth retardation model have suggested that the Sin3/HDAC1 complex may negatively regulate the expression level of pancreatic and duodenal homeobox 1 (PDX1), which is known as an important transcription factor for the development of pancreas and β-cell maturation32 via histone modification of its proximal promoter33. A T2D risk allele at the FAM60A locus might contribute to disease susceptibility by impairing the transcriptional regulation of genes that are important for glucose metabolism.
INAFM2 encodes InaF-motif containing 2 and has previously been known as Osteogenesis upregulated transcript 1 (OGU1) or long intergenic non-protein coding RNA 984 (LINC00984), which is a putative long non-coding RNA. Although the expression of OGU1 has been shown to be upregulated during osteogenesis34, the function of INAFM2 encoding protein is still unknown. Around rs67839313, there are two plausible genes for susceptibility to T2D: PLCB2 and DISP2. PLCB2 encodes phospholipase C isoform β-2 and phospholipase C is a known regulator of insulin secretion through hydrolysis of islet phosphoinositide pools35. Therefore, it is feasible that this locus is associated with impaired glucose-stimulated insulin secretion machinery. DISP2 encodes dispatched homologue 2, which is a cell surface marker on insulin-positive cells36. Although the functional role of this molecule in glucose homeostasis is not well understood, it is potentially involved in the maturation of pancreatic β-cells or it might have a role in already matured pancreatic β-cells.
The effect size for the T2D association of rs1575972 near DMRTA1 was similar among all populations in this study, except for the Mexican/Latino population (Table 2 and Supplementary Table 10). The risk allele of rs1575972 in the DMRTA1 locus was nominally correlated with a decrease in FPI (Supplementary Table 8), which suggests that this locus might contribute to T2D susceptibility through affecting insulin secretion in pancreatic β-cells. The DMRTA1 encodes doublesex and mab-3-related transcription factor-like family A1, which has been recently reported to be involved in neuronal development by regulating the Pax6-Neurog2 transcriptional cascade37. Although the relevance of DMRTA1 to pancreatic development has not been established, DMRTA1 might play a role in β-cell development, because Pax6 and Neurog3, other member of the neurogenin subfamily, are key transcriptional regulators of pancreatic endocrine cell differentiation38.
INS, IGF-2 and TH are located at approximately rs7107784 near the MIR4686 locus. IGF-2 plays a key role in embryonic growth and may also influence body weight in adulthood39, and TH (tyrosine hydroxylase) has been shown to play a role in β-cell development40. This locus is known to be associated with risk of type-1 diabetes (rs1004446-C, 45 kbp from rs7107784; r2=0.003, D′=0.275)41. The risk allele of rs7107784-G is nominally associated with the increase of FPI levels in the European population (MAGIC data; Supplementary Table 8) and an increase of HOMA-IR in our Japanese data set (Supplementary Table 7). This suggests that the effects of rs7107784-G are probably not mediated by an impairment of insulin production or secretion, but rather by an impairment of insulin sensitivity.
rs67156297 near ATP8B2 was nominally associated with T2D in the transethnic replication meta-analysis (Table 2 and Supplementary Table 10). ATP8B2 encodes a member of the P4 family of ATPases (type 4P-type ATPase), which are multispan transmembrane proteins that have been implicated in phospholipid translocation from the exoplasmic to the cytoplasmic membrane leaflet42. The role of ATP8B2 in the pathogenesis of T2D has not been established. However, another member of the P4 ATPase family, atp10a, has been shown to be important for the biogenesis and/or membrane-directed trafficking of Glut4 receptors, and loss-of-function of atp10a induces IR and obesity in mice42.
The remaining two loci, rs1116357 near CCDC85A and rs9309245 near ASB3, were not associated with T2D (P>0.05) in the replication meta-analysis for non-Japanese populations, which suggests that the effect of these loci might be specific to the Japanese population. As heterogeneity in effect sizes was observed for rs1116357 or rs9309245 between Japanese and other ethnic groups, including European, South Asian and Mexican (Supplementary Tables 25 and 27), two possibilities might exist for the two SNP loci: (1) the LD between the causal alleles and the Japanese lead SNPs are consistent across the populations, but the risk alleles have effects only in the Japanese, and (2) the causal alleles are in LD with these SNPs only in the Japanese. By a systematic evaluation for effect sizes and LDs within these loci, we did not identify any SNPs associated with T2D in European populations, which are in LD with our lead SNPs in the Japanese, whereas not in LD in European populations (Supplementary Fig. 8 and Supplementary Tables 26 and 27). Therefore, the causal allele in the two loci might have an effect only in Japanese populations; however, further evaluation is required to elucidate the precise mechanism how these loci contribute to T2D susceptibility in the Japanese.
While searching for potential drug targets for T2D using a systematic bioinformatics approach, 83 overlapping genes were identified from 752 genes (40 biological T2D GWAS genes and 712 genes that encode products in direct PPI with 40 biological T2D GWAS genes) and 871 drug target genes for various human diseases27. Of these, 5 were T2D GWAS genes: PPARG, KCNJ11, ABCC8, GCK and KIF11. PPARG, KCNJ11 and ABCC8 have approved T2D treatment options. In addition, a GCK activator is currently undergoing clinical trials for the treatment of T2D. KIF11, which encodes kinesin family member 11 (also known as EG5), has been shown to be involved in regulating cell mitosis and inhibitors targeting this gene product have been developed as chemotherapeutic agents in the treatment of cancer43. Although the role of KIF11 in the regulation of glucose metabolism has not been well established, a recent study reported that knockdown of KIF11 using small interfering RNA resulted in increased glycogenesis in human primary hepatocytes44. Thus, a KIF11 inhibitor might ameliorate glucose homeostasis by suppressing gluconeogenesis from the liver.
We identified two genes, GSK3B and JUN, which directly interact with multiple biological T2D susceptibility genes. GSK3B encodes glycogen synthase kinase 3β, which is a constitutively active multifunctional serine/threonine kinase and is involved in diverse physiological pathways, including metabolism, cell cycle regulation, gene expression, development, oncogenesis and neuroprotection45. Several studies using Gsk3b-modified mouse models have suggested that inhibition of GSK3B function may have beneficial effects on glucose metabolism through pancreatic β-cell preservation or enhancement of insulin-stimulated glycogen synthase regulation and glycogen deposition45,46,47. Currently, GSK3B inhibitors are under clinical trial for the treatment of cancers (Supplementary Table 23), but these compounds could also be potential treatments for T2D.
JUN encodes the proto-oncogene c-Jun and the role of c-Jun in the pathogenesis of T2D is not well understood. However, c-Jun has been shown to decrease the expression of the human insulin gene by repressing insulin promoter activity48. c-Jun is a transactivation component of the heterodimeric transcription factor AP-1 and activated through phosphorylation of serines 63 and 73 by Jun N-terminal kinase 2 (ref. 49), and inhibition of JNK has been shown to ameliorate glucose intolerance in a mouse model for T2D50. Currently, AP-1 inhibitor is under clinical trial for the treatment of rheumatoid arthritis (Supplementary Table 23) and might also be potential treatments for T2D.
Although these results suggest that these loci are potential therapeutic targets for treating T2D, the pipeline used to identify these genes has some limitations. As eQTL effects have often been observed for genes far from each locus, it is possible that some biological genes located outside of LD block in each locus were overlooked. In addition, the selection criteria for PubMed text-mining or knockout mouse studies were based on the known functions; therefore, T2D-associated genes whose functions have not been established may have been missed. The number of criteria that were met for individual genes were simply summed for scoring, although the relative impact of the six criteria used here on biological significance may not be equal. We used the previously described scoring method27, to prioritize genes in an objective manner; however, it would be worthwhile to refine the pipeline by modifying the selection criteria for genes in future studies. Finally, the potential therapeutic targets or treatments identified through the in silico pipeline have not yet been validated through an experimental approach. Furthermore, in vivo evaluation is essential to clarify the therapeutic effect of these potential T2D treatments.
In conclusion, we have identified seven novel T2D susceptibility loci using a large-scale Japanese GWAS meta-analysis. The T2D association for four of these was also observed in non-Japanese populations. In addition, we have proposed several new potential pharmacological targets for T2D treatment using a systematic bioinformatics approach. These results indicate that expansion of single ethnic GWAS is still useful to identify novel susceptibility loci to complex traits not only for ethnicity-specific but also for common loci across different ethnicities. Moreover, systematic approaches for integrating the findings of genetic, biological and pharmacological studies could be useful for developing new T2D treatments, although additional pipeline refinement would be required.
Methods
Subjects
Discovery stage (Stage-1). We selected T2D cases from individuals registered in BioBank Japan as having T2D (set-1 cases, n=9,817). Control groups consisted of individuals registered in BioBank Japan as not having T2D but with diseases other than T2D (cerebral aneurysm, oesophageal cancer, endometrial cancer, chronic pulmonary emphysema or glaucoma) or volunteers from the Osaka-Midosuji Rotary Club and Pharma SNP consortium (set-1 controls, n=6,763; Supplementary Table 24). We also used case and control individuals registered in the BioBank Japan that were previously analysed and reported (set-2 cases, n=5,646 and set-2 controls, n=19,420)7. There was no overlap in individuals in set-1 and set-2.
Validation analysis (Stage-2). We examined 7,936 T2D cases from the BioBank Japan that were not included in the discovery stage and from subjects with T2D, who visited outpatient clinics at The University of Tokyo, Juntendo University, National Center for Global Health and Medicine, Hiranuma Clinic, St Marianna University School of Medicine, The Hiroshima Atomic Bomb Casualty Council Health Management Center, Kawasaki Medical School, Toyama University Hospital or Shiga University of Medical Science. We also examined 5,539 controls from individuals that enrolled during an annual health check-up at six institutions: The Hiroshima Atomic Bomb Casualty Council Health Management Center, The National Center for Global Health and Medicine, Keio University, Hiranuma Clinic, St Marianna University School of Medicine and Toyama University Hospital. T2D was diagnosed according to World Health Organization criteria51. We excluded individuals who were positive for antibodies against glutamic acid decarboxylase and those with diabetes due to liver dysfunction, steroids and other drugs that might raise glucose levels, malignancy or a monogenic disorder known to cause diabetes.
Clinical characteristics of Stage-1and Stage-2 participants are shown in Supplementary Table 24. Genomic DNA was extracted from peripheral leukocytes using the standard procedure. All individuals provided written informed consent to participate in this study. The protocol of this study conformed to the provisions of the Declaration of Helsinki and was approved by the ethical committees at the RIKEN Yokohama Institute and all other institutions.
Genotyping and quality control in the discovery stage
Set-1 samples were genotyped using the Human Omni Express Exome Bead Chip. There were 535,686 autosomal SNPs that passed quality control, with a call rate ≥0.99, for Hardy–Weinberg equilibrium test P ≥1 × 10−6 in controls and minor allele frequency (MAF) ≥0.01. Set-2 samples were genotyped using the Illumina Human 610K SNP array. There were 480,426 autosomal SNPs that passed quality control and were used for further analysis. For sample quality control, we evaluated cryptic relatedness for each sample using an identity-by-state method and removed samples that exhibited second-degree or closer relatedness. We further performed principal component analysis to select individuals within the major Japanese (Hondo) cluster as reported previously5,6,7,52, and data for 16,580 individuals (9,817 T2D cases and 6,763 controls) in set-1 and 25,066 individuals (5,646 T2D cases and 19,420 controls) in set-2 were used in subsequent analyses. To evaluate the potential effect of population stratification, we used a quantile–quantile plot of the observed P-values (Supplementary Fig. 1A,B).
Imputation
We performed genotype imputation using MACH and Minimac53,54 with individuals from the 1000 Genomes Project (phased JPT, CHB and Han Chinese South data n=275, March 2012) as reference populations55. We selected SNPs with MAF ≥0.01 and a Minimac software quality score (r2)≥0.7. Individual genotype dosage data were used for association studies using mach2dat53,54.
Genotyping and quality control in the Stage-2 analysis
We genotyped 7,936 individuals with T2D and 5,539 controls using a multiplex PCR-Invader assay, as described previously3,5,6,7. Genotyping success rates <95% or concordance rates <99.9% were excluded from further evaluation.
Follow-up analyses
We obtained follow-up analysis data (n=up to 223,966: 65,936 T2Ds and 158,030 controls) from multiple cohorts or a publicly available database, as described below.
East Asian populations. We obtained genotype data for up to 29,937 individuals (12,554 T2Ds and 17,383 controls), de novo genotyping from 2 cohorts and in silico replication data from 9 cohorts (Supplementary Table 9).
South Asian populations. We obtained in silico genotype data for a total of up to 24,965 individuals (10,587 T2Ds and 14,378 controls) from 6 cohorts (Supplementary Table 9).
European populations. We obtained genotype data for up to 160,850 individuals (38,947 T2Ds and 121,903 controls), de novo genotyping data from the Danish case–control study and from a publicly available database (DIAGRAM3 http://diagram-consortium.org/downloads.html)10. The Danish case–control study consisted of individuals from the Inter99 cohort56, Health2006 cohort57, Vejle Biobank58, T2D cases from the Danish ADDITION screening cohort59 and a T2D case–control study obtained at Steno Diabetes Center (SDC). Two SNPs were genotyped by Illumina MetaboChip in 8,781 individuals from Inter99, Health2006 and SDC, whereas four SNPs were genotyped by LGC Genomics, UK, in individuals from Inter99, Vejle Biobank, ADDITION and SDC samples (Supplementary Table 9).
Mexican/Latino population. We obtained in silico genotype data for up to 8,214 individuals (3,848 T2Ds and 4,366 controls) from the SIGMA Type 2 Diabetes Consortium (Supplementary Table 9).
Ethnicity was self-reported by the enroled individuals. For each study, approval was obtained from the institutional review boards of the participating institutions and written informed consent was obtained from all participants. We excluded association data obtained by imputed genotyped data with a low quality of imputation (r2<0.7 or info <0.7). Details of the study samples are described in Supplementary Table 9.
Statistical analysis
The association between each SNP and T2D was assessed using the logistic regression test with an additive model with or without adjusting for age, sex and log-transformed BMI. We combined data from the each GWAS and our validation analyses using an inverse variance method and examined heterogeneity with a Cochran’s Q test using METAL60. Regional association plots were generated using LocusZoom61.
We also performed quantitative traits analysis for fasting plasma glucose, HOMA-β and HOMA-IR using multiple linear regression analysis in an additive association model with or without adjusting for age, sex and log-transformed BMI. The Japanese samples studied here show skewed distribution values for BMI, HOMA-IR and HOMA-β; therefore, we have analysed the quantitative traits using log-transformed BMI, HOMA-IR and HOMA-β.
Drug discovery
We performed a search for potential drug targets using genetic information of confirmed T2D susceptibility loci and publicly available bioinformatics tools29,30 and databases28,62,63,64,65 using a method that has been previously described by Okada et al.27 (Supplementary Note).
Additional information
Data availability: Summary statistics of 2 Japanese GWAS (study 1: 9,817 cases, 6,763 controls; study2: 5,646 cases, 19,420 controls) for directly genotyped data are available through a NBDC Human Database website (http://humandbs.biosciencedbc.jp/en/.
How to cite this article: Imamura, M. et al. Genome-wide association studies in the Japanese population identifies seven novel loci for type 2 diabetes. Nat. Commun. 7:10531 doi: 10.1038/ncomms10531 (2016).
References
McCarthy, M. I. Genomics, type 2 diabetes, and obesity. N. Engl. J. Med. 363, 2339–2350 (2010).
Imamura, M. et al. Genetics of type 2 diabetes: the GWAS era and future perspectives. Endocr. J. 58, 723–739 (2011).
Unoki, H. et al. SNPs in KCNQ1 are associated with susceptibility to type 2 diabetes in East Asian and European populations. Nat. Genet. 40, 1098–1102 (2008).
Yasuda, K. et al. Variants in KCNQ1 are associated with susceptibility to type 2 diabetes mellitus. Nat. Genet. 40, 1092–1097 (2008).
Yamauchi, T. et al. A genome-wide association study in the Japanese population identifies susceptibility loci for type 2 diabetes at UBE2E2 and C2CD4A-C2CD4B. Nat. Genet. 42, 864–868 (2010).
Imamura, M. et al. A single-nucleotide polymorphism in ANK1 is associated with susceptibility to type 2 diabetes in Japanese populations. Hum. Mol. Genet 21, 3042–3049 (2012).
Hara, K. et al. Genome-wide association study identifies three novel loci for type 2 diabetes. Hum. Mol. Genet. 23, 239–446 (2014).
Voight, B. F. et al. Twelve type 2 diabetes susceptibility loci identified through large-scale association analysis. Nat. Genet. 42, 579–589 (2010).
Dupuis, J. et al. New genetic loci implicated in fasting glucose homeostasis and their impact on type 2 diabetes risk. Nat. Genet. 42, 105–116 (2010).
Morris, A. P. et al. Large-scale association analysis provides insights into the genetic architecture and pathophysiology of type 2 diabetes. Nat. Genet. 44, 981–990 (2012).
Scott, R. A. et al. Large-scale association analyses identify new loci influencing glycemic traits and provide insight into the underlying biological pathways. Nat. Genet. 44, 991–1005 (2012).
Mahajan, A. et al. Genome-wide trans-ancestry meta-analysis provides insight into the genetic architecture of type 2 diabetes susceptibility. Nat. Genet. 46, 234–244 (2014).
Kooner, J. S. Genome-wide association study in individuals of South Asian ancestry identifies six new type 2 diabetes susceptibility loci. Nat. Genet. 43, 984–989 (2011).
Cho, Y. S. Meta-analysis of genome-wide association studies identifies eight new loci for type 2 diabetes in East Asians. Nat. Genet. 44, 67–72 (2011).
The SIGMA Type 2 Diabetes Consortium. Sequence variants in SLC16A11 are a common risk factor for type 2 diabetes in Mexico. Nature 506, 97–101 (2014).
Steinthorsdottir, V. et al. Identification of low-frequency and rare sequence variants associated with elevated or reduced risk of type 2 diabetes. Nat. Genet. 46, 294–298 (2014).
Moltke, I. et al. A common Greenlandic TBC1D4 variant confers muscle insulin resistance and type 2 diabetes. Nature 512, 190–193 (2014).
Saxena, R. et al. Genome-wide association study identifies a novel locus contributing to type 2 diabetes susceptibility in Sikhs of Punjabi origin from India. Diabetes 62, 1746–1755 (2013).
Stahl, E. A. et al. Bayesian inference analyses of the polygenic architecture of rheumatoid arthritis. Nat. Genet. 44, 483–489 (2012).
Chan, J. C. N. et al. Diabetes in Asia: epidemiology, risk factors, and pathophysiology. JAMA 301, 2129–2140 (2009).
Freedman, M. L. et al. Assessing the impact of population stratification on genetic association studies. Nat. Genet. 36, 388–393 (2004).
Saxena, R. et al. Genome-wide association analysis identifies loci for type 2 diabetes and triglyceride levels. Science 316, 1331–1336 (2007).
Zeggini, E. et al. Replication of genome-wide association signals in UK samples reveals risk loci for type 2 diabetes. Science 316, 1336–1341 (2007).
Scott, L. J. et al. A genome-wide association study of type 2 diabetes in Finns detects multiple susceptibility variants. Science 316, 1341–1345 (2007).
Manning, A. K. et al. A genome-wide approach accounting for body mass index identifies genetic variants influencing fasting glycemic traits and insulin resistance. Nat. Genet. 44, 659–669 (2012).
Soranzo, N. et al. Common variants at 10 genomic loci influence hemoglobin A1(C) levels via glycemic and nonglycemic pathways. Diabetes 59, 3229–3239 (2010).
Okada, Y. et al. Genetics of rheumatoid arthritis contributes to biology and drug discovery. Nature 20, 376–381 (2014).
Eppig, J. T. et al. The Mouse Genome Database (MGD): comprehensive resource for genetics and genomics of the laboratory mouse. Nucleic Acids Res. 40, D881–D886 (2012).
Raychaudhuri, S. et al. Identifying relationships among genomic disease regions: predicting genes at pathogenic SNP associations and rare deletions. PLoS Genet. 5, e1000534 (2009).
Rossin, E. J. et al. Proteins encoded in genomic regions associated with immune-mediated disease physically interact and suggest underlying biology. PLoS Genet. 7, e1001273 (2011).
Rouse, R. et al. Complex SIN3-HDAC1 histone deacetylase cycle-fluctuating regulator of the protein is a cell family with sequence similarity 60A gene regulation. J. Biol. Chem. 287, 32346–32353 (2012).
Holland, A. M. et al. Experimental control of pancreatic development and maintenance. Proc. Natl Acad. Sci. USA 99, 12236–12241 (2011).
Park, J. H. Development of type 2 diabetes following intrauterine growth retardation in rats is associated with progressive epigenetic silencing of Pdx1. J. Clin. Invest. 118, 2316–2324 (2008).
Kikuchi, K. et al. Transcripts of unknown function in multiple-signaling pathways involved in human stem cell differentiation. Nucleic Acids Res. 37, 4987–5000 (2009).
Walstar, S. et al. Regulation of insulin secretion by phospholipase C. Am. J. Physiol. 271, E409–E416 (1996).
Hald, J. et al. Pancreatic islet and progenitor cell surface markers with cell sorting potential. Diabetologia 55, 154–165 (2012).
Kikkawa, T. et al. Dmrta1 regulates proneural gene expression downstream of Pax6 in the mammalian telencephalon. Genes Cells 18, 636–649 (2013).
Wilson, M. E. et al. Gene expression cascades in pancreatic development. Mech. Dev. 120, 65–80 (2003).
O'Dell, S. D. et al. Molecules in focus Insulin-like growth factor II (IGF-II). Int. J. Biochem. Cell Biol. 30, 767–771 (1998).
Vázquez, P. et al. Non-neural tyrosine hydroxylase, via modulation of endocrine pancreatic precursors, is required for normal development of beta cells in the mouse pancreas. Diabetologia 57, 2339–2347 (2014).
Hakonarson, H. et al. A genome-wide association study identifies KIAA0350 as a type 1 diabetes gene. Nature 448, 591–594 (2007).
Flomer, D. E. P4 ATPases—Lipid flippases and their role in disease. Biochim. Biophys. Acta 1791, 628–635 (2009).
Salmela, A. L. et al. Mitosis as an anti-cancer drug target. Chromosoma 122, 431–449 (2013).
Haney, S. et al. RNAi screening in primary human hepatocytes of genes implicated in genome-wide association studies for roles in type 2 diabetes identifies roles for CAMK1D and CDKAL1, among others, in hepatic glucose regulation. PLoS ONE 8, e64946 (2013).
Rayasam, G. V. et al. Glycogen synthase kinase 3: more than a namesake. Br. J. Pharmacol. 156, 885–898 (2009).
Liu, Y. et al. Conditional ablation of Gsk-3β in islet beta cells results in expanded mass and resistance to fat feeding-induced diabetes in mice. Diabetologia 53, 2600–2610 (2010).
Patel, S. et al. Tissue-specific role of glycogen synthase kinase 3beta in glucose homeostasis and insulin action. Mol. Cell Biol. 28, 6314–6328 (2008).
Inagaki, N. et al. c-Jun represses the human insulin promoter activity that depends on multiple cAMP response elements. Proc. Natl Acad. Sci. USA 89, 1045–1049 (1992).
Meng, Q. H. et al. c-JUN, at the crossroad of the signaling network. Protein Cell 2, 889–898 (2011).
Bennett, B. L. et al. JNK: a new therapeutic target for diabetes. Curr. Opin. Pharmacol. 3, 420–425 (2003).
Alberti, K. G. et al. Definition, diagnosis and classification of diabetes mellitus and its complications. Part 1: diagnosis and classification of diabetes mellitus provisional report of a WHO consultation. Diabet. Med. 15, 539–553 (1998).
Yamaguchi-Kabata, Y. et al. Japanese population structure, based on SNP genotypes from 7003 individuals compared to other ethnic groups: effects on population-based association studies. Am. J. Hum. Genet. 83, 445–456 (2008).
Li, Y. et al. MaCH: using sequence and genotype data to estimate haplotypes and unobserved genotypes. Genet. Epidemiol. 34, 816–834 (2010).
Howie, B. et al. Fast and accurate genotype imputation in genome-wide association studies through pre-phasing. Nat. Genet. 44, 955–959 (2012).
1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Jørgensen, T. et al. A randomized non-pharmacological intervention study for prevention of ischaemic heart disease: baseline results Inter99(1). Eur. J. Cardiovasc. Prev. Rehab. 10, 377–386 (2003).
Thuesen, B. H. et al. Cohort Profile: the Health2006 cohort, Research Centre for Prevention and Health. Int. J. Epidemiol. 43, 568–575 (2014).
Albrechtsen, A. et al. Exome sequencing-driven discovery of coding polymorphisms associated with common metabolic phenotypes. Diabetologia 56, 298–310 (2013).
Lauritzen, T. et al. The ADDITION study: proposed trial of the cost-effectiveness of an intensive multifactorial intervention on morbidity and mortality among people with Type 2 diabetes detected by screening. Int. J. Obes. Relat. Metab. Disord. 24, S6–S11 (2000).
Cristen, J. W. et al. METAL: fast and efficient meta-analysis of genome wide association scan. Bioinformatics 26, 2190–2191 (2010).
Pruim, R. J. et al. LocusZoom: regional visualization of genome-wide association scan results. Bioinformatics 26, 2336–2337 (2010).
Grundberg, E. et al. Mapping cis- and trans-regulatory effects across multiple tissues in twins. Nat. Genet. 44, 1084–1089 (2012).
Schadt, E. E. et al. Mapping the genetic architecture of gene expression in human liver. PLoS Biol. 6, e107 (2008).
Wishart, D. S. et al. DrugBank: a comprehensive resource for in silico drug discovery and exploration. Nucleic Acids Res. 34, D668–D672 (2006).
Zhu, F. et al. Therapeutic target database update 2012: a resource for facilitating target-oriented drug discovery. Nucleic Acids Res. 40, D1128–D1136 (2012).
Acknowledgements
The Mexican/Latino association data were provided by SIGMA T2D Consortium (see Supplementary Note for the contributors list). Data on glycaemic traits in European populations have been contributed by MAGIC investigators and have been downloaded from www.magicinvestigators.org/. Data on GWAS meta-analysis for T2D in European populations have been contributed by DIAGRAM consortium and have been downloaded from http://diagram-consortium.org/. This work was partly supported by a grant from the Leading Project of Ministry of Education, Culture, Sports, Science and Technology-Japan. The work of the Shanghai Jiao Tong University was supported from grants from the National 973 Program (2011CB504001), 863 Program (2012AA02A509) and National Science Foundation of China (81322010). R.C.W.M. and J.C.N.C. acknowledge support from the Hong Kong Foundation for Research and Development in Diabetes, established under the auspices of the Chinese University of Hong Kong, the Innovation and Technology Fund (ITS/088/08 and ITS/487/09FP)), and the Research Grants Council Theme-based Research Scheme (T12–402/13-N). The work by the Shanghai Diabetes Genetic Study (SDGS) was supported in part by the US National Institutes of Health grants R37CA070867, R01CA124558, R01CA64277 and UL1 RR024975, the Department of Defense Idea Award BC050791, Vanderbilt Ingram professorship funds and the Allen Foundation Fund. We thank the dedicated investigators and staff members from research teams at Vanderbilt University, Shanghai Cancer Institute and the Shanghai Institute of Preventive Medicine, and especially the study participants for their contributions in the studies. This study was provided with data from the Korean Genome Analysis Project (4845-301), the Korean Genome and Epidemiology Study (4851-302) and Korea Biobank Project (4851-307, KBP-2013-11 and KBP-2014-68) that were supported by the Korea Center for Disease Control and Prevention, Republic of Korea. This research was supported by an intramural grant from the Korea National Institute of Health (2014-NI73001-00), Republic of Korea. This study was supported by a grant of the Korea Health Technology R&D Project, Ministry of Health and Welfare, Republic of Korea (HI14C0060). The Novo Nordisk Foundation Center for Basic Metabolic Research is an independent Research Center at the University of Copenhagen partially funded by an unrestricted donation from the Novo Nordisk Foundation (www.metabol.ku.dk). The Danish studies, Inter99 and Health2006, were partly funded by the Lundbeck Foundation and produced by The Lundbeck Foundation Centre for Applied Medical Genomics in Personalised Disease Prediction, Prevention and Care (LuCamp, www.lucamp.org). The Asian Indian Diabetic Heart Study/Sikh Diabetes Study (AIDHS/SDS) was supported by the National Institute of Health grants KO1TW006087 funded by the Fogarty International Center, R01DK082766 funded by National Institute of Diabetes and Digestive and Kidney Diseases, and a seed grant from University of Oklahoma Health Sciences Center, Oklahoma City, USA. We thank the research participants for their contribution and support for making this study possible. A.H.C. was supported by a fellowship from CONACyT-Mexico. J.M.M. was supported by Sara Borrell Fellowship from the Instituto Carlos III, grant SEV-2011-00067 of Severo Ochoa Program and EMBO short-term fellowship, EFSD/Lilly research fellowship and Beatriu de Pinós fellowship from the Agency for Management of University and Research Grants (AGAUR). SIGMA study was supported by the Slim Foundation. Y.S.C. acknowledges support from the National Research Foundation of Korea (NRF) grant funded by the Korea government (MEST) (2012R1A2A1A03006155). Field-work, genotyping and standard clinical chemistry assays in PROMIS were principally supported by grants awarded to the University of Cambridge from the British Heart Foundation, UK Medical Research Council, Wellcome Trust, EU Framework 6-funded Bloodomics Integrated Project, Pfizer, Novartis and Merck. J.D. acknowledges that this work was funded by the UK Medical Research Council (G0800270), British Heart Foundation (SP/09/002), UK National Institute for Health Research Cambridge Biomedical Research Centre, European Research Council (268834) and European Commission Framework Programme 7 (HEALTH-F2-2012-279233).
Author information
Authors and Affiliations
Contributions
S. Maeda and T.K. supervised the study. M. Imamura, T.Y., K.H., S. Maeda and T.K. conceived and designed the experiments. M. Imamura, T.Y., K.H., S. Maeda and T.K. wrote the manuscript. K.Y., M. Iwata, H.H., H.M., C.I., K.K., H.W., Y.T., K.T., R.K. and M.K. collected Japanese samples and generated genotype data. M.Imamura, A.T., J.H., N.S., M. Iwasaki, H.F. and K.S. performed statistical analyses for the Japanese GWAS meta-analysis. C.H., R.Z., W.J., J.L., X.-O.S., W. Zheng, R.C.W.M., J.C.N.C. and C.H.T.T. contributed in design, sample collection, GWAS assays or statistical analyses for the Chinese GWA studies. S. Moon, Y.J.K., B.-J.K., S.H.K., K.S.P. and Y.S.C. performed statistical analyses for the Korean GWAS data. N.G., T.J., M.E.J., D.R.W., I.B., C.C., T.H. and O.P. performed statistical analyses for the Danish study. E.S.T. and X.W. performed statistical analyses for the Singapore GWAS data. A.H.C., J.M.M. and J.F. performed statistical analyses for the Mexican/Latin GWAS data. W. Zhao, J.D., D. Saleheen, A.R., P.F., J.R.S., R.S. and D. Sanghera contributed study data sets and performed statistical analyses for the South Asian GWAS data. M. Imamura and Y.O. performed in silico analyses for drug discovery.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1-8, Supplementary Tables 1-27, Supplementary Note and Supplementary References (PDF 11743 kb)
Supplementary Data 1
Summary of GWAS meta-analysis (Stage-1: set-1 + set-2) for 83 established T2D susceptibility loci. (XLSX 49 kb)
Supplementary Data 2
Mouse knockout phenotype (XLSX 30 kb)
Supplementary Data 3
cis-eQTL analyses (XLSX 21 kb)
Supplementary Data 4
Biological scores of the genes in the T2D risk loci (XLSX 31 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Imamura, M., Takahashi, A., Yamauchi, T. et al. Genome-wide association studies in the Japanese population identify seven novel loci for type 2 diabetes. Nat Commun 7, 10531 (2016). https://doi.org/10.1038/ncomms10531
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms10531
This article is cited by
-
Glucocorticoid receptor-NECAB1 axis can negatively regulate insulin secretion in pancreatic β-cells
Scientific Reports (2023)
-
Transcriptomics-driven drug repositioning for the treatment of diabetic foot ulcer
Scientific Reports (2023)
-
The role of FTO variant rs1421085 in the relationship with obesity: a systematic review and meta-analysis
Eating and Weight Disorders - Studies on Anorexia, Bulimia and Obesity (2022)
-
Suppressive role of E3 ubiquitin ligase FBW7 in type I diabetes in non-obese diabetic mice through mediation of ubiquitination of EZH2
Cell Death Discovery (2021)
-
Polymorphic genetic markers and how they are associated with clinical and metabolic indicators of type 2 diabetes mellitus in the Kazakh population
Journal of Diabetes & Metabolic Disorders (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.