Abstract
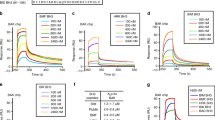
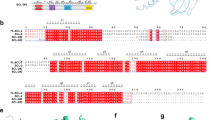
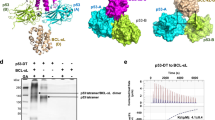
Following DNA damage, nuclear p53 induces the expression of PUMA, a BH3-only protein that binds and inhibits the antiapoptotic BCL-2 repertoire, including BCL-xL. PUMA, unique among BH3-only proteins, disrupts the interaction between cytosolic p53 and BCL-xL, allowing p53 to promote apoptosis via direct activation of the BCL-2 effector molecules BAX and BAK. Structural investigations using NMR spectroscopy and X-ray crystallography revealed that PUMA binding induced partial unfolding of two α-helices within BCL-xL. Wild-type PUMA or a PUMA mutant incapable of causing binding-induced unfolding of BCL-xL equivalently inhibited the antiapoptotic BCL-2 repertoire to sensitize for death receptor–activated apoptosis, but only wild-type PUMA promoted p53-dependent, DNA damage–induced apoptosis. Our data suggest that PUMA-induced partial unfolding of BCL-xL disrupts interactions between cytosolic p53 and BCL-xL, releasing the bound p53 to initiate apoptosis. We propose that regulated unfolding of BCL-xL provides a mechanism to promote PUMA-dependent signaling within the apoptotic pathways.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Green, D.R. & Kroemer, G. Cytoplasmic functions of the tumour suppressor p53. Nature 458, 1127–1130 (2009).
Chipuk, J.E., Bouchier-Hayes, L., Kuwana, T., Newmeyer, D.D. & Green, D.R. PUMA couples the nuclear and cytoplasmic proapoptotic function of p53. Science 309, 1732–1735 (2005).
Nakano, K. & Vousden, K.H. PUMA, a novel proapoptotic gene, is induced by p53. Mol. Cell 7, 683–694 (2001).
Yu, J., Zhang, L., Hwang, P.M., Kinzler, K.W. & Vogelstein, B. PUMA induces the rapid apoptosis of colorectal cancer cells. Mol. Cell 7, 673–682 (2001).
Jeffers, J.R. et al. Puma is an essential mediator of p53-dependent and -independent apoptotic pathways. Cancer Cell 4, 321–328 (2003).
Chen, L. et al. Differential targeting of prosurvival Bcl-2 proteins by their BH3-only ligands allows complementary apoptotic function. Mol. Cell 17, 393–403 (2005).
Kuwana, T. et al. BH3 domains of BH3-only proteins differentially regulate Bax-mediated mitochondrial membrane permeabilization both directly and indirectly. Mol. Cell 17, 525–535 (2005).
Chipuk, J.E. et al. Direct activation of Bax by p53 mediates mitochondrial membrane permeabilization and apoptosis. Science 303, 1010–1014 (2004).
Leu, J.I., Dumont, P., Hafey, M., Murphy, M.E. & George, D.L. Mitochondrial p53 activates Bak and causes disruption of a Bak–Mcl1 complex. Nat. Cell Biol. 6, 443–450 (2004).
Chipuk, J.E. et al. Mechanism of apoptosis induction by inhibition of the anti-apoptotic BCL-2 proteins. Proc. Natl. Acad. Sci. USA 105, 20327–20332 (2008).
Hinds, M.G. et al. Bim, Bad and Bmf: intrinsically unstructured BH3-only proteins that undergo a localized conformational change upon binding to prosurvival Bcl-2 targets. Cell Death Differ. 14, 128–136 (2007).
Sattler, M. et al. Structure of Bcl-xL–Bak peptide complex: recognition between regulators of apoptosis. Science 275, 983–986 (1997).
Petros, A.M. et al. Rationale for Bcl-xL/Bad peptide complex formation from structure, mutagenesis, and biophysical studies. Protein Sci. 9, 2528–2534 (2000).
Feng, W., Huang, S., Wu, H. & Zhang, M. Molecular basis of Bcl-xL′s target recognition versatility revealed by the structure of Bcl-xL in complex with the BH3 domain of Beclin-1. J. Mol. Biol. 372, 223–235 (2007).
Liu, X., Dai, S., Zhu, Y., Marrack, P. & Kappler, J.W. The structure of a Bcl-xL/Bim fragment complex: implications for Bim function. Immunity 19, 341–352 (2003).
Petros, A.M., Gunasekera, A., Xu, N., Olejniczak, E.T. & Fesik, S.W. Defining the p53 DNA-binding domain/Bcl-xL-binding interface using NMR. FEBS Lett. 559, 171–174 (2004).
Hagn, F. et al. BclxL changes conformation upon binding to wild-type but not mutant p53 DNA binding domain. J. Biol. Chem. 285, 3439–3450 (2010).
Muchmore, S.W. et al. X-ray and NMR structure of human Bcl-xL, an inhibitor of programmed cell death. Nature 381, 335–341 (1996).
Berjanskii, M. & Wishart, D.S. NMR: prediction of protein flexibility. Nat. Protoc. 1, 683–688 (2006).
O′Neill, J.W., Manion, M.K., Maguire, B. & Hockenbery, D.M. BCL-XL dimerization by three-dimensional domain swapping. J. Mol. Biol. 356, 367–381 (2006).
Denisov, A.Y., Sprules, T., Fraser, J., Kozlov, G. & Gehring, K. Heat-induced dimerization of BCL-xL through α-helix swapping. Biochemistry 46, 734–740 (2007).
Day, C.L. et al. Structure of the BH3 domains from the p53-inducible BH3-only proteins Noxa and Puma in complex with Mcl-1. J. Mol. Biol. 380, 958–971 (2008).
Smits, C., Czabotar, P.E., Hinds, M.G. & Day, C.L. Structural plasticity underpins promiscuous binding of the prosurvival protein A1. Structure 16, 818–829 (2008).
Nikolova, P.V., Henckel, J., Lane, D.P. & Fersht, A.R. Semirational design of active tumor suppressor p53 DNA binding domain with enhanced stability. Proc. Natl. Acad. Sci. USA 95, 14675–14680 (1998).
Joerger, A.C., Allen, M.D. & Fersht, A.R. Crystal structure of a superstable mutant of human p53 core domain. Insights into the mechanism of rescuing oncogenic mutations. J. Biol. Chem. 279, 1291–1296 (2004).
Xu, H. et al. The MDM2-binding region in the transactivation domain of p53 also acts as a Bcl-X-L–binding motif. Biochemistry 48, 12159–12168 (2009).
Letai, A. et al. Distinct BH3 domains either sensitize or activate mitochondrial apoptosis, serving as prototype cancer therapeutics. Cancer Cell 2, 183–192 (2002).
Kim, H. et al. Stepwise activation of BAX and BAK by tBID, BIM, and PUMA initiates mitochondrial apoptosis. Mol. Cell 36, 487–499 (2009).
Ren, D. et al. BID, BIM, and PUMA are essential for activation of the BAX- and BAK-dependent cell death program. Science 330, 1390–1393 (2010).
Miyashita, O., Onuchic, J.N. & Wolynes, P.G. Nonlinear elasticity, proteinquakes, and the energy landscapes of functional transitions in proteins. Proc. Natl. Acad. Sci. USA 100, 12570–12575 (2003).
Swain, J.F. & Gierasch, L.M. The changing landscape of protein allostery. Curr. Opin. Struct. Biol. 16, 102–108 (2006).
Frederick, K.K., Marlow, M.S., Valentine, K.G. & Wand, A.J. Conformational entropy in molecular recognition by proteins. Nature 448, 325–329 (2007).
Henzler-Wildman, K. & Kern, D. Dynamic personalities of proteins. Nature 450, 964–972 (2007).
Tzeng, S.R. & Kalodimos, C.G. Dynamic activation of an allosteric regulatory protein. Nature 462, 368–372 (2009).
Boehr, D.D., McElheny, D., Dyson, H.J. & Wright, P.E. The dynamic energy landscape of dihydrofolate reductase catalysis. Science 313, 1638–1642 (2006).
Masterson, L.R. et al. Dynamically committed, uncommitted, and quenched states encoded in protein kinase A revealed by NMR spectroscopy. Proc. Natl. Acad. Sci. USA 108, 6969–6974 (2011).
Smock, R.G. & Gierasch, L.M. Sending signals dynamically. Science 324, 198–203 (2009).
Wang, Y., Filippov, I., Richter, C., Luo, R. & Kriwacki, R.W. Solution NMR studies of an intrinsically unstructured protein within a dilute, 75 kDa eukaryotic protein assembly; probing the practical limits for efficiently assigning polypeptide backbone resonances. ChemBioChem 6, 2242–2246 (2005).
Laue, T.M., Shah, B.D., Ridgeway, T.M. & Pelletier, S.L. Computer-aided interpretation of analytical sedimentation data for proteins in Analytical Ultracentrifugation in Biochemistry and Polymer Science (eds. S.E. Harding, A.J. Rowe & J.C. Horton) 90–125 (The Royal Society of Chemistry, 1992).
Schuck, P. Size-distribution analysis of macromolecules by sedimentation velocity ultracentrifugation and lamm equation modeling. Biophys. J. 78, 1606–1619 (2000).
Schuck, P., Perugini, M.A., Gonzales, N.R., Howlett, G.J. & Schubert, D. Size-distribution analysis of proteins by analytical ultracentrifugation: strategies and application to model systems. Biophys. J. 82, 1096–1111 (2002).
Balbo, A., Brown, P.H., Braswell, E.H. & Schuck, P. Measuring protein-protein interactions by equilibrium sedimentation. Curr. Prot. Immunol. 18, 18.8 (2007).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Keller, R.J.J. Optimizing the Process of Nuclear Magnetic Resonance Spectrum Analysis and Computer Aided Resonance Assignment. PhD thesis, Eidgenossische Technishe Hochschule (2005).
Religa, T.L., Ruschak, A.M., Rosenzweig, R. & Kay, L.E. Site-directed methyl group labeling as an NMR probe of structure and dynamics in supramolecular protein systems: applications to the proteasome and to the ClpP protease. J. Am. Chem. Soc. 133, 9063–9068 (2011).
Tugarinov, V. & Kay, L.E. Ile, Leu, and Val methyl assignments of the 723-residue malate synthase G using a new labeling strategy and novel NMR methods. J. Am. Chem. Soc. 125, 13868–13878 (2003).
Tugarinov, V. & Kay, L.E. An isotope labeling strategy for methyl TROSY spectroscopy. J. Biomol. NMR 28, 165–172 (2004).
Tugarinov, V., Choy, W.Y., Orekhov, V.Y. & Kay, L.E. Solution NMR-derived global fold of a monomeric 82-kDa enzyme. Proc. Natl. Acad. Sci. USA 102, 622–627 (2005).
Güntert, P. Automated NMR structure calculation with CYANA. Methods Mol. Biol. 278, 353–378 (2004).
Cornilescu, G., Delaglio, F. & Bax, A. Protein backbone angle restraints from searching a database for chemical shift and sequence homology. J. Biomol. NMR 13, 289–302 (1999).
Pettersen, E.F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Laskowski, R.A., Rullmannn, J.A., MacArthur, M.W., Kaptein, R. & Thornton, J.M. AQUA and PROCHECK-NMR: programs for checking the quality of protein structures solved by NMR. J. Biomol. NMR 8, 477–486 (1996).
Kiefer, F., Arnold, K., Kunzli, M., Bordoli, L. & Schwede, T. The SWISS-MODEL Repository and associated resources. Nucleic Acids Res. 37, D387–D392 (2009).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Zwart, P.H. et al. Automated structure solution with the PHENIX suite. Methods Mol. Biol. 426, 419–435 (2008).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Acknowledgements
We thank J.T. Opferman and S.W.G. Tait (St. Jude Children's Research Hospital) for the Mx-Cre bak−/− baxf/− animals and MCF7 SMAC-GFP cells, respectively. The prokaryotic expression vectors for PUMAβ and BCL-xLΔLΔC were kindly provided by E. Eldering (Academic Medical Center, Amsterdam) and G. Wagner (Harvard University), respectively. We would like to acknowledge D. Miller (St. Jude Children's Research Hospital) for help with synchrotron data collection. Southeast Regional Collaborative Access Team (SER-CAT) supporting institutions may be found at http://www.ser.aps.anl.gov. Use of the SER-CAT Advanced Photon Source was supported by the US Department of Energy, Office of Science, Office of Basic Energy Sciences, under contract no. W-31-109-Eng-38. We would like to thank Bruker BioSpin and R. Weismann and W. Bermel for access to a 950-MHz NMR spectrometer. We also thank M. Madan Babu and A. Venkatakrishnan (Medical Research Council Laboratory of Molecular Biology, Cambridge, UK) for stimulating discussions and comments on the manuscript. This work was supported by NIH R01CA082491 and 1R01GM083159 (to R.W.K.), NIH R01GM52735 and R01GM96208 (to D.R.G.), NIH R01 CA157740 (to J.E.C.), the JJR Foundation (to J.E.C.), the William A. Spivak Fund (to J.E.C.), the Fridolin Charitable Trust (to J.E.C.), a National Cancer Institute Cancer Center Support Grant P30CA21765 (at St. Jude Children's Research Hospital), research grant no. 5-FY11-74 from the March of Dimes Foundation (to J.E.C.) and the American Lebanese Syrian Associated Charities. J.C.F. is a recipient of the Alma and Hal Reagan Cancer Research fellowship provided by the University of Tennessee Health Sciences Center.
Author information
Authors and Affiliations
Contributions
A.V.F., J.E.C. and J.C.F. as well as D.R.G. and R.W.K. contributed equally to this work. A.V.F. performed experiments, analyzed data and wrote the paper; J.E.C. performed experiments, analyzed data and wrote the paper; J.C.F. performed experiments, analyzed data and wrote the paper; M.-K.Y., C.R.G., A.N., K.B., L.O. and L.M. performed experiments and analyzed data; and S.W.W., D.R.G. and R.W.K. analyzed data and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results (PDF 3113 kb)
Rights and permissions
About this article
Cite this article
Follis, A., Chipuk, J., Fisher, J. et al. PUMA binding induces partial unfolding within BCL-xL to disrupt p53 binding and promote apoptosis. Nat Chem Biol 9, 163–168 (2013). https://doi.org/10.1038/nchembio.1166
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1166
This article is cited by
-
Structures of p53/BCL-2 complex suggest a mechanism for p53 to antagonize BCL-2 activity
Nature Communications (2023)
-
Tolfenamic acid inhibits ROS-generating oxidase Nox1-regulated p53 activity in intrastriatal injection of malonic acid rats
The Journal of Physiological Sciences (2022)
-
A systematic review on the role of MSC-derived exosomal miRNAs in the treatment of heart failure
Molecular Biology Reports (2022)
-
The role of P53 up-regulated modulator of apoptosis (PUMA) in ovarian development, cardiovascular and neurodegenerative diseases
Apoptosis (2021)
-
EZH2 inhibitors reverse resistance to gefitinib in primary EGFR wild-type lung cancer cells
BMC Cancer (2020)