Abstract
The polarity protein Scribble (SCRIB) regulates apical–basal polarity, directional migration and tumour suppression in Drosophila and mammals1,2,3,4,5,6. Here we report that SCRIB is an important regulator of myeloid cell functions including bacterial infection and inflammation. SCRIB interacts directly with the NADPH oxidase (NOX) complex in a PSD95/Dlg/ZO-1 (PDZ)-domain-dependent manner and is required for NOX-induced reactive oxygen species (ROS) generation in culture and in vivo. On bacterial infection, SCRIB localized to phagosomes in a leucine-rich repeat-dependent manner and promoted ROS production within phagosomes to kill bacteria. Unexpectedly, SCRIB loss promoted M1 macrophage polarization and inflammation. Thus, SCRIB uncouples ROS-dependent bacterial killing activity from M1 polarization and inflammatory functions of macrophages. Modulating the SCRIB–NOX pathway can therefore identify ways to manage infection and inflammation with implications for chronic inflammatory diseases, sepsis and cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Bilder, D., Li, M. & Perrimon, N. Cooperative regulation of cell polarity and growth by Drosophila tumor suppressors. Science 289, 113–116 (2000).
Cordenonsi, M. et al. The Hippo transducer TAZ confers cancer stem cell-related traits on breast cancer cells. Cell 147, 759–772 (2011).
Feigin, M. E. et al. Mislocalization of the cell polarity protein scribble promotes mammary tumorigenesis and is associated with basal breast cancer. Cancer Res. 74, 3180–3194 (2014).
Zhan, L. et al. Deregulation of scribble promotes mammary tumorigenesis and reveals a role for cell polarity in carcinoma. Cell 135, 865–878 (2008).
Pearson, H. B. et al. SCRIB expression is deregulated in human prostate cancer, and its deficiency in mice promotes prostate neoplasia. J. Clin. Invest. 121, 4257–4267 (2011).
Elsum, I. A. et al. Scrib heterozygosity predisposes to lung cancer and cooperates with KRas hyperactivation to accelerate lung cancer progression in vivo. Oncogene 33, 5523–5533 (2014).
Baker, L. et al. SCRIBBLE is required for pregnancy-induced alveologenesis in the adult mammary gland. J. Cell Sci. 129, 2307–2315 (2016).
Meng, T. C., Fukada, T. & Tonks, N. K. Reversible oxidation and inactivation of protein tyrosine phosphatases in vivo. Mol. Cell 9, 387–399 (2002).
Nishinaka, Y. et al. A new sensitive chemiluminescence probe, L-012, for measuring the production of superoxide anion by cells. Biochem. Biophys. Res. Commun. 193, 554–559 (1993).
Babior, B. M., Kipnes, R. S. & Curnutte, J. T. Biological defense mechanisms. The production by leukocytes of superoxide, a potential bactericidal agent. J. Clin. Invest. 52, 741–744 (1973).
Park, H. S. et al. Cutting edge: direct interaction of TLR4 with NAD(P)H oxidase 4 isozyme is essential for lipopolysaccharide-induced production of reactive oxygen species and activation of NF-κB. J. Immunol. 173, 3589–3593 (2004).
Segal, A. W. & Abo, A. The biochemical basis of the NADPH oxidase of phagocytes. Trends Biochem. Sci. 18, 43–47 (1993).
Knaus, U. G., Heyworth, P. G., Evans, T., Curnutte, J. T. & Bokoch, G. M. Regulation of phagocyte oxygen radical production by the GTP-binding protein Rac 2. Science 254, 1512–1515 (1991).
Nourry, C., Grant, S. G. & Borg, J. P. PDZ domain proteins: plug and play! Sci. STKE 2003, re7 (2003).
Bedard, K. & Krause, K. H. The NOX family of ROS-generating NADPH oxidases: physiology and pathophysiology. Physiol. Rev. 87, 245–313 (2007).
Groemping, Y., Lapouge, K., Smerdon, S. J. & Rittinger, K. Molecular basis of phosphorylation-induced activation of the NADPH oxidase. Cell 113, 343–355 (2003).
Lee, H. J. & Zheng, J. J. PDZ domains and their binding partners: structure, specificity, and modification. Cell Commun. Signal. 8, 8 (2010).
Birrane, G., Chung, J. & Ladias, J. A. Novel mode of ligand recognition by the Erbin PDZ domain. J. Biol. Chem. 278, 1399–1402 (2003).
Segal, A. W. How neutrophils kill microbes. Annu. Rev. Immunol. 23, 197–223 (2005).
Bilder, D. & Perrimon, N. Localization of apical epithelial determinants by the basolateral PDZ protein Scribble. Nature 403, 676–680 (2000).
Audebert, S. et al. Mammalian Scribble forms a tight complex with the βPIX exchange factor. Curr. Biol. 14, 987–995 (2004).
Lam, G. Y. et al. Listeriolysin O suppresses phospholipase C-mediated activation of the microbicidal NADPH oxidase to promote Listeria monocytogenes infection. Cell Host Microbe 10, 627–634 (2011).
Briggs, R. T., Drath, D. B., Karnovsky, M. L. & Karnovsky, M. J. Localization of NADH oxidase on the surface of human polymorphonuclear leukocytes by a new cytochemical method. J. Cell Biol. 67, 566–586 (1975).
Benoit, M., Desnues, B. & Mege, J. L. Macrophage polarization in bacterial infections. J. Immunol. 181, 3733–3739 (2008).
Lawrence, T. The nuclear factor NF-κB pathway in inflammation. Cold Spring Harb. Perspect. Biol. 1, a001651 (2009).
Matthews, J. R., Wakasugi, N., Virelizier, J. L., Yodoi, J. & Hay, R. T. Thioredoxin regulates the DNA binding activity of NF-κB by reduction of a disulphide bond involving cysteine 62. Nucleic Acids Res. 20, 3821–3830 (1992).
Sattler, M. et al. Hematopoietic growth factors signal through the formation of reactive oxygen species. Blood 93, 2928–2935 (1999).
Sheng, K. C., Pietersz, G. A., Tang, C. K., Ramsland, P. A. & Apostolopoulos, V. Reactive oxygen species level defines two functionally distinctive stages of inflammatory dendritic cell development from mouse bone marrow. J. Immunol. 184, 2863–2872 (2010).
Dinauer, M. C. & Orkin, S. H. Chronic granulomatous disease. Annu. Rev. Med. 43, 117–124 (1992).
Rieber, N., Hector, A., Kuijpers, T., Roos, D. & Hartl, D. Current concepts of hyperinflammation in chronic granulomatous disease. Clin. Dev. Immunol. 2012, 252460 (2012).
Winkelstein, J. A. et al. Chronic granulomatous disease. Report on a national registry of 368 patients. Medicine 79, 155–169 (2000).
Han, W. et al. NADPH oxidase limits lipopolysaccharide-induced lung inflammation and injury in mice through reduction-oxidation regulation of NF-κB activity. J. Immunol. 190, 4786–4794 (2013).
Wilson, C., Nunez, M. T. & Gonzalez-Billault, C. Contribution of NADPH oxidase to the establishment of hippocampal neuronal polarity in culture. J. Cell Sci. 128, 2989–2995 (2015).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Lutz, M. B. et al. An advanced culture method for generating large quantities of highly pure dendritic cells from mouse bone marrow. J. Immunol. Methods 223, 77–92 (1999).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Johnson, B. A. & Blevins, R. A. NMR View: A computer program for the visualization and analysis of NMR data. J. Biomol. NMR 4, 603–614 (1994).
Tsuchiya, S. et al. Induction of maturation in cultured human monocytic leukemia cells by a phorbol diester. Cancer Res. 42, 1530–1536 (1982).
Acknowledgements
We would like to thank S. Courtneidge (Oregon Health and Science University, USA) for Tks5, NOXA1 and NoxO1 cDNA, I. Harris for assistance with ROS assays, G. M. C. Gasmi-Seabrook for assistance with NMR experiments, P. B. Stathopulos for assistance in structural modelling and its analysis, and members of the Muthuswamy laboratory for critical discussions. This work was supported by CA098830, BC075024, Era of Hope Scholar award from DOD Breast Cancer Research Program; Rita Allen Foundation, Lee K Margaret Lau Chair for breast cancer research and Campbell Family Institute for Breast cancer research to S.K.M. M.U. received fellowships from The Uehara Memorial Foundation, The Naito Foundation, and the Terry Fox Foundation EIRR21 at CIHR, and a JSPS postdoctoral fellowship for research abroad. This was also funded in part by the Ontario Ministry of Health and Long Term Care. The views expressed do not necessarily reflect those of the OMOHLTC.
Author information
Authors and Affiliations
Contributions
W.Z. was responsible for the study, performed the majority of the experiments, analysed data and assisted with writing of the manuscript. M.U., N.I. and M.I. planned, performed, analysed and wrote the NMR studies. I.J. performed image acquisition and analysis and worked together with W.Z. and assisted with writing the manuscript. K.A. and M.B. were involved in maintaining the mouse colony. C.W.T. and P.S.O. planned, performed and analysed dendritic cell studies. S.K.M. conceptualized and executed the study and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 SCRIB is required for reactive oxygen species (ROS) generation.
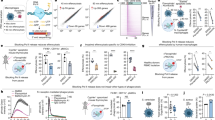
(a) Schematic diagram of the tetracycline-inducible knock-in constructs used to generate SCRIB shRNA mice. (b) Bone marrow derived macrophages were imaged with confocal microscope. Representative images are shown. Scale bar represents 50 μm.. Bone marrow cells were isolated from WT + Dox or − Dox, ishSCRIB + Dox or − Dox mice, and then neutrophils were isolated. Cells were then stained for flow cytometry and analyzed by flow cytometry checking the purity of neutrophils (c), or the expression of ishSCRIB (GFP) (d). (e) Immunoblot analysis of PDGF (50 ng ml−1) induced PDGFR-β tyrosine phosphorylation in MEFs at indicated time points. Immune cells isolated from bone marrow, spleen, thymus from WT and ishSCRIB mice (8 to 12 weeks old) with normal food or Dox food were analyzed GFP signal (f), Splenic cells stained with indicated surface marker for dendritic cells, neutrophils, and monocytes (g); bone marrow cells stained with surface marker for neutrophils, and monocytes (h); splenic cells and thymic cells were stained with T cell surface maker (i); bone marrow cells stained with B cell surface marker (j); and splenic cells stained with B cell surface maker (k).
Supplementary Figure 2 SCRIB is a component of the NOX complex.
(a) Quantitation of SCRIB binding to β-PIX. (b) Wild type (WT), Flagp22phox (Flag-p22) or Flag-p22phox without last four amino acid (Flag-p22Δ) were co-transfected with T7 tagged full length SCRIB. Flag immunoprecipitants were recovered and analyzed by anti-Flag and anti-T7 immunoblots. n = 3 biologically independent experiments (mean ± SD). (two-tailed, unpaired t-test with equal variances: P = 0.001). (c) Components of the four NOX complexes, NOX1-4. (d) Total cell lysates (TCL) from 293T cells transfected with Myc-tagged NOX complex proteins or anti-Myc immunoprecipitates were analyzed for the presence of endogenous SCRIB. (e) Total cell lysates (TCL) from 293T cells transfected with Myc-tagged NOX complex proteins or anti-Myc immunoprecipitates were analyzed for the presence of Myc-tagged proteins. (f) SCRIB immunoprecipitation followed by anti p22phox immunoblots using lysates obtained from parental RAW 264.7 cell line. Quantitation of p22phox binding to SCRIB in RAW 264.7 cell line (g) or in bone marrow derived macrophages (h) stimulated with PMA. Each experiment was performed in triplicate and data presented is an average of n = 3 independent experiments (mean ± SD). Unprocessed scans of full blots/gels are shown in Supplementary Fig. 6.
Supplementary Figure 3 SCRIB interacts directly with p22phox, which is required for PMA-induced generation of ROS.
(a) Schematic showing the full length SCRIB protein with Leucine Rich Repeats (LRR) and PDZ domains and the 6xHIS-tagged individual SCRIB PDZ domains or GST-tagged p22phox C-terminal tail (GST-p22tail). (b) 1H-15N heteronuclear single quantum coherence (HSQC) spectrum of 15N- labeled ScrPDZ4 (0.2 mM) in the absence (black) and the presence of varying concentration ratio of p22phox tail peptide/SCRIBPDZ4: 1.5 (red), 3.0 (green), 4.5 (blue), 8.9 (yellow), 17.8 (magenta) and 26.7-fold (cyan), are shown. Assignments were transferred from human SCRIB PDZ4 domain (BMRB number 10103) and further confirmed by 3D NMR experiments. Residues exhibiting large chemical shift (> 0.2 normalized chemical shift change in Supplementary Fig. 3c) are indicated. An asterisk (∗) indicates the peak from N-Acetylglycine-15N. Repeated 4 times and the representative experiment was shown as a figure. (c) NMR characterization of the interaction between p22phox tail and His-SCRIBPDZ4. The chemical shift change of each SCRIBPDZ4 residue is normalized to the observed change for G1112 in the presence of the p22phox tail peptide (26.7-fold). Blank regions indicate residues for which the value cannot be determined because of overlapped or unassigned peaks. Residues exhibiting large chemical shifts (> 0.2 normalized chemical shift change) are labeled. (d) SCRIBPDZ4 residues affected by the addition of p22phox tail peptide are mapped and indicated on the apo SCRIBPDZ4 structure (that is residues 8–105 from PDB ID 1UJU). (e) Electrostatic potential surface representation of SCRIBPDZ4 at neutral pH. Basic and acidic potentials are shown in blue (+5 kT/e) and red (− 5 kT/e), respectively. Basic residues near p22phox binding pocket are indicated. The electrostatic surface potential was generated using the APBS tools in Pymol, with partial charges determined by the PDB2PQR server (http://nbcr-222.ucsd.edu/pdb2pqr_1.8). (f) Conservation of SCRIBPDZ4 residues. Percent conservation of each residue position in human SCRIBPDZ4 (Residues within R1099-C1189) compared to the alignment of mouse, rat, zebrafish and fly SCRIB PDZ4 protein sequences. The residues affected by p22phox tail binding are shown. (g) 1H-15N HSQC spectrum of 15N-labeled ScrPDZ4 (0.2 mM) in the absence (black) and the presence (blue) of the full p22phox cytoplasmic region (that is residues 131–195, 5.0-fold excess). Residues exhibiting large chemical shift in Supplementary Fig. 2 are indicated. Repeated twice and the representative experiment was shown as a figure. (h) 1H-15N HSQC spectrum of 15N-labeled SCRIBPDZ4 (0.2 mM) in the absence (black) and the presence (green) of the p22phox cytoplasmic region with a C-terminal tail truncation (that is residues 131–185, 5.0- fold excess). An asterisk (∗) indicates peaks from N-Acetylglicine-15N. Repeated twice and the representative experiment was shown as a figure. HSQC spectra of wild-type i and G1112A/R1178A double mutant (j) 15N-labeled ScrPDZ4 (0.1 mM) in the absence (black) and the presence (red) of p22phox tail peptide (32.4-fold) are shown. Repeated twice and the representative experiment was shown as a figure. k CD spectra of the wild type (WT) and G1112A/R1178A double mutant forms of Scrib-PDZ4. (l) Comparison of response curves of WT and G1112A/R1178A variants of ScribPDZ4. Response curve was obtained with 20 μM ScribPDZ4 proteins.
Supplementary Figure 4 SCRIB required for clearing S. aureus infection in vivo and is recruited to phagosomes to mediate production of ROS.
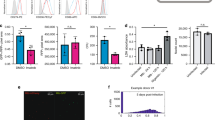
(a) Immunoblot analysis for SCRIB and GFP expression in whole blood from WT, ishSCRIB mice. (b) Whole blood from WT, ishSCRIB + Dox or − Dox mice was incubated with S. aureus for 20 min. Lysotaphin was added at 20 min of incubation and aliquots were removed at 60 min for enumeration of S. aureus CFU. Blood from n = 3 independent mice were used and error bar represents mean ± SD. (two-tailed, unpaired t-test with equal variances: P = 0.001. (c) Cells were imaged every 40 s for 2.0 h. FITC containing phagosomes were detected at f + 80 s (where f is frame 116 out of 271, an arbitrary frame chosen that precedes observation of phagocytic events after addition of S. aureus). Concomitant accumulation of RFP-SCRIB around the FITC-S. aureus(t = 480 s) intensity that localized to and around the site of bacterial engulfment as monitored with the FITC S. aureus signal, white arrow. Scale bar is 5 μm. d Quantification of FITC S. aureus (green line) and RFP-SCRIB (red line) signal intensity at the point of phagocytosis that correlates with the time frames shown in c. Increases in RFP-SCRIB intensity largely follow the same pattern and colocalize with FITC S. aureus at the time of phagocytosis. e Cells were incubated with phrodo-S. aureus for 1 h and then fixed for immunofluorescence staining. Representative and magnified inset images are shown. Green is pHrodo-S. aureus; red is SCRIB. Scale bar is 5μm. f RAW 264.7shSCRIB cells lacking endogenous SCRIB were used to re-express wild type SCRIB (WT.Rescue), or with SCRIB with point mutation in the LRR domain (P305L) (P305L.Rescue). n = 3 biologically independent experiments (mean ± SD). (two-tailed, unpaired t-test with equal variances: P = 0.01). Unprocessed scans of full blots/gels are shown in Supplementary Fig. 6.
Supplementary Figure 5 SCRIB regulated inflammatory response.
(a) Bone marrow cells were isolated from ishSCRIB mice with normal food. Cells were split into two groups: in present of Doxycycline (1 μg ml−1) or not in present of Doxycycline. After 7 day of culture, cells were treated with LPS (100 ng ml−1) for 2.0 h. RNA were isolated from untreated cells and treated cells and then followed by reverse transcripts PCRs. cDNA were used for Taqman array mouse inflammation gene set. Data shown is average of results from 2 mice. (b) Lysates from RAW 264.7shLuc (control) or shScrib treated with LPS (100 ng ml−1) for indicated time point, immunoblotted for p-IkBα, IkBα, and Actin. (c) Confocal microscope images of RAW 264.7 cells before or after stimulated with LPS for 5 min, 15 min, 30 min or 3 h, fixed and stained for p65 (green) and DAPI (blue). Representative images were shown on the left panel (scale bars represent 50 μm) and quantification of p65 translocate to the nuclear were shown on the bottom right panel. n = 3 biologically independent experiments (mean ± SD). (two-tailed, unpaired t-test with equal variances: P = 0.01). BMDCs from WT or ishSCRIB mice were grown as described in Methods with or without Dox. On day 8, DCs were harvested and stimulated with LPS d or CpG e for 18h at the concentrations indicated. Data shown is representative of reproducible results from 2 mice d DC cells were then stained for indicated markers and analyzed by flow cytometry. Unprocessed scans of full blots/gels are shown in Supplementary Fig. 6.
Supplementary information
Supplementary Information
Supplementary Information (PDF 7026 kb)
Supplementary Table 1
Supplementary Information (XLSX 13 kb)
Supplementary Table 2
Supplementary Information (XLSX 49 kb)
Supplementary Table 3
Supplementary Information (XLS 568 kb)
Rights and permissions
About this article
Cite this article
Zheng, W., Umitsu, M., Jagan, I. et al. An interaction between Scribble and the NADPH oxidase complex controls M1 macrophage polarization and function. Nat Cell Biol 18, 1244–1252 (2016). https://doi.org/10.1038/ncb3413
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3413
This article is cited by
-
Hepatocellular carcinoma-derived high mobility group box 1 triggers M2 macrophage polarization via a TLR2/NOX2/autophagy axis
Scientific Reports (2020)
-
The Scribble family in cancer: twentieth anniversary
Oncogene (2020)
-
MicroRNA-296: a promising target in the pathogenesis of atherosclerosis?
Molecular Medicine (2018)