Abstract
MicroRNAs play diverse roles in both normal and malignant stem cells. Focusing on miRs and/or miR∗s abundant in squamous cell carcinoma (SCC) stem cells, we engineer an efficient, strand-specific expression library, and apply functional genomics screening in mice to identify which of 169 cancer-associated miRs are key drivers in malignant progression. Not previously linked functionally to cancer, miR-21∗ was the second top hit, surfacing in >12% of tumours. miR-21∗ also correlates with poor prognosis in human SCCs and enhances tumour progression in xenografts. On deleting the miR-21 gene and rescuing each strand separately, we document the dual, but independent, oncogenicity of miR-21 and miR-21∗. A cohort of predicted miR-21∗ targets inversely correlate with miR-21∗ in SCCs. Of particular interest is Phactr4, which we show is a miR-21∗ target in SCCs, acting through the Rb/E2F cell cycle axis. Through in vivo physiological miR screens, our findings add an interesting twist to an increasingly important oncomiR locus.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Change history
10 December 2015
In the version of this Article originally published online, the third sentence of the abstract should have read “Not previously linked functionally to cancer, miR-21* was the second top hit, surfacing in >12% of tumours”. This is now correct in all versions of the Article.
References
Bartel, D. P. MicroRNAs: target recognition and regulatory functions. Cell. 136, 215–233 (2009).
Lu, J. et al. MicroRNA expression profiles classify human cancers. Nature. 435, 834–838 (2005).
Lujambio, A. & Lowe, S. W. The microcosmos of cancer. Nature. 482, 347–355 (2012).
Di Leva, G., Garofalo, M. & Croce, C. M. MicroRNAs in cancer. Annu. Rev. Pathol. 9, 287–314 (2014).
Adams, B. D., Kasinski, A. L. & Slack, F. J. Aberrant regulation and function of MicroRNAs in cancer. Curr. Biol. 24, R762–R776 (2014).
Darido, C. et al. Targeting of the tumor suppressor GRHL3 by a miR-21-dependent proto-oncogenic network results in PTEN loss and tumorigenesis. Cancer Cell 20, 635–648 (2011).
Ma, X. et al. Loss of the miR-21 allele elevates the expression of its target genes and reduces tumorigenesis. Proc. Natl Acad. Sci. USA 108, 10144–10149 (2011).
Bhandari, A. et al. The Grainyhead transcription factor Grhl3/Get1 suppresses miR-21 expression and tumorigenesis in skin: modulation of the miR-21 target MSH2 by RNA-binding protein DND1. Oncogene 32, 1497–1507 (2013).
Benaich, N. et al. Rewiring of an epithelial differentiation factor, miR-203, to inhibit human squamous cell carcinoma metastasis. Cell Rep. 9, 104–117 (2014).
Zhang, L., Ge, Y. & Fuchs, E. miR-125b can enhance skin tumor initiation and promote malignant progression by repressing differentiation and prolonging cell survival. Genes Dev. 28, 2532–2546 (2014).
Schober, M. & Fuchs, E. Tumor-initiating stem cells of squamous cell carcinomas and their control by TGF-β and integrin/focal adhesion kinase (FAK) signaling. Proc. Natl Acad. Sci. USA 108, 10544–10549 (2011).
Lapouge, G. et al. Skin squamous cell carcinoma propagating cells increase with tumour progression and invasiveness. Embo J. 31, 4563–4575 (2012).
Oshimori, N., Oristian, D. & Fuchs, E. TGF-β promotes heterogeneity and drug resistance in squamous cell carcinoma. Cell 160, 963–976 (2015).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Adam, R. C. et al. Pioneer factors govern super-enhancer dynamics in stem cell plasticity and lineage choice. Nature 521, 366–370 (2015).
Beronja, S., Livshits, G., Williams, S. & Fuchs, E. Rapid functional dissection of genetic networks via tissue-specific transduction and RNAi in mouse embryos. Nat. Med. 16, 821–827 (2010).
Beronja, S. et al. RNAi screens in mice identify physiological regulators of oncogenic growth. Nature 501, 185–190 (2013).
Schramek, D. et al. Direct in vivo RNAi screen unveils myosin IIa as a tumor suppressor of squamous cell carcinomas. Science 343, 309–313 (2014).
Quintanilla, M., Brown, K., Ramsden, M. & Balmain, A. Carcinogen-specific mutation and amplification of Ha-ras during mouse skin carcinogenesis. Nature 322, 78–80 (1986).
Nassar, D., Latil, M., Boeckx, B., Lambrechts, D. & Blanpain, C. Genomic landscape of carcinogen-induced and genetically induced mouse skin squamous cell carcinoma. Nat. Med. 21, 946–954 (2015).
Ma, L., Teruya-Feldstein, J. & Weinberg, R. A. Tumour invasion and metastasis initiated by microRNA-10b in breast cancer. Nature. 449, 682–688 (2007).
Gregory, P. A. et al. The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat. Cell Biol. 10, 593–601 (2008).
Taylor, M. A., Sossey-Alaoui, K., Thompson, C. L., Danielpour, D. & Schiemann, W. P. TGF-β upregulates miR-181a expression to promote breast cancer metastasis. J. Clin. Invest. 123, 150–163 (2013).
Chen, X. et al. Endogenous expression of Hras(G12V) induces developmental defects and neoplasms with copy number imbalances of the oncogene. Proc. Natl Acad. Sci. USA 106, 7979–7984 (2009).
Hatley, M. E. et al. Modulation of K-Ras-dependent lung tumorigenesis by microRNA-21. Cancer Cell 18, 282–293 (2010).
Medina, P. P., Nolde, M. & Slack, F. J. OncomiR addiction in an in vivo model of microRNA-21-induced pre-B-cell lymphoma. Nature 467, 86–90 (2010).
Fujita, S. et al. miR-21 gene expression triggered by AP-1 is sustained through a double-negative feedback mechanism. J. Mol. Biol. 378, 492–504 (2008).
Talotta, F. et al. An autoregulatory loop mediated by miR-21 and PDCD4 controls the AP-1 activity in RAS transformation. Oncogene 28, 73–84 (2009).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
John, B. et al. Human MicroRNA targets. PLoS Biol. 2, e363 (2004).
Lewis, B. P., Burge, C. B. & Bartel, D. P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120, 15–20 (2005).
Frankel, L. B. et al. Programmed cell death 4 (PDCD4) is an important functional target of the microRNA miR-21 in breast cancer cells. J. Biol. Chem. 283, 1026–1033 (2008).
Solimini, N. L. et al. STOP gene Phactr4 is a tumor suppressor. Proc. Natl Acad. Sci. USA 110, E407-414 (2013).
Kim, T. H., Goodman, J., Anderson, K. V. & Niswander, L. Phactr4 regulates neural tube and optic fissure closure by controlling PP1-, Rb-, and E2F1-regulated cell-cycle progression. Dev Cell. 13, 87–102 (2007).
Pink, R. C. et al. The passenger strand, miR-21-3p, plays a role in mediating cisplatin resistance in ovarian cancer cells. Gynecol. Oncol. 137, 143–151 (2015).
Yi, R. et al. Morphogenesis in skin is governed by discrete sets of differentially expressed microRNAs. Nat. Genet. 38, 356–362 (2006).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Ge, Y., Sun, Y. & Chen, J. IGF-II is regulated by microRNA-125b in skeletal myogenesis. J. Cell Biol. 192, 69–81 (2011).
Keyes, B. E. et al. Nfatc1 orchestrates aging in hair follicle stem cells. Proc. Natl Acad. Sci. USA 110, E4950–E4959 (2013).
Acknowledgements
We especially thank D. Schramek for intellectual input into the design of the allograft assay. We also thank D. Schramek and A. Sendoel for advice regarding data presentation; J. Racelis for technical assistance; A. Aldeguer, L. Polak, J. Levorse, S. Hacker, M. Sribour, D. Oristian and N. Stokes for assistance with mouse handling and experiments; and Z. Shen, N. Oshimori, S. Luo, K. Lay, M. Kadaja, B. Keyes, A. Asare, M. Laurin, R. Adam and A. Kulukian for helpful discussions. We thank the Comparative Bioscience Center (AAALAC accredited) for care of mice in accordance with National Institutes of Health (NIH) guidelines; Rockefeller Genomics Resource Center (C. Zhao, Director) and Weill Cornell Genomics Resource Center (J. Xiang, Director) for sequencing; Flow Cytometry facility (S. Mazel, Director) for FACS sorting. E.F. is an Investigator of the Howard Hughes Medical Institute. Y.G. is the recipient of a Women & Science Postdoctoral Fellowship, and a Department of Defense Breast Cancer Postdoctoral Fellowship.
Author information
Authors and Affiliations
Contributions
Y.G. and E.F. designed the experiments and wrote the manuscript. L.Z. designed and characterized the SA-miR vector. Y.G. and L.Z. performed the miR sequencing on FACS-sorted cells and made the lentiviral SA-miR constructs (Figs 1a and 2). Y.G. performed all other experiments and analyses (Figs 1b, c and 3–8). B.R. carried out the TCGA data analyses on miR-21 and miR-21∗ targets. M.N. assisted in CRISPR/CAS of Phactr4. All authors provided intellectual input, and vetted and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 miRNA profiling of skin progenitors from different stages of development, homeostasis and tumourigenesis.
(a) FACS scheme for normal stem cells and progenitors at different stages of skin development, and for papilloma and SCC progenitors and their progenies at different stages of malignant progression. Please refer to Methods for detailed gates. (b,c) Quantitative PCR to quantify the relative mRNA levels for a group of cell-type specific markers to validate the purity of FACS isolated populations: Lhx2 and Pdzrn3 for HF progenitors (E17, P0), Cd34 and Lhx2 for adult HF stem cells, Trp63 for E17 Epidermal progenitors, Sca1 for P0 and adult epidermal progenitors; Ccnd1, Cd44, Krt6 and Vim for SCC-SC upregulated markers and Krt10, Lhx2 and Nfatc1 for SCC-SC downregulated markers compared to HF-SCs. Paired t-tests were used. At least three biological replicates were performed; shown are mean ± s.d. n = 3 independent experiments. ∗P < 0.05.∗∗P < 0.01. (d) SCC signature miRNAs (cut off 100 RRMM) that are conserved between mouse and human and exhibit ≥2X change in basal cells (BCs) of SCCs relative to normal SCs (DEseq P value < 0.05). (e) qPCRs confirmed a subset of miRNA signatures discovered in miRNA profiling. Paired t-tests were used. At least three biological replicates were performed; shown are mean ± s.d. n = 3 independent experiments. ∗P < 0.05.∗∗P < 0.01. (f) miRNA in situ hybridizations of known and SCC-related miRNAs in development and malignant transformation. These results showed that miR-205, an essential miRNA known to govern skin epithelium homeostasis, is broadly and abundantly expressed throughout skin development and transformation; miR-203, known to promote skin stratification and suppress metastasis, is specifically enriched in the suprabasal layers at all stages; miR-125b known to maintain normal SC self-renewal and drive skin tumourigenesis is transiently upregulated during early stage of benign tumourigenesis; miR-744 is broadly expressed during embryogenesis and later becomes more restricted to suprabasal layers during adult and tumour development. At least three biological replicates were performed; shown are representative images. Scale bar = 50 μm.
Supplementary Figure 2 Design and characterization of lentiviral SA-miRNA expression vectors.
(a) Schematic to show differences in miR-21 and miR-21∗ sequences and the corresponding strand-specific SA-miRs.(b) SA-miRNAs are pooled and packaged into lentivirus to transduce cultured mouse SCC cells. Cells were isolated 2 days post infection, sorted with endogenous GFP, and subjected to deep sequencing. The normalized reads for position 1 and 2 isomiRs were plotted to demonstrate faithful 5’ end processing using SA-miRNA system. Grey bar represents control cells transduced with SA vector driving the expression of a scrambled sequence alone, reflecting endogenous isomiR levels. A different Scrambled sequence was used from the SA-Scr shown in the graph. (c) To compare the efficacy of our SA-miRs with oft-used SH-miRs (pENTR/pSM2 (U6), Addgene 17387)40, we performed luciferase reporter assays to examine the effects of miR-125b and miR-203 on their established target genes Scarb1 and p6310,41, respectively, expressed in either the SH-miR or the SA-miR backbone. All vectors were co-transfected with miR mimics at equal concentration in controls and experiments. Similar sensor-mediated inhibition (achieved by anti-sense sequences of miR-125b or miR-203) was observed from both, but stronger target inhibition was observed only from SA-miRNA mediated miR-125b and miR-203 expression when using 3’UTR reporters. Paired t-tests were used to compare SH-miR to SH-control, and SA-miR to SA-control, respectively. At least three biological replicates were performed; shown are mean ± s.d. n = 3 independent experiments. ∗∗P < 0.01. N.S. Not Significant.
Supplementary Figure 3 More candidate oncomiRs in skin SCCs identified from in vivo pooled screen with SA-miRNA library.
(a) Caspase 3 and Ki67 staining of tumours show unchanged apoptosis and heightened mitosis, respectively, in oncogenic HRas tumours derived from the SA-miRNA library transduced cohort compared to that of SA-Scr library. Scale bar = 50 μm. At least five biological replicates were performed; shown are representative images. Ki67 positive nuclei as a percentage of total nuclei was quantified and shown. Paired t-tests were used to compare SA-miR to SA-Scr. Shown are mean ± s.d. n = 5 independent experiments. ∗∗P < 0.01). (b) Targeted deep sequencing of SA-miRNA inserts in the tumours emerging from animals transduced with the SA-miR library showed that three different miRNA families had miRs that were overrepresented in tumours. OncomiRs miR-125b and miR-205 were also captured. Clonal dominance was not observed in regions of the backskin adjacent to the tumour (shown) or in E12.5 or P50 skin (see Fig. 3). Numbers of tumours are shown in parentheses. The frequency of clonal dominance in these tumours was significantly less than that seen for miR-21 and miR-21∗ (see Fig. 3).
Supplementary Figure 4 Validation of miR-21∗ in driving SCC progression in vivo.
(a) Strand-specific expression of SA-miR-21 and SA-miR-21∗ vectors in vivo. Note that SA-miR-21 elevated miR-21 but did not affect miR-21∗ levels; SA-miR-21∗ elevated miR-21∗, but did not affect miR-21 levels in the skin epidermis 3 days after infection. Paired t-tests were used to compare SA-miR to SA-Scr. At least three biological replicates were performed; shown are mean ± s.d. n = 3 independent experiments. ∗P < 0.05,∗∗P < 0.01. (b) In situ hybridizations on day 7 post-wound mouse backskin, showing specific miR-21 and miR-21∗ probe signals in heterozygous but not miR-21-/- skin. MiR-21 locus was transcriptionally activated during wounding15 and has been proposed to be regulated by a number of transcription factors in the context of normal skin and cancer42. Scale bar = 50 μm. (c) SCs purified from the bulge of HFs from WT skin and then transduced with either SA-Scr, SA-miR-21 or SA-miR-21∗ were subjected to colony formation assays in culture. Colonies were stained with Rhodamine B, 2 weeks after seeding at the concentrations indicated at left. Numbers of holoclones (defined by clones >20 μm in diameter and typical of stem cells) are shown in the graph below. Note elevation in the number of stem cells, particularly when miR-21∗ is elevated. (d) SA-miR overexpression of miR-21 or miR-21∗ in keratinocytes resulted in their increased proliferation (top graph) and decreased apoptosis in suspension cultures (bottom graph). Paired t-tests were used to compare SA-miRs to SA-Scr. At least three biological replicates were performed; shown are mean ± s.d. n = 3 independent experiments. If not associated with passenger strand miR, ∗ denotes statistical significance. ∗P < 0.05,∗∗P < 0.01.
Supplementary Figure 5 CRISPR/CAS strategy to knockout miR-21 locus.
Surveyor assay: the CRISPR/CAS-edited mir-21 locus is amplified by PCR from ΔSCC genomic DNA and then digested by an endonuclease enzyme that only cuts the non-matched DNA position (introduced by CRISPR editing). If editing is successful, the 718 bp PCR product is cleaved into 462 and 256 bp smaller bands.
Supplementary Figure 6 miR-21∗ expression and CRIPSR/CAS in human SCCs.
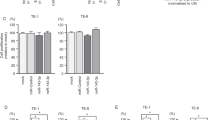
(a) miRNA sequencing data of SCCs show that miR-21∗ but not miR-31∗ or miR-205∗ is stabilized in mouse skin SCCs and human HNSCCs, even though all of these miR guide counterparts are abundant in these tumours. Shown are mean ± s.d. n = 3 independent experiments. (b) In situ hybridization of miR-21 and miR-21∗ in paraffin-fixed human tissue arrays with HNSCCs and normal tissues. Note that hybridizations are not as elegant as with frozen sections (see main Figures) but the data are still clear. Scale bar = 100 μm. (c) Demonstration that the SA-miR vectors used to drive miR-21 and miR-21∗ achieve elevated expression in human FaDu and HaCaT cells. At least five biological replicates were performed; shown are mean ± s.d. n = 3 independent experiments. ∗P < 0.05,∗∗P < 0.01. N.S. Not Significant. Paired t-tests were used to compare SA-miRs to SA-Scr. (d–f) CRISPR/CAS edited human A431 SCC cells, surveyor assay, and deficiency in tumourigenesis. For both mouse and human SCCs, top three predicted off-target sites were sequenced and no mutations were detected in the three clones tested (see Methods). Shown are mean ± s.d. n = 3 independent experiments. Two-way ANOVA with repeated measurement was performed.
Supplementary Figure 7 More miR-21∗ and miR-21 putative targets in SCCs.
(a) 8 of miR-21∗ and 6 of miR-21 predicted targets are down-regulated in HNSCCs compared to normal tissues. Shown are mean ± s.d. n = 31 for normal tissues and n = 263 for tumours. Unpaired t-test was used to compare normal to tumour samples. (b) Predicted targets that inversely correlate with miR-21∗ or miR-21 levels in tumours, respectively. Shown are mean ± s.d. n = 26 for tumour samples. Paired t-test was used to compare low to high expression tumour samples. (c) In addition to miR-21∗ target PHACTR4 (see main text), RGMA (miR-21∗ target), PDCD4(miR-21 target) and YOD1(miR-21 target) expression levels inversely correlated with poor prognosis in HNSCCs n = 7 ∼ 330 tumour samples. Log-rank test method was used to compare patient survival between high and low expression samples.
Supplementary Figure 8 miR-21∗ targets Phactr4 in SCCs.
(a,b) Western blot showing suppressed levels of Phactr4 resulting from SA-miR-21∗ overexpression (a), and Phactr4 levels from WT or ΔPP1 mutant expression (b). In a, the numbers indicate densitometry values normalized against α-tubulin and measured by ImageJ. At least three biological replicates were performed; shown are representative images.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1137 kb)
Supplementary Table 1
Supplementary Information (XLSX 42 kb)
Supplementary Table 2
Supplementary Information (XLSX 39 kb)
Supplementary Table 3
Supplementary Information (XLSX 9 kb)
Supplementary Table 4
Supplementary Information (XLSX 291 kb)
Supplementary Table 5
Supplementary Information (XLSX 57 kb)
Supplementary Table 6
Supplementary Information (XLSX 68 kb)
Supplementary Table 7
Supplementary Information (XLSX 35 kb)
Supplementary Table 8
Supplementary Information (XLSX 48 kb)
Rights and permissions
About this article
Cite this article
Ge, Y., Zhang, L., Nikolova, M. et al. Strand-specific in vivo screen of cancer-associated miRNAs unveils a role for miR-21∗ in SCC progression. Nat Cell Biol 18, 111–121 (2016). https://doi.org/10.1038/ncb3275
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3275
This article is cited by
-
MicroRNA-21 guide and passenger strand regulation of adenylosuccinate lyase-mediated purine metabolism promotes transition to an EGFR-TKI-tolerant persister state
Cancer Gene Therapy (2022)
-
A stress-induced miR-31–CLOCK–ERK pathway is a key driver and therapeutic target for skin aging
Nature Aging (2021)
-
miR-22 promotes stem cell traits via activating Wnt/β-catenin signaling in cutaneous squamous cell carcinoma
Oncogene (2021)
-
HIF1A activates the transcription of lncRNA RAET1K to modulate hypoxia-induced glycolysis in hepatocellular carcinoma cells via miR-100-5p
Cell Death & Disease (2020)
-
Regulation of PTEN expression by noncoding RNAs
Journal of Experimental & Clinical Cancer Research (2018)


