Abstract
Dense multicilia in higher vertebrates are important for luminal flow and the removal of thick mucus. To generate hundreds of basal bodies for multiciliogenesis, specialized terminally differentiated epithelial cells undergo massive centriole amplification. In proliferating cells, however, centriole duplication occurs only once per cell cycle. How cells ensure proper regulation of centriole biogenesis in different contexts is poorly understood. We report that the centriole amplification is controlled by two duplicated genes, Cep63 and Deup1. Cep63 regulates mother-centriole-dependent centriole duplication. Deup1 governs deuterosome assembly to mediate large-scale de novo centriole biogenesis. Similarly to Cep63, Deup1 binds to Cep152 and then recruits Plk4 to activate centriole biogenesis. Phylogenetic analyses suggest that Deup1 diverged from Cep63 in a certain ancestor of lobe-finned fishes during vertebrate evolution and was subsequently adopted by tetrapods. Thus, the Cep63 gene duplication has enabled mother-centriole-independent assembly of the centriole duplication machinery to satisfy different requirements for centriole number.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Fliegauf, M., Benzing, T. & Omran, H. When cilia go bad: cilia defects and ciliopathies. Nat. Rev. Mol. Cell. Biol. 8, 880–893 (2007).
Nigg, E. A. & Raff, J. W. Centrioles, centrosomes, and cilia in health and disease. Cell 139, 663–678 (2009).
Beisson, J. & Wright, M. Basal body/centriole assembly and continuity. Curr. Opin. Cell Biol. 15, 96–104 (2003).
Cunha-Ferreira, I., Bento, I. & Bettencourt-Dias, M. From zero to many: control of centriole number in development and disease. Traffic 10, 482–498 (2009).
Nigg, E. A. & Stearns, T. The centrosome cycle: centriole biogenesis, duplication and inherent asymmetries. Nat. Cell Biol. 13, 1154–1160 (2011).
Lemullois, M., Boisvieux-Ulrich, E., Laine, M. C., Chailley, B. & Sandoz, D. Development and functions of the cytoskeleton during ciliogenesis in metazoa. Biol. Cell 63, 195–208 (1988).
Anderson, R. G. & Brenner, R. M. The formation of basal bodies (centrioles) in the Rhesus monkey oviduct. J. Cell Biol. 50, 10–34 (1971).
Sorokin, S. P. Reconstructions of centriole formation and ciliogenesis in mammalian lungs. J. Cell Sci. 3, 207–230 (1968).
Gonczy, P. Towards a molecular architecture of centriole assembly. Nat. Rev. Mol. Cell Biol. 13, 425–435 (2012).
Azimzadeh, J. & Marshall, W. F. Building the centriole. Curr. Biol. 20, R816–825 (2010).
Hatch, E. M., Kulukian, A., Holland, A. J., Cleveland, D. W. & Stearns, T. Cep152 interacts with Plk4 and is required for centriole duplication. J. Cell Biol. 191, 721–729 (2010).
Cizmecioglu, O. et al. Cep152 acts as a scaffold for recruitment of Plk4 and CPAP to the centrosome. J. Cell Biol. 191, 731–739 (2010).
Dzhindzhev, N. S. et al. Asterless is a scaffold for the onset of centriole assembly. Nature 467, 714–718 (2010).
Sir, J. H. et al. A primary microcephaly protein complex forms a ring around parental centrioles. Nat. Genet. 43, 1147–1153 (2011).
Lawo, S., Hasegan, M., Gupta, G. D. & Pelletier, L. Subdiffraction imaging of centrosomes reveals higher-order organizational features of pericentriolar material. Nat. Cell Biol. 14, 1148–1158 (2012).
Mennella, V. et al. Subdiffraction-resolution fluorescence microscopy reveals a domain of the centrosome critical for pericentriolar material organization. Nat. Cell Biol. 14, 1159–1168 (2012).
Sonnen, K. F., Schermelleh, L., Leonhardt, H. & Nigg, E. A. 3D-structured illumination microscopy provides novel insight into architecture of human centrosomes. Biol. Open 1, 965–976 (2012).
Kleylein-Sohn, J. et al. Plk4-induced centriole biogenesis in human cells. Dev. Cell 13, 190–202 (2007).
Tang, C. J. et al. The human microcephaly protein STIL interacts with CPAP and is required for procentriole formation. EMBO J. 30, 4790–4804 (2011).
Tang, C. J., Fu, R. H., Wu, K. S., Hsu, W. B. & Tang, T. K. CPAP is a cell-cycle regulated protein that controls centriole length. Nat. Cell Biol. 11, 825–831 (2009).
Kohlmaier, G. et al. Overly long centrioles and defective cell division on excess of the SAS-4-related protein CPAP. Curr. Biol. 19, 1012–1018 (2009).
Schmidt, T. I. et al. Control of centriole length by CPAP and CP110. Curr. Biol. 19, 1005–1011 (2009).
Arquint, C., Sonnen, K. F., Stierhof, Y. D. & Nigg, E. A. Cell-cycle-regulated expression of STIL controls centriole number in human cells. J. Cell Sci. 125, 1342–1352 (2012).
Vulprecht, J. et al. STIL is required for centriole duplication in human cells. J. Cell Sci. 125, 1353–1362 (2012).
Strnad, P. et al. Regulated HsSAS-6 levels ensure formation of a single procentriole per centriole during the centrosome duplication cycle. Dev. Cell 13, 203–213 (2007).
Dirksen, E. R. Centriole morphogenesis in developing ciliated epithelium of the mouse oviduct. J. Cell Biol. 51, 286–302 (1971).
You, Y., Richer, E. J., Huang, T. & Brody, S. L. Growth and differentiation of mouse tracheal epithelial cells: selection of a proliferative population. Am. J. Physiol. Lung. Cell Mol. Physiol. 283, L1315–1321 (2002).
Vladar, E. K. & Stearns, T. Molecular characterization of centriole assembly in ciliated epithelial cells. J. Cell Biol. 178, 31–42 (2007).
Cao, J. et al. miR-129-3p controls cilia assembly by regulating CP110 and actin dynamics. Nat. Cell Biol. 14, 697–706 (2012).
Piperno, G. & Fuller, M. T. Monoclonal antibodies specific for an acetylated form of alpha-tubulin recognize the antigen in cilia and flagella from a variety of organisms. J. Cell Biol. 101, 2085–2094 (1985).
Avidor-Reiss, T. & Gopalakrishnan, J. Building a centriole. Curr. Opin. Cell Biol. 25, 72–77 (2013).
Habedanck, R., Stierhof, Y. D., Wilkinson, C. J. & Nigg, E. A. The Polo kinase Plk4 functions in centriole duplication. Nat. Cell Biol. 7, 1140–1146 (2005).
Blacque, O. E. et al. Loss of C. elegans BBS-7 and BBS-8 protein function results in cilia defects and compromised intraflagellar transport. Genes. Dev. 18, 1630–1642 (2004).
Krock, B. L. & Perkins, B. D. The intraflagellar transport protein IFT57 is required for cilia maintenance and regulates IFT-particle-kinesin-II dissociation in vertebrate photoreceptors. J. Cell Sci. 121, 1907–1915 (2008).
Nakagawa, Y., Yamane, Y., Okanoue, T. & Tsukita, S. Outer dense fibre 2 is a widespread centrosome scaffold component preferentially associated with mother centrioles: its identification from isolated centrosomes. Mol. Biol. Cell 12, 1687–1697 (2001).
Zardoya, R. & Meyer, A. The complete DNA sequence of the mitochondrial genome of a ‘living fossil,’ the coelacanth (Latimeria chalumnae). Genetics 146, 995–1010 (1997).
Amemiya, C. T. et al. The African coelacanth genome provides insights into tetrapod evolution. Nature 496, 311–316 (2013).
Bertrand, S. & Escriva, H. Evolutionary crossroads in developmental biology: amphioxus. Development 138, 4819–4830 (2011).
Canestro, C., Bassham, S. & Postlethwait, J.H. Seeing chordate evolution through the Ciona genome sequence. Genome. Biol. 4, 208 (2003).
Hayes, J. M. et al. Identification of novel ciliogenesis factors using a new in vivo model for mucociliary epithelial development. Dev. Biol. 312, 115–130 (2007).
Zhao, C. & Malicki, J. Nephrocystins and MKS proteins interact with IFT particle and facilitate transport of selected ciliary cargos. EMBO J. 30, 2532–2544 (2011).
Wu, S., Ma, L., Wu, Y., Zeng, R. & Zhu, X. Nudel is crucial for the WAVE complex assembly in vivo by selectively promoting subcomplex stability and formation through direct interactions. Cell Res. 22, 1270–1284 (2012).
Wickner, S., Maurizi, M. R. & Gottesman, S. Posttranslational quality control: folding, refolding, and degrading proteins. Science 286, 1888–1893 (1999).
Zeng, L. et al. The m-subunit of murine translation initiation factor eIF3 maintains the integrity of the eIF3 complex and is required for embryonic development, homeostasis, and organ size control. J. Biol. Chem. 288, 30087–30093 (2013).
Pearson, C. G. & Winey, M. Basal body assembly in ciliates: the power of numbers. Traffic 10, 461–471 (2009).
Azimzadeh, J., Wong, M. L., Downhour, D. M., Sanchez Alvarado, A. & Marshall, W. F. Centrosome loss in the evolution of planarians. Science 335, 461–463 (2012).
Meyer, A. & Schartl, M. Gene and genome duplications in vertebrates: the one-to-four (-to-eight in fish) rule and the evolution of novel gene functions. Curr. Opin. Cell Biol. 11, 699–704 (1999).
Kramer-Zucker, A. G. et al. Cilia-driven fluid flow in the zebrafish pronephros, brain and Kupffer’s vesicle is required for normal organogenesis. Development 132, 1907–1921 (2005).
Kemp, A. The role of epidermal cilia in development of the Australian lungfish, Neoceratodus forsteri (Osteichthyes: Dipnoi). J. Morphol. 228, 203–221 (1996).
Schermelleh, L., Heintzmann, R. & Leonhardt, H. A guide to super-resolution fluorescence microscopy. J. Cell Biol. 190, 165–175 (2010).
Wang, F., Zhang, Q., Cao, J., Huang, Q. & Zhu, X. The microtubule plus end-binding protein EB1 is involved in Sertoli cell plasticity in testicular seminiferous tubules. Exp. Cell Res. 314, 213–226 (2008).
Liu, Y., Pathak, N., Kramer-Zucker, A. & Drummond, I. A. Notch signalling controls the differentiation of transporting epithelia and multiciliated cells in the zebrafish pronephros. Development 134, 1111–1122 (2007).
Showell, C. & Conlon, F. L. Natural mating and tadpole husbandry in the western clawed frog Xenopus tropicalis. Cold. Spring Harbor. Protocols. 2009 pdb prot5292 (2009).
Boulter, E., Grall, D., Cagnol, S. & Van Obberghen-Schilling, E. Regulation of cell-matrix adhesion dynamics and Rac-1 by integrin linked kinase. FASEB J. 20, 1489–1491 (2006).
Sun, Q., Yu, X. C. & Margolin, W. Assembly of the FtsZ ring at the central division site in the absence of the chromosome. Mol. Microbiol. 29, 491–503 (1998).
Shan, Y. et al. Nudel and FAK as antagonizing strength modulators of nascent adhesions through paxillin. PLoS Biol. 7 e1000116 (2009).
Acknowledgements
The authors thank Y. Zheng, Y. Shen and J. Ten for helpful suggestions; E. Nigg for Cep152 antibody; S. Li and Z. Li for technical support on 3D-SIM and H. Dai, T. Zhang, F. Qin and X. Zhu for technical assistance. This work was supported by the Chinese Academy of Sciences (XDA01010107), the National Basic Research Program of China (2012CB945003 and 2010CB912102), the National Science Foundation of China (31330045, 31271427 and 91129000) and Shanghai Municipal Science and Technology Commission (12JC1409900). X.Y. acknowledges the support of the SA-SIBS Scholarship Program.
Author information
Authors and Affiliations
Contributions
X.Y. and X.Z. conceived and directed the project. H.Z. carried out most experimental work. L.Z. carried out the frog and transmission EM experiments. Y.Z did the experiments in bacteria and drew all illustrative models. J.C. established the culture and infection conditions for MTECs. S.L. found that CEP152 RNAi downregulated CEP63. Q.H. generated the homemade antibodies. T.X. and X.H. provided the 3D-SIM and frog systems. X.Z., X.Y. and H.Z. designed experiments, interpreted data, and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Sequence alignment of Deup1 and Cep63.
The protein sequence alignment was generated by the Clustal X 2.0 multiple sequence alignment program using default parameters. The protein sequences were from GenBank accessions numbers NM_181645 (human DEUP1), KC211186 (mouse Deup1), NM_025180 (human CEP63), and KC211185 (mouse Cep63). The histogram depicts conservation between the two proteins. Related to Fig. 1.
Supplementary Figure 2 The aggregates formed by GFP–Cep63 or GFP–Cep152 do not support de novo procentriole formation.
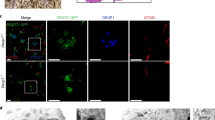
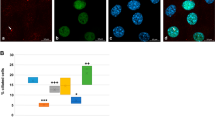
(a) Expression of GFP-tagged Deup1, Cep63 or Cep152 in U2OS cells. (b,c) Analyses for procentriole formation. The arrowheads and arrows point to typical mother centrioles and GFP-positive aggregates, respectively. The bright SAS-6 foci, which mark the cartwheels, are used as procentriole markers in the quantification analyses. Only interphase cells containing MCD procentrioles, thus in the S or G2 phase, were scored (n = 30 cells from triplicates). GFP–Deup1-expressing cells served as the positive control for de dovo procentriole biogenesis. (d) Endogenous CEP152 exhibits punctate distributions in the GFP–Cep63 aggregates. Such a pattern is in sharp contrast to the ring-shaped distributions of CEP152 in the deuterosomes (arrow in the GFP–Deup1-positive cell) and MCD cradles (arrowheads) (also see Fig. 1h), suggesting that the GFP–Cep63 aggregates are disorganised. Apparently, the punctate CEP152 is unable to induce the cartwheel assembly, not to mention de novo centriole formation. Related to Fig. 1.
Supplementary Figure 3 Immunoblotting results for domain mapping.
(a) Mapping of the Deup1- and CEP63-interactiing domains of Cep152. The numbers in the diagrams indicate amino acid positions. Exogenous Cep152, its mutants, or firefly luciferase (Luc) was expressed alone or together with Flag-Deup1 in HEK293T cells, as indicated. Coimmunoprecipitation was then performed using anti-Flag resin. Luciferase served as the negative control. The asterisks indicate the positions of full-length fusion proteins. (b) Mapping of the CEP152-interacting domain of Deup1. Flag-tagged Deup1 or mutants were expressed in HEK293T cells. Coimmunoprecipitation was then performed. (c) Mapping of the CEP152-interacting domain of Cep63. Flag-tagged Cep63 or mutants were expressed in HEK293T cells. Coimmunoprecipitation was then performed. These results are summarised in Fig. 1j.
Supplementary Figure 4 Effect of Deup1 or Cep63 RNAi in MTECs at ALI d 3.
(a) Cep152 failed to exhibit the ring-shaped deuterosome localization upon Deup1 RNAi. The arrows indicate MCD procentriole. The insets are magnified 2× to show typical de novo procentrioles (arrowheads). (b-d) Ablation of Cep63 alone in MTECs does not block deuterosome formation and centriole amplification. MTECs were infected with lentivirus to express control (Ctrl-i) or Cep63-specific (63-i1 or 63-i2) shRNA together with GFP–Cetn1. The GFP-positive cells at ALI d 3 were sorted out using FACS and subjected to immunoblotting to show RNAi efficiency. α-Tubulin served as the loading control. The insets are magnified to show MCD procentrioles. The statistical results for centriole numbers are presented in Fig. 6d.
Supplementary Figure 5 Validation of the specificity of Dp-MO1 and Dp-MO2 using a GFP reporter.
(a) Diagrams ofin vitro-transcribed mRNAs. The red bars indicate the locations of the MO-targeting sites. (b) Dp-MO1 and Dp-MO2 specifically blocked the translation of the xDpATG-GFP mRNA. Xenopus embryos at the two- or four-cell stage were microinjected with a mixture of 10 ng MO, 100 pg GFP or xDpATG-GFP mRNA, and 50 pg RFP-F mRNA (as an injection marker). The autofluorescence of GFP or RFP was visualised at approximately the developmental stage 26. Related to Fig. 8d.
Supplementary Figure 6 Full scans of original blots.
The boxed regions indicate the areas shown in the figures.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1522 kb)
Supplementary Table 1
Supplementary Information (XLSX 13 kb)
Supplementary Table 2
Supplementary Information (XLSX 12 kb)
Supplementary Table 3
Supplementary Information (XLS 29 kb)
3D model of a part of a cell in stage II immunostained for Deup1 (green), Cep152 (blue), and Centrin (red).
The top and side views are represented in Fig. 4c. (MOV 5587 kb)
3D model of a part of a cell in stage III immunostained for Deup1 (green), Cep152 (blue), and Centrin (red).
The top and side views are represented in Fig. 4c. (MOV 8021 kb)
3D model of a part of a cell in stage IV immunostained for Deup1 (green), Cep152 (blue), and Centrin (red).
The top and side views are represented in Fig. 4c. (MOV 9609 kb)
3D model of a part of a cell in stage V immunostained for Deup1 (green), Cep152 (blue), and Centrin (red).
The top and side views are represented in Fig. 4c. (MOV 9753 kb)
3D model of a part of a cell in stage VI immunostained for Deup1 (green), Cep152 (blue), and Centrin (red).
The top and side views are represented in Fig. 4c. (MOV 7096 kb)
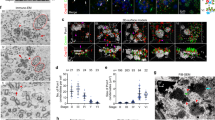
EM images from four consecutive sections of a control (Ctrl-i) and a Deup1-knockdown (Dp-i1) MTEC at ALI d3.
The thickness of each section is 70 nm. Deuterosomes (d), mother centrioles (m), and daughter procentrioles around mother (p) are numbered according to the sequence of appearance. The overlaid images are shown in Fig. 7a. (MOV 1085 kb)
3D model for deuterosome-like structures formed by His-Deup1 (green) in E. coli.
The protein expression was induced with 10 μM IPTG. See Fig. 7d for the original 3D-SIM image. (MOV 4525 kb)
3D model for deuterosome-like structures formed by His-Deup1 (green) in E. coli.
The protein expression was induced with 50 μM IPTG. See Fig. 7d for the original 3D-SIM image. (MOV 5332 kb)
Rights and permissions
About this article
Cite this article
Zhao, H., Zhu, L., Zhu, Y. et al. The Cep63 paralogue Deup1 enables massive de novo centriole biogenesis for vertebrate multiciliogenesis. Nat Cell Biol 15, 1434–1444 (2013). https://doi.org/10.1038/ncb2880
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2880
This article is cited by
-
Dual-color live imaging unveils stepwise organization of multiple basal body arrays by cytoskeletons
EMBO Reports (2024)
-
A local translation program regulates centriole amplification in the airway epithelium
Scientific Reports (2023)
-
Architectural basis for cylindrical self-assembly governing Plk4-mediated centriole duplication in human cells
Communications Biology (2023)
-
Appearing and disappearing acts of cilia
Journal of Biosciences (2023)
-
Genome-wide association study biomarkers in T-cell mediated rejection: selective effect according to the Banff classification
Journal of Nephrology (2022)