Abstract
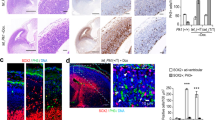
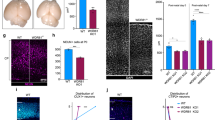
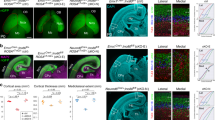
Primary microcephaly 1 is a neurodevelopmental disorder caused by mutations in the MCPH1 gene, whose product MCPH1 (also known as microcephalin and BRIT1) regulates DNA-damage response. Here we show that Mcph1 disruption in mice results in primary microcephaly, mimicking human MCPH1 symptoms, owing to a premature switching of neuroprogenitors from symmetric to asymmetric division. MCPH1-deficiency abrogates the localization of Chk1 to centrosomes, causing premature Cdk1 activation and early mitotic entry, which uncouples mitosis and the centrosome cycle. This misorients the mitotic spindle alignment and shifts the division plane of neuroprogenitors, to bias neurogenic cell fate. Silencing Cdc25b, a centrosome substrate of Chk1, corrects MCPH1-deficiency-induced spindle misalignment and rescues the premature neurogenic production in Mcph1-knockout neocortex. Thus, MCPH1, through its function in the Chk1–Cdc25–Cdk1 pathway to couple the centrosome cycle with mitosis, is required for precise mitotic spindle orientation and thereby regulates the progenitor division mode to maintain brain size.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Tang, B. L. Molecular genetic determinants of human brain size. Biochem. Biophys. Res. Commun. 345, 911–916 (2006).
Evans, P. D. et al. Microcephalin, a gene regulating brain size, continues to evolve adaptively in humans. Science 309, 1717–1720 (2005).
Wang, Y. Q. & Su, B. Molecular evolution of microcephalin, a gene determining human brain size. Hum. Mol. Genet. 13, 1131–1137 (2004).
Woods, C. G., Bond, J. & Enard, W. Autosomal recessive primary microcephaly (MCPH): a review of clinical, molecular, and evolutionary findings. Am. J.Hum. Genet. 76, 717–728 (2005).
Thornton, G. K. & Woods, C. G. Primary microcephaly: do all roads lead to Rome? Trends Genet. 25, 501–510 (2009).
Nicholas, A. K. et al. WDR62 is associated with the spindle pole and is mutated in human microcephaly. Nat. Genet. 42, 1010–1014 (2010).
Guernsey, D. L. et al. Mutations in centrosomal protein CEP152 in primary microcephaly families linked to MCPH4. Am. J. Hum. Genet. 87, 40–51 (2010).
Yu, T. W. et al. Mutations in WDR62, encoding a centrosome-associated protein, cause microcephaly with simplified gyri and abnormal cortical architecture. Nat. Genet. 42, 1015–1020 (2010).
Neitzel, H. et al. Premature chromosome condensation in humans associated with microcephaly and mental retardation: a novel autosomal recessive condition. Am. J. Hum. Genet. 70, 1015–1022 (2002).
Trimborn, M. et al. Mutations in microcephalin cause aberrant regulation of chromosome condensation. Am. J. Hum. Genet. 75, 261–266 (2004).
Wood, J.L., Liang, Y., Li, K. & Chen, J. Microcephalin/MCPH1 associates with the Condensin II complex to function in homologous recombination repair. J. Biol. Chem. 283, 29586–29592 (2008).
Trimborn, M., Schindler, D., Neitzel, H. & Hirano, T. Misregulated chromosome condensation in MCPH1 primary microcephaly is mediated by condensin II. Cell Cycle 5, 322–326 (2006).
Jeffers, L. J., Coull, B. J., Stack, S. J. & Morrison, C. G. Distinct BRCT domains in Mcph1/Brit1 mediate ionizing radiation-induced focus formation and centrosomal localization. Oncogene 27, 139–144 (2008).
Lin, S. Y., Rai, R., Li, K., Xu, Z. X. & Elledge, S. J. BRIT1/MCPH1 is a DNA damage responsive protein that regulates the Brca1-Chk1 pathway, implicating checkpoint dysfunction in microcephaly. Proc. Natl Acad. Sci. USA 102, 15105–15109 (2005).
Rai, R. et al. BRIT1 regulates early DNA damage response, chromosomal integrity, and cancer. Cancer Cell 10, 145–157 (2006).
Alderton, G. K. et al. Regulation of mitotic entry by microcephalin and its overlap with ATR signalling. Nat. Cell Biol. 8, 725–733 (2006).
Xu, X., Lee, J. & Stern, D. F. Microcephalin is a DNA damage response protein involved in regulation of CHK1 and BRCA1. J. Biol. Chem. 279, 34091–34094 (2004).
Peng, G. et al. BRIT1/MCPH1 links chromatin remodelling to DNA damage response. Nat. Cell Biol. 11, 865–872 (2009).
Yang, S. Z., Lin, F. T. & Lin, W. C. MCPH1/BRIT1 cooperates with E2F1 in the activation of checkpoint, DNA repair and apoptosis. EMBO Rep. 9, 907–915 (2008).
McKinnon, P. J. DNA repair deficiency and neurological disease. Nat. Rev. Neurosci. 10, 100–112 (2009).
Griffith, E. et al. Mutations in pericentrin cause Seckel syndrome with defective ATR-dependent DNA damage signaling. Nat. Genet. 40, 232–236 (2008).
Bond, J. & Woods, C. G. Cytoskeletal genes regulating brain size. Curr. Opin.Cell Biol. 18, 95–101 (2006).
Huttner, W. B. & Kosodo, Y. Symmetric versus asymmetric cell division during neurogenesis in the developing vertebrate central nervous system. Curr. Opin.Cell Biol. 17, 648–657 (2005).
Kriegstein, A., Noctor, S. & Martinez-Cerdeno, V. Patterns of neural stem and progenitor cell division may underlie evolutionary cortical expansion. Nat. Rev. Neurosci. 7, 883–890 (2006).
Jackson, A. P. et al. Identification of microcephalin, a protein implicated in determining the size of the human brain. Am. J. Hum. Genet. 71, 136–142 (2002).
Liang, Y. et al. BRIT1/MCPH1 is essential for mitotic and meiotic recombinationDNA repair and maintaining genomic stability in mice. PLoS Genet. 6, e1000826 (2010).
Knoblich, J. A. Mechanisms of asymmetric stem cell division. Cell 132, 583–597 (2008).
Zhong, W. & Chia, W. Neurogenesis and asymmetric cell division. Curr. Opin. Neurobiol. 18, 4–11 (2008).
Fish, J. L., Dehay, C., Kennedy, H. & Huttner, W. B. Making bigger brains-the evolution of neural-progenitor-cell division. J. Cell Sci. 121, 2783–2793 (2008).
Buchman, J. J. & Tsai, L. H. Spindle regulation in neural precursors of flies and mammals. Nat. Rev. Neurosci. 8, 89–100 (2007).
Kosodo, Y. et al. Asymmetric distribution of the apical plasma membrane during neurogenic divisions of mammalian neuroepithelial cells. EMBO J. 23, 2314–2324 (2004).
Wang, X. et al. Asymmetric centrosome inheritance maintains neural progenitors in the neocortex. Nature 461, 947–955 (2009).
Higginbotham, H. R. & Gleeson, J. G. The centrosome in neuronal development. Trends Neurosci. 30, 276–283 (2007).
Doxsey, S., Zimmerman, W. & Mikule, K. Centrosome control of the cell cycle. Trends Cell Biol. 15, 303–311 (2005).
Tibelius, A. et al. Microcephalin and pericentrin regulate mitotic entry via centrosome-associated Chk1. J. Cell Biol. 185, 1149–1157 (2009).
Kramer, A. et al. Centrosome-associated Chk1 prevents premature activation of cyclin-B-Cdk1 kinase. Nat. Cell Biol. 6, 884–891 (2004).
Nigg, E. A. & Raff, J. W. Centrioles, centrosomes, and cilia in health and disease. Cell 139, 663–678 (2009).
Rebollo, E. et al. Functionally unequal centrosomes drive spindle orientation in asymmetrically dividing Drosophila neural stem cells. Dev. Cell 12, 467–474 (2007).
Fukasawa, K. Oncogenes and tumour suppressors take on centrosomes. Nat. Rev. Cancer 7, 911–924 (2007).
Zachos, G. et al. Chk1 is required for spindle checkpoint function. Dev. Cell 12, 247–260 (2007).
Loffler, H. et al. Chk1-dependent regulation of Cdc25B functions to coordinate mitotic events. Cell Cycle 5, 2543–2547 (2006).
Chen, Y. & Poon, R. Y. The multiple checkpoint functions of CHK1 and CHK2 in maintenance of genome stability. Front. Biosci. 13, 5016–5029 (2008).
Roth, G. & Dicke, U. Evolution of the brain and intelligence. Trends Cogn. Sci. 9, 250–257 (2005).
Vitale, I., Galluzzi, L., Castedo, M. & Kroemer, G. Mitotic catastrophe: a mechanism for avoiding genomic instability. Nat. Rev. Mol. Cell Biol. 12, 385–392 (2011).
Castedo, M. et al. Cell death by mitotic catastrophe: a molecular definition. Oncogene 23, 2825–2837 (2004).
Rauch, A. et al. Mutations in the pericentrin (PCNT) gene cause primordial dwarfism. Science 319, 816–819 (2008).
Yingling, J. et al. Neuroepithelial stem cell proliferation requires LIS1 for precise spindle orientation and symmetric division. Cell 132, 474–486 (2008).
Lizarraga, S. B. et al. Cdk5rap2 regulates centrosome function and chromosome segregation in neuronal progenitors. Development 137, 1907–1917 (2010).
Fish, J. L., Kosodo, Y., Enard, W., Paabo, S. & Huttner, W. B. Aspm specifically maintains symmetric proliferative divisions of neuroepithelial cells. Proc. Natl Acad. Sci. USA 103, 10438–10443 (2006).
Pulvers, J. N. et al. Mutations in mouse Aspm (abnormal spindle-like microcephaly associated) cause not only microcephaly but also major defects in the germline. Proc. Natl Acad. Sci. USA 107, 16595–16600 (2010).
Wakefield, J. G., Bonaccorsi, S. & Gatti, M. The Drosophila protein asp is involved in microtubule organization during spindle formation and cytokinesis. J. Cell Biol. 153, 637–648 (2001).
Lucas, E. P. & Raff, J. W. Maintaining the proper connection between the centrioles and the pericentriolar matrix requires Drosophila centrosomin. J. Cell Biol. 178, 725–732 (2007).
Basto, R. et al. Flies without centrioles. Cell 125, 1375–1386 (2006).
Brunk, K. et al. Microcephalin coordinates mitosis in the syncytial Drosophila embryo. J. Cell Sci. 120, 3578–3588 (2007).
Buchman, J. J. et al. Cdk5rap2 interacts with pericentrin to maintain the neural progenitor pool in the developing neocortex. Neuron 66, 386–402 (2010).
Trimborn, M. et al. Establishment of a mouse model with misregulated chromosome condensation due to defective Mcph1 function. PLoS ONE 5, e9242 (2010).
Gonczy, P. Mechanisms of asymmetric cell division: flies and worms pave the way. Nat. Rev. Mol. Cell Biol. 9, 355–366 (2008).
Kosodo, Y. et al. Cytokinesis of neuroepithelial cells can divide their basal process before anaphase. EMBO J. 27, 3151–3163 (2008).
Zhou, Z. et al. PRMT5 regulates Golgi apparatus structure through methylation of the golgin GM130. Cell Res. 20, 1023–1033 (2010).
Arai, Y. et al. Neural stem and progenitor cells shorten S-phase on commitment to neuron production. Nat. Commun. 2, 154.
Nowakowski, R. S., Lewin, S. B. & Miller, M. W. Bromodeoxyuridine immunohistochemical determination of the lengths of the cell cycle and the DNA-synthetic phase for an anatomically defined population. J. Neurocytol. 18, 311–318 (1989).
Saito, T. In vivo electroporation in the embryonic mouse central nervous system. Nat. Protoc. 1, 1552–1558 (2006).
Frappart, P. O. et al. An essential function for NBS1 in the prevention of ataxia and cerebellar defects. Nat. Med. 11, 538–544 (2005).
Akhter, S. et al. Deficiency in SNM1 abolishes an early mitotic checkpoint induced by spindle stress. Mol. Cell. Biol. 24, 10448–10455 (2004).
Acknowledgements
We thank L. Frappart for the advice and the discussion on histological analysis and Z. Xu for the introduction into the in utero electroporation technology. We are grateful to M. Baldauf for his histological assistance, M. Welzel for her technical support and D. Galendo, C. Birch-Hirschfeld and C. Mueller for their assistance in the maintenance of the animal colonies. We are also grateful to M. Trajkovic-Arsic for the advice on the in situ hybridization techniques and A. Gompf for the FACS. We also would like to thank P. Herrlich, H. Heuer, T. Li and Y. Yang for their critical reading of the manuscript. Further thanks go to E. Stoeckl for editing the manuscript. We are also grateful to many other members of our laboratory for the helpful discussions. R.G., M.S. and Z.W.Z. are supported by a fellowship of the Leibniz Graduate School on Ageing (LGSA) Programme. P-O.F. is supported by an Emmy Noether Grant from the Deutschen Forschungsgemeinschaft (DFG). Z-Q.W. is supported in part by the DFG, Germany.
Author information
Authors and Affiliations
Contributions
R.G. carried out most of the experiments, analysed data and prepared the figures and the manuscript; Z.W.Z. carried out in situ hybridization, in utero electroporation, immunoblot analysis and analysed data; M.S. carried out gene targeting in embryonic stem cells and analysed gene expression; P-O.F. assisted with neurosphere experiments; T.J. contributed to mouse colony maintenance; Z-Q.W. designed experiments, analysed data and composed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1188 kb)
Rights and permissions
About this article
Cite this article
Gruber, R., Zhou, Z., Sukchev, M. et al. MCPH1 regulates the neuroprogenitor division mode by coupling the centrosomal cycle with mitotic entry through the Chk1–Cdc25 pathway. Nat Cell Biol 13, 1325–1334 (2011). https://doi.org/10.1038/ncb2342
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2342
This article is cited by
-
Aging-induced MCPH1 translocation activates necroptosis and impairs hematopoietic stem cell function
Nature Aging (2024)
-
Genetics of human brain development
Nature Reviews Genetics (2024)
-
GID complex regulates the differentiation of neural stem cells by destabilizing TET2
Frontiers of Medicine (2023)
-
An updated view on the centrosome as a cell cycle regulator
Cell Division (2022)
-
Endosomal trafficking defects alter neural progenitor proliferation and cause microcephaly
Nature Communications (2022)