Abstract
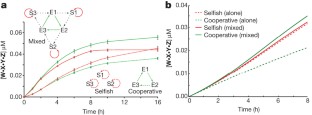
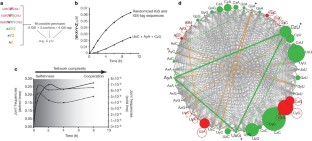
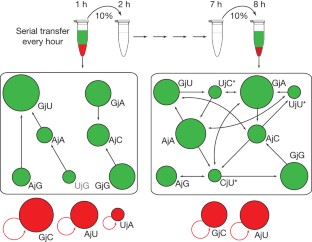
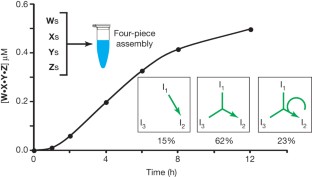
The origins of life on Earth required the establishment of self-replicating chemical systems capable of maintaining and evolving biological information. In an RNA world, single self-replicating RNAs would have faced the extreme challenge of possessing a mutation rate low enough both to sustain their own information and to compete successfully against molecular parasites with limited evolvability. Thus theoretical analyses suggest that networks of interacting molecules were more likely to develop and sustain life-like behaviour. Here we show that mixtures of RNA fragments that self-assemble into self-replicating ribozymes spontaneously form cooperative catalytic cycles and networks. We find that a specific three-membered network has highly cooperative growth dynamics. When such cooperative networks are competed directly against selfish autocatalytic cycles, the former grow faster, indicating an intrinsic ability of RNA populations to evolve greater complexity through cooperation. We can observe the evolvability of networks through in vitro selection. Our experiments highlight the advantages of cooperative behaviour even at the molecular stages of nascent life.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
Change history
31 October 2012
A minor typo in Fig. 1 was corrected.
References
Joyce, G. F. RNA evolution and the origins of life. Nature 338, 217–224 (1989)
Zaher, H. S. & Unrau, P. J. Selection of an improved RNA polymerase ribozyme with superior extension and fidelity. RNA 13, 1017–1026 (2007)
Wochner, A., Attwater, J., Coulson, A. & Holliger, P. Ribozyme-catalyzed transcription of an active ribozyme. Science 332, 209–212 (2011)
Eigen, M. & Schuster, P. The hypercycle. A principle of natural self-organization. Part A: emergence of the hypercycle. Naturwissenschaften 64, 541–565 (1977)
Kauffman, S. A. The Origins of Order: Self-Organization and Selection in Evolution (Oxford Univ. Press, 1993)
Sievers, D. & von Kiedrowski, G. Self-replication of complementary nucleotide-based oligomers. Nature 369, 221–224 (1994)
Levy, M. & Ellington, A. D. The descent of polymerization. Nature Struct. Biol. 8, 580–582 (2001)
Szathmáry, E. The origin of replicators and reproducers. Phil. Trans. Royal Soc. B 361, 1761–1776 (2006)
Nowak, M. A. & Ohtsuki, H. Preevolutionary dynamics and the origin of evolution. Proc. Natl Acad. Sci. USA 105, 14924–14927 (2008)
Eigen, M. Selforganization of matter and the evolution of biological macromolecules. Naturwissenschaften 58, 465–523 (1971)
Maynard Smith, J. Hypercycles and the origin of life. Nature 280, 445–446 (1979)
Kun, Á., Santos, M. & Szathmáry, E. Real ribozymes suggest a relaxed error threshold. Nature Genet. 37, 1008–1011 (2005)
Doudna, J. A. & Szostak, J. W. RNA-catalysed synthesis of complementary-strand RNA. Nature 339, 519–522 (1989)
Joyce, G. F. Forty years of in vitro evolution. Angew. Chem. Int. Ed. 46, 6420–6436 (2007)
Lincoln, T. A. & Joyce, G. F. Self-sustained replication of an RNA enzyme. Science 323, 1229–1232 (2009)
Levy, M. & Ellington, A. D. Exponential growth by cross-catalytic cleavage of deoxyribozymogens. Proc. Natl Acad. Sci. USA 100, 6416–6421 (2003)
Lee, D. H., Severin, K., Yokobayashi, Y. & Ghadiri, M. R. Emergence of symbiosis in peptide self-replication through a hypercyclic network. Nature 390, 591–594 (1997)
Lee, D. H., Severin, K. & Ghadiri, M. R. Autocatalytic networks: the transition from molecular self-replication to molecular ecosystems. Curr. Opin. Chem. Biol. 1, 491–496 (1997)
Voytek, S. B. & Joyce, G. F. Niche partitioning in the coevolution of 2 distinct RNA enzymes. Proc. Natl Acad. Sci. USA 106, 7780–7785 (2009)
Kim, D.-E. & Joyce, G. F. Cross-catalytic replication of an RNA ligase ribozyme. Chem. Biol. 11, 1505–1512 (2004)
Eigen, M. & Schuster, P. The hypercycle. A principle of natural self-organization. Part C: the realistic hypercycle. Naturwissenschaften 65, 341–369 (1978)
Reinhold-Hurek, B. & Shub, D. A. Self-splicing introns in tRNA genes of widely divergent bacteria. Nature 357, 173–176 (1992)
Hayden, E. J. & Lehman, N. Self-assembly of a group I intron from inactive oligonucleotide fragments. Chem. Biol. 13, 909–918 (2006)
Hayden, E. J., von Kiedrowski, G. & Lehman, N. Systems chemistry on ribozyme self-construction: evidence for anabolic autocatalysis in a recombination network. Angew. Chem. Int. Ed. 47, 8424–8428 (2008)
Draper, W. E., Hayden, E. J. & Lehman, N. Mechanisms of covalent self-assembly of the Azoarcus ribozyme from four fragment oligonucleotides. Nucleic Acids Res. 36, 520–531 (2008)
Nowak, M. A. Evolutionary Dynamics: Exploring the Equations of Life (Harvard Univ. Press, 2006)
Hordijk, W. & Steel, M. Detecting autocatalytic, self-containing sets in chemical reaction systems. J. Theor. Biol. 227, 451–461 (2004)
Weiner, A. M. & Maizels, N. 3′ terminal tRNA-like structures tag genomic RNA molecules for replication: implications for the origin of protein synthesis. Proc. Natl Acad. Sci. USA 84, 7383–7387 (1987)
Boerlijst, M. C. & Hogeweg, P. Spiral wave structures in prebiotic evolution: hypercycles stable against parasites. Physica D 48, 17–28 (1991)
Szathmáry, E. & Demeter, L. Group selection of early replicators and the origin of life. J. Theor. Biol. 128, 463–486 (1987)
Lynch, M., Burger, R., Butcher, D. & Gabriel, W. The mutational meltdown in asexual populations. J. Hered. 84, 339–344 (1993)
Eigen, M. & Schuster, P. The hypercycle. A principle of natural self-organization. Part B: the abstract hypercycle. Naturwissenschaften 65, 7–41 (1978)
Acknowledgements
We would like to thank A. Burton, R. Ghadiri, P. Higgs, B. Larson, K. Chacón and A. López García de Lomana for help during preparation of this manuscript. This work was supported by NASA grant NNX10AR15G to N.L., the Center for Life in Extreme Environments at Portland State University, and a fellowship from the Human Frontier Science Program Organization to R.X.-B.
Author information
Authors and Affiliations
Contributions
N.L. and N.V. conceived the basic idea and conducted the experiments; E.J.H. and I.A.C. contributed to the evaluation of the results; I.A.C., M.L.M. and R.X.-B. constructed the mathematical models; N.L. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Methods, Supplementary Text and Data, a Supplementary Discussion, Supplementary Figures 1-14, Supplementary Tables 1-3 and additional references. (PDF 4848 kb)
Rights and permissions
About this article
Cite this article
Vaidya, N., Manapat, M., Chen, I. et al. Spontaneous network formation among cooperative RNA replicators. Nature 491, 72–77 (2012). https://doi.org/10.1038/nature11549
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11549
This article is cited by
-
Quasispecies productivity
The Science of Nature (2024)
-
Multispecies autocatalytic RNA reaction networks in coacervates
Communications Chemistry (2023)
-
Self-organization of primitive metabolic cycles due to non-reciprocal interactions
Nature Communications (2023)
-
Emergence of autocatalytic sets in a simple model of technological evolution
Journal of Evolutionary Economics (2023)
-
Experimental exploration of a ribozyme neutral network using evolutionary algorithm and deep learning
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.