Abstract
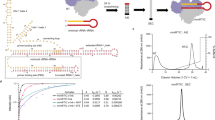
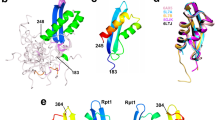
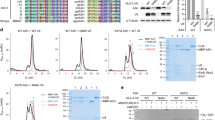
Regulation of the expression of the human immunodeficiency virus (HIV) genome is accomplished in large part by controlling transcription elongation. The viral protein Tat hijacks the host cell’s RNA polymerase II elongation control machinery through interaction with the positive transcription elongation factor, P-TEFb, and directs the factor to promote productive elongation of HIV mRNA. Here we describe the crystal structure of the Tat·P-TEFb complex containing HIV-1 Tat, human Cdk9 (also known as CDK9), and human cyclin T1 (also known as CCNT1). Tat adopts a structure complementary to the surface of P-TEFb and makes extensive contacts, mainly with the cyclin T1 subunit of P-TEFb, but also with the T-loop of the Cdk9 subunit. The structure provides a plausible explanation for the tolerance of Tat to sequence variations at certain sites. Importantly, Tat induces significant conformational changes in P-TEFb. This finding lays a foundation for the design of compounds that would specifically inhibit the Tat·P-TEFb complex and block HIV replication.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Onafuwa-Nuga, A. & Telesnitsky, A. The remarkable frequency of human immunodeficiency virus type 1 genetic recombination. Microbiol. Mol. Biol. Rev. 73, 451–80 (2009)
Letvin, N. L. Virology. Moving forward in HIV vaccine development. Science 326, 1196–1198 (2009)
Zhu, Y. et al. Transcription elongation factor P-TEFb is required for HIV-1 tat transactivation in vitro . Genes Dev. 11, 2622–2632 (1997)
Mancebo, H. S. et al. P-TEFb kinase is required for HIV Tat transcriptional activation in vivo and in vitro . Genes Dev. 11, 2633–2644 (1997)
Garber, M. E., Wei, P. & Jones, K. A. HIV-1 Tat interacts with cyclin T1 to direct the P-TEFb CTD kinase complex to TAR RNA. Cold Spring Harb. Symp. Quant. Biol. 63, 371–380 (1998)
Peterlin, B. M. & Price, D. H. Controlling the elongation phase of transcription with P-TEFb. Mol. Cell 23, 297–305 (2006)
Peng, J., Zhu, Y., Milton, J. T. & Price, D. H. Identification of multiple cyclin subunits of human P-TEFb. Genes Dev. 12, 755–762 (1998)
Marshall, N. F., Peng, J., Xie, Z. & Price, D. H. Control of RNA polymerase II elongation potential by a novel carboxyl-terminal domain kinase. J. Biol. Chem. 271, 27176–27183 (1996)
Fu, T. J., Peng, J., Lee, G., Price, D. H. & Flores, O. Cyclin K functions as a CDK9 regulatory subunit and participates in RNA polymerase II transcription. J. Biol. Chem. 274, 34527–34530 (1999)
Chao, S. H. & Price, D. H. Flavopiridol inactivates P-TEFb and blocks most RNA polymerase II transcription in vivo . J. Biol. Chem. 276, 31793–31799 (2001)
Zhou, Q. & Yik, J. H. The Yin and Yang of P-TEFb regulation: implications for human immunodeficiency virus gene expression and global control of cell growth and differentiation. Microbiol. Mol. Biol. Rev. 70, 646–659 (2006)
Michels, A. A. & Bensaude, O. RNA-driven cyclin-dependent kinase regulation: when CDK9/cyclin T subunits of P-TEFb meet their ribonucleoprotein partners. Biotechnol. J. 3, 1022–1032 (2008)
Sedore, S. C. et al. Manipulation of P-TEFb control machinery by HIV: recruitment of P-TEFb from the large form by Tat and binding of HEXIM1 to TAR. Nucleic Acids Res. 35, 4347–4358 (2007)
Wimmer, J. et al. Interactions between Tat and TAR and human immunodeficiency virus replication are facilitated by human cyclin T1 but not cyclins T2a or T2b. Virology 255, 182–189 (1999)
Bieniasz, P. D., Grdina, T. A., Bogerd, H. P. & Cullen, B. R. Recruitment of a protein complex containing Tat and cyclin T1 to TAR governs the species specificity of HIV-1 Tat. EMBO J. 17, 7056–7065 (1998)
Fujinaga, K. et al. The ability of positive transcription elongation factor B to transactivate human immunodeficiency virus transcription depends on a functional kinase domain, cyclin T1, and Tat. J. Virol. 72, 7154–7159 (1998)
Zhou, M. et al. Tat modifies the activity of CDK9 to phosphorylate serine 5 of the RNA polymerase II carboxyl-terminal domain during human immunodeficiency virus type 1 transcription. Mol. Cell. Biol. 20, 5077–5086 (2000)
Pavletich, N. P. Mechanisms of cyclin-dependent kinase regulation: structures of Cdks, their cyclin activators, and Cip and INK4 inhibitors. J. Mol. Biol. 287, 821–828 (1999)
Garber, M. E. et al. The interaction between HIV-1 Tat and human cyclin T1 requires zinc and a critical cysteine residue that is not conserved in the murine CycT1 protein. Genes Dev. 12, 3512–3527 (1998)
Janin, J. Specific versus non-specific contacts in protein crystals. Nature Struct. Biol. 4, 973–974 (1997)
Baumli, S. et al. The structure of P-TEFb (CDK9/cyclin T1), its complex with flavopiridol and regulation by phosphorylation. EMBO J. 27, 1907–1918 (2008)
Kuppuswamy, M., Subramanian, T., Srinivasan, A. & Chinnadurai, G. Multiple functional domains of Tat, the trans-activator of HIV-1, defined by mutational analysis. Nucleic Acids Res. 17, 3551–3561 (1989)
Sadaie, M. R., Mukhopadhyaya, R., Benaissa, Z. N., Pavlakis, G. N. & Wong-Staal, F. Conservative mutations in the putative metal-binding region of human immunodeficiency virus Tat disrupt virus replication. AIDS Res. Hum. Retroviruses 6, 1257–1263 (1990)
Ruben, S. et al. Structural and functional characterization of human immunodeficiency virus tat protein. J. Virol. 63, 1–8 (1989)
Jeang, K. T., Xiao, H. & Rich, E. A. Multifaceted activities of the HIV-1 transactivator of transcription, Tat. J. Biol. Chem. 274, 28837–28840 (1999)
Rice, A. P. & Carlotti, F. Mutational analysis of the conserved cysteine-rich region of the human immunodeficiency virus type 1 Tat protein. J. Virol. 64, 1864–1868 (1990)
Anand, K., Schulte, A., Vogel-Bachmayr, K., Scheffzek, K. & Geyer, M. Structural insights into the cyclin T1-Tat-TAR RNA transcription activation complex from EIAV. Nature Struct. Mol. Biol. 15, 1287–1292 (2008)
Taube, R. et al. Interactions between equine cyclin T1, Tat, and TAR are disrupted by a leucine-to-valine substitution found in human cyclin T1. J. Virol. 74, 892–898 (2000)
Derse, D. & Newbold, S. H. Mutagenesis of EIAV TAT reveals structural features essential for transcriptional activation and TAR element recognition. Virology 194, 530–536 (1993)
Ippolito, J. A. & Steitz, T. A. A 1.3-Å resolution crystal structure of the HIV-1 trans-activation response region RNA stem reveals a metal ion-dependent bulge conformation. Proc. Natl Acad. Sci. USA 95, 9819–9824 (1998)
Richter, S., Ping, Y. H. & Rana, T. M. TAR RNA loop: a scaffold for the assembly of a regulatory switch in HIV replication. Proc. Natl Acad. Sci. USA 99, 7928–7933 (2002)
Campbell, G. R. & Loret, E. P. What does the structure-function relationship of the HIV-1 Tat protein teach us about developing an AIDS vaccine? Retrovirology 6, 50 (2009)
Hemelaar, J., Gouws, E., Ghys, P. D. & Osmanov, S. Global and regional distribution of HIV-1 genetic subtypes and recombinants in 2004. AIDS 20, W13–W23 (2006)
Crooks, G. E., Hon, G., Chandonia, J. M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004)
Kuiken, C. et al. HIV Sequence Compendium 2009, LA-UR 09–03280 (Theoretical Biology and Biophysics Group, Los Alamos National Laboratory, USA). (2009)
D’Orso, I. & Frankel, A. D. Tat acetylation modulates assembly of a viral-host RNA-protein transcription complex. Proc. Natl Acad. Sci. USA 106, 3101–3106 (2009)
Barboric, M. et al. Tat competes with HEXIM1 to increase the active pool of P-TEFb for HIV-1 transcription. Nucleic Acids Res. 35, 2003–2012 (2007)
Michels, A. A. et al. MAQ1 and 7SK RNA interact with CDK9/cyclin T complexes in a transcription-dependent manner. Mol. Cell. Biol. 23, 4859–4869 (2003)
Yik, J. H. et al. Inhibition of P-TEFb (CDK9/Cyclin T) kinase and RNA polymerase II transcription by the coordinated actions of HEXIM1 and 7SK snRNA. Mol. Cell 12, 971–982 (2003)
Russo, A. A., Jeffrey, P. D., Patten, A. K., Massague, J. & Pavletich, N. P. Crystal structure of the p27Kip1 cyclin-dependent-kinase inhibitor bound to the cyclin A-Cdk2 complex. Nature 382, 325–331 (1996)
Li, Q. et al. Analysis of the large inactive P-TEFb complex indicates that it contains one 7SK molecule, a dimer of HEXIM1 or HEXIM2, and two P-TEFb molecules containing Cdk9 phosphorylated at threonine 186. J. Biol. Chem. 280, 28819–28826 (2005)
Chen, R., Yang, Z. & Zhou, Q. Phosphorylated positive transcription elongation factor b (P-TEFb) is tagged for inhibition through association with 7SK snRNA. J. Biol. Chem. 279, 4153–4160 (2004)
Wang, Y., Liu, X. Y. & De Clercq, E. Role of the HIV-1 positive elongation factor P-TEFb and inhibitors thereof. Mini Rev. Med. Chem. 9, 379–385 (2009)
Jancarik, J., Pufan, R., Hong, C., Kim, S. H. & Kim, R. Optimum solubility (OS) screening: an efficient method to optimize buffer conditions for homogeneity and crystallization of proteins. Acta Crystallogr. D 60, 1670–1673 (2004)
Tahirov, T. H. et al. High-resolution crystals of methionine aminopeptidase from Pyrococcus furiosus obtained by water-mediated transformation. J. Struct. Biol. 121, 68–72 (1998)
Otwinowski, Z. & Minor, W. Processing of X-ray Diffraction Data Collected in Oscillation Mode (eds. Carter, J. C. W. & Sweet, R. M.). (1997)
Brünger, A. T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Vassylyev, D. G., Vassylyeva, M. N., Perederina, A., Tahirov, T. H. & Artsimovitch, I. Structural basis for transcription elongation by bacterial RNA polymerase. Nature 448, 157–162 (2007)
Acknowledgements
We thank J. Lovelace and G. E. Borgstahl for maintenance and management of the Eppley Institute’s X-ray Crystallography facility; D. G. Vassylyev for the zonal scaling instruction files. This work is supported by the NIH grants GM35500 and AI074392 to D.H.P., by Nebraska Department of Health and Human Services grant LB506 and in part by NIH grant GM082923 to T.H.T. This work is also based on research conducted at the Advanced Photon Source on the Northeastern Collaborative Access Team beamlines, which are supported by award RR-15301 from the National Center for Research Resources at the National Institutes of Health. Use of the Advanced Photon Source is supported by the US Department of Energy, Office of Basic Energy Sciences, under Contract No. DE-AC02-06CH11357. The Eppley Institute’s X-ray Crystallography facility is supported by Cancer Center Support Grant P30CA036727.
Author information
Authors and Affiliations
Contributions
T.H.T. managed the crystallization and structure determination part of the project, solved the crystal structures and prepared the manuscript. N.D.B. obtained the crystals. Diffraction data collection was performed by N.D.B. and T.H.T. Protein cloning, expression, purification and writing the corresponding methods sections were performed by S.C.S., K.V. and J.J.C., respectively. D.H.P. managed the protein production part of the project, and helped generate and edit the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-12 with legends, Supplementary Tables 1-2 and References. (PDF 14705 kb)
Rights and permissions
About this article
Cite this article
Tahirov, T., Babayeva, N., Varzavand, K. et al. Crystal structure of HIV-1 Tat complexed with human P-TEFb. Nature 465, 747–751 (2010). https://doi.org/10.1038/nature09131
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature09131
This article is cited by
-
Live-cell imaging reveals the spatiotemporal organization of endogenous RNA polymerase II phosphorylation at a single gene
Nature Communications (2021)
-
CDK9 keeps RNA polymerase II on track
Cellular and Molecular Life Sciences (2021)
-
Differences in Transcriptional Dynamics Between T-cells and Macrophages as Determined by a Three-State Mathematical Model
Scientific Reports (2020)
-
Regulation of cyclin T1 during HIV replication and latency establishment in human memory CD4 T cells
Virology Journal (2019)
-
Design of Tat-Activated Cdk9 Inhibitor
International Journal of Peptide Research and Therapeutics (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.