Abstract
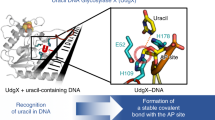
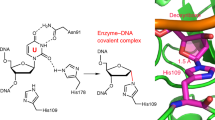
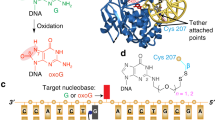
The genomes of aerobic organisms suffer chronic oxidation of guanine to the genotoxic product 8-oxoguanine (oxoG)1. Replicative DNA polymerases misread oxoG residues and insert adenine instead of cytosine opposite the oxidized base. Both bases in the resulting A·oxoG mispair are mutagenic lesions, and both must undergo base-specific replacement to restore the original C·G pair. Doing so represents a formidable challenge to the DNA repair machinery, because adenine makes up roughly 25% of the bases in most genomes. The evolutionarily conserved enzyme adenine DNA glycosylase (called MutY in bacteria and hMYH in humans) initiates repair of A·oxoG to C·G by removing the inappropriately paired adenine base from the DNA backbone. A central issue concerning MutY function is the mechanism by which A·oxoG mispairs are targeted among the vast excess of A·T pairs. Here we report the use of disulphide crosslinking2 to obtain high-resolution crystal structures of MutY–DNA lesion-recognition complexes. These structures reveal the basis for recognizing both lesions in the A·oxoG pair and for catalysing removal of the adenine base.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lindahl, T. Instability and decay of the primary structure of DNA. Nature 362, 709–715 (1993)
Norman, D. P. G. & Verdine, G. L. Covalent trapping of protein–DNA complexes. Annu. Rev. Biochem. 72, 337–366 (2003)
Gogos, A., Cillo, J., Clarke, N. D. & Lu, A. L. Specific recognition of A/G and A/7,8-dihydro-8-oxoguanine (8-oxoG) mismatches by Escherichia coli MutY: removal of the C-terminal domain preferentially affects A/8-oxoG recognition. Biochemistry 35, 16665–16671 (1996)
Noll, D. M., Gogos, A., Granek, J. A. & Clarke, N. D. The C-terminal domain of the adenine-DNA glycosylase MutY confers specificity for 8-oxoguanine·adenine mispairs and may have evolved from MutT, an 8-oxo-dGTPase. Biochemistry 38, 6374–6379 (1999)
Guan, Y. et al. MutY catalytic core, mutant and bound adenine structures define specificity for DNA repair enzyme superfamily. Nature Struct. Biol. 5, 1058–1064 (1998)
Volk, D. E. et al. Structural similarities between MutT and the C-terminal domain of MutY. Biochemistry 39, 7331–7336 (2000)
Hollis, T., Ichikawa, Y. & Ellenberger, T. DNA bending and a flip-out mechanism for base excision by the helix–hairpin–helix DNA glycosylase. Escherichia coli AlkA. EMBO J. 19, 758–766 (2000)
Bruner, S. D., Norman, D. P. & Verdine, G. L. Structural basis for recognition and repair of the endogenous mutagen 8-oxoguanine in DNA. Nature 403, 859–866 (2000)
Fromme, J. C. & Verdine, G. L. Structure of a trapped endonuclease III–DNA covalent intermediate. EMBO J. 22, 3461–3471 (2003)
Bernards, A. S., Miller, J. K., Bao, K. K. & Wong, I. Flipping duplex DNA inside out: a double base-flipping reaction mechanism by Escherichia coli MutY adenine glycosylase. J. Biol. Chem. 277, 20960–20964 (2002)
Kouchakdjian, M. et al. NMR structural studies of the ionizing radiation adduct 7-hydro-8-oxodeoxyguanosine (8-oxo-7H-dG) opposite deoxyadenosine in a DNA duplex. 8-Oxo-7H-dG(syn)·dA(anti) alignment at lesion site. Biochemistry 30, 1403–1412 (1991)
McAuley-Hecht, K. E. et al. Crystal structure of a DNA duplex containing 8-hydroxydeoxyguanine–adenine base pairs. Biochemistry 33, 10266–10270 (1994)
Lin, J., Abeygunawardana, C., Frick, D. N., Bessman, M. J. & Mildvan, A. S. Solution structure of the quaternary MutT–M2+–AMPCPP–M2+ complex and mechanism of its pyrophosphohydrolase action. Biochemistry 36, 1199–1211 (1997)
Massiah, M. A., Saraswat, V., Azurmendi, H. F. & Mildvan, A. S. Solution structure and NH exchange studies of the MutT pyrophosphohydrolase complexed with Mg2+ and 8-oxo-dGMP, a tightly bound product. Biochemistry 42, 10140–10154 (2003)
Plum, G. E., Grollman, A. P., Johnson, F. & Breslauer, K. J. Influence of the oxidatively damaged adduct 8-oxodeoxyguanosine on the conformation, energetics, and thermodynamic stability of a DNA duplex. Biochemistry 34, 16148–16160 (1995)
Porello, S. L., Leyes, A. E. & David, S. S. Single-turnover and pre-steady-state kinetics of the reaction of the adenine glycosylase MutY with mismatch-containing DNA substrates. Biochemistry 37, 14756–14764 (1998)
Werner, R. M. & Stivers, J. T. Kinetic isotope effect studies of the reaction catalyzed by uracil DNA glycosylase: evidence for an oxocarbenium ion–uracil anion intermediate. Biochemistry 39, 14054–14064 (2000)
Dinner, A. R., Blackburn, G. M. & Karplus, M. Uracil-DNA glycosylase acts by substrate autocatalysis. Nature 413, 752–755 (2001)
Porello, S. L., Williams, S. D., Kuhn, H., Michaels, M. L. & David, S. S. Specific recognition of substrate analogs by the DNA mismatch repair enzyme MutY. J. Am. Chem. Soc. 118, 10684–10692 (1996)
Al-Tassan, N. et al. Inherited variants of MYH associated with somatic G:C → T:A mutations in colorectal tumors. Nature Genet. 30, 227–232 (2002)
Jones, S. et al. Biallelic germline mutations in MYH predispose to multiple colorectal adenoma and somatic G:C → T:A mutations. Hum. Mol. Genet. 11, 2961–2967 (2002)
Sieber, O. M. et al. Multiple colorectal adenomas, classic adenomatous polyposis, and germ-line mutations in MYH. N. Engl. J. Med. 348, 791–799 (2003)
Chmiel, N. H., Livingston, A. L. & David, S. S. Insight into the functional consequences of inherited variants of the hMYH adenine glycosylase associated with colorectal cancer: complementation assays with hMYH variants and pre-steady-state kinetics of the corresponding mutated E. coli enzymes. J. Mol. Biol. 327, 431–443 (2003)
Nicholls, A., Sharp, K. A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins Struct. Funct. Genet. 11, 281–296 (1991)
Acknowledgements
We thank H. Nash for the reduced abasic phosphoramidite; Y. Korkhin for assistance with data collection and processing; M. Becker for beamline assistance; S. Bruner and J. J. Miranda for critically reading the manuscript; and Enanta Pharmaceuticals for use of their X-ray generator and detector. Some data for this study were measured at beamline X25 of the National Synchrotron Light Source; financial support for this beamline comes from the NIH and the United States Department of Energy.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
41586_2004_BFnature02306_MOESM3_ESM.pdf
Supplementary figure 3: MutY Sequence alignment showing the amino acid sequences from humans, the bacterium Escherichia coli (E.co.), the thermophilic bacterium Bacillus stearothermophilus (B.st., the ortholog used in this work), and the fission yeast Schizosaccharomyces pombe (S.po.). (PDF 93 kb)
Rights and permissions
About this article
Cite this article
Fromme, J., Banerjee, A., Huang, S. et al. Structural basis for removal of adenine mispaired with 8-oxoguanine by MutY adenine DNA glycosylase. Nature 427, 652–656 (2004). https://doi.org/10.1038/nature02306
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature02306
This article is cited by
-
Dynamic basis for dA•dGTP and dA•d8OGTP misincorporation via Hoogsteen base pairs
Nature Chemical Biology (2023)
-
8-Oxoguanine: from oxidative damage to epigenetic and epitranscriptional modification
Experimental & Molecular Medicine (2022)
-
The trajectory of intrahelical lesion recognition and extrusion by the human 8-oxoguanine DNA glycosylase
Nature Communications (2020)
-
A combined Far-FTIR, FTIR Spectromicroscopy, and DFT Study of the Effect of DNA Binding on the [4Fe4S] Cluster Site in EndoIII
Scientific Reports (2020)
-
Automated AFM analysis of DNA bending reveals initial lesion sensing strategies of DNA glycosylases
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.