Abstract
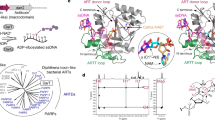
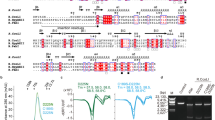
A uracil DNA glycosylase (UDG) from Mycobacterium smegmatis (MsmUdgX) shares sequence similarity with family 4 UDGs and forms exceedingly stable complexes with single-stranded uracil-containing DNAs (ssDNA-Us) that are resistant to denaturants. However, MsmUdgX has been reported to be inactive in excising uracil from ssDNA-Us and the underlying structural basis is unclear. Here, we report high-resolution crystal structures of MsmUdgX in the free, uracil- and DNA-bound forms, respectively. The structural information, supported by mutational and biochemical analyses, indicates that the conserved residue His109 located on a characteristic loop forms an irreversible covalent linkage with the deoxyribose at the apyrimidinic site of ssDNA-U, thus rendering the enzyme unable to regenerate. By proposing the catalytic pathway and molecular mechanism for MsmUdgX, our studies provide an insight into family 4 UDGs and UDGs in general.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Barnes, D. E. & Lindahl, T. Repair and genetic consequences of endogenous DNA base damage in mammalian cells. Annu. Rev. Genet. 38, 445–476 (2004).
Visnes, T. et al. Uracil in DNA and its processing by different DNA glycosylases. Philos. Trans. R. Soc. Lond. B 364, 563–568 (2009).
Lindahl, T., Ljungquist, S., Siegert, W., Nyberg, B. & Sperens, B. DNA N-glycosidases: properties of uracil-DNA glycosidase from Escherichia coli. J. Biol. Chem. 252, 3286–3294 (1977).
Li, J., Yang, Y., Guevara, J., Wang, L. & Cao, W. Identification of a prototypical single-stranded uracil DNA glycosylase from Listeria innocua. DNA Repair 57, 107–115 (2017).
Xia, B. et al. Correlated mutation in the evolution of catalysis in uracil DNA glycosylase superfamily. Sci. Rep. 7, 45978 (2017).
Xia, B. et al. Specificity and catalytic mechanism in family 5 uracil DNA glycosylase. J. Biol. Chem. 289, 18413–18426 (2014).
Parikh, S. S., Putnam, C. D. & Tainer, J. A. Lessons learned from structural results on uracil-DNA glycosylase. Mutat. Res. 460, 183–199 (2000).
Pearl, L. H. Structure and function in the uracil-DNA glycosylase superfamily. Mutat. Res. 460, 165–181 (2000).
Mol, C. D. et al. Crystal structure of human uracil-DNA glycosylase in complex with a protein inhibitor: protein mimicry of DNA. Cell 82, 701–708 (1995).
Slupphaug, G. et al. A nucleotide-flipping mechanism from the structure of human uracil-DNA glycosylase bound to DNA. Nature 384, 87–92 (1996).
Maiti, A., Morgan, M. T., Pozharski, E. & Drohat, A. C. Crystal structure of human thymine DNA glycosylase bound to DNA elucidates sequence-specific mismatch recognition. Proc. Natl Acad. Sci. USA 105, 8890–8895 (2008).
Zhang, L. et al. Thymine DNA glycosylase specifically recognizes 5-carboxylcytosine-modified DNA. Nat. Chem. Biol. 8, 328–330 (2012).
Wibley, J. E., Waters, T. R., Haushalter, K., Verdine, G. L. & Pearl, L. H. Structure and specificity of the vertebrate anti-mutator uracil-DNA glycosylase SMUG1. Mol. Cell 11, 1647–1659 (2003).
Hoseki, J. et al. Crystal structure of a family 4 uracil-DNA glycosylase from Thermus thermophilus HB8. J. Mol. Biol. 333, 515–526 (2003).
Kawai, A. et al. Crystal structure of family 4 uracil-DNA glycosylase from Sulfolobus tokodaii and a function of tyrosine 170 in DNA binding. FEBS Lett. 589, 2675–2682 (2015).
Kosaka, H., Hoseki, J., Nakagawa, N., Kuramitsu, S. & Masui, R. Crystal structure of family 5 uracil-DNA glycosylase bound to DNA. J. Mol. Biol. 373, 839–850 (2007).
Venkatesh, J., Kumar, P., Krishna, P. S., Manjunath, R. & Varshney, U. Importance of uracil DNA glycosylase in Pseudomonas aeruginosa and Mycobacterium smegmatis, G+C-rich bacteria, in mutation prevention, tolerance to acidified nitrite, and endurance in mouse macrophages. J. Biol. Chem. 278, 24350–24358 (2003).
Sang, P. B., Srinath, T., Patil, A. G., Woo, E. J. & Varshney, U. A unique uracil-DNA binding protein of the uracil DNA glycosylase superfamily. Nucleic Acids Res. 43, 8452–8463 (2015).
Saikrishnan, K. et al. Domain closure and action of uracil DNA glycosylase (UDG): structures of new crystal forms containing the Escherichia coli enzyme and a comparative study of the known structures involving UDG. Acta Crystallogr. D 58, 1269–1276 (2002).
Hashimoto, H., Zhang, X. & Cheng, X. D. Excision of thymine and 5-hydroxymethyluracil by the MBD4 DNA glycosylase domain: structural basis and implications for active DNA demethylation. Nucleic Acids Res. 40, 8276–8284 (2012).
Woods, R. D. et al. Structure and stereochemistry of the base excision repair glycosylase MutY reveal a mechanism similar to retaining glycosidases. Nucleic Acids Res. 44, 801–810 (2016).
Stivers, J. T. & Jiang, Y. L. A mechanistic perspective on the chemistry of DNA repair glycosylases. Chem. Rev. 103, 2729–2759 (2003).
Champoux, J. J. DNA topoisomerases: structure, function, and mechanism. Annu. Rev. Biochem. 70, 369–413 (2001).
Gao, Z., Maloney, D. J., Dedkova, L. M. & Hecht, S. M. Inhibitors of DNA polymerase beta: activity and mechanism. Bioorg. Med. Chem. 16, 4331–4340 (2008).
Gros, C. et al. DNA methylation inhibitors in cancer: recent and future approaches. Biochimie 94, 2280–2296 (2012).
Haracska, L., Prakash, L. & Prakash, S. A mechanism for the exclusion of low-fidelity human Y-family DNA polymerases from base excision repair. Genes Dev. 17, 2777–2785 (2003).
Hazra, T. K. et al. Oxidative DNA damage repair in mammalian cells: a new perspective. DNA Repair 6, 470–480 (2007).
Heidrun, I., Hong Jing, C. & Champoux, J. J. Human Tdp1 cleaves a broad spectrum of substrates, including phosphoamide linkages. J. Biol. Chem. 280, 36518 (2005).
Ide, H. & Kotera, M. Human DNA glycosylases involved in the repair of oxidatively damaged DNA. Biol. Pharm. Bull. 27, 480–485 (2004).
Khodyreva, S. N. et al. Apurinic/apyrimidinic (AP) site recognition by the 5'-dRP/AP lyase in poly(ADP-ribose) polymerase-1 (PARP-1). Proc. Natl Acad. Sci. USA 107, 22090–22095 (2010).
Steven, A. R. et al. Ku is a 5'-dRP/AP lyase that excises nucleotide damage near broken ends. Nature 464, 1214 (2010).
Tubbs, J. L., Pegg, A. E. & Tainer, J. A. DNA binding, nucleotide flipping, and the helix-turn-helix motif in base repair by O6-alkylguanine-DNA alkyltransferase and its implications for cancer chemotherapy. DNA Repair 6, 1100–1115 (2007).
Zhang, Z., Cheng, B. & Tse-Dinh, Y. C. Crystal structure of a covalent intermediate in DNA cleavage and rejoining by Escherichia coli DNA topoisomerase I. Proc Natl Acad. Sci. USA 108, 6939–6944 (2011).
Pluta, R. et al. Structural basis of a histidine-DNA nicking/joining mechanism for gene transfer and promiscuous spread of antibiotic resistance. Proc. Natl Acad. Sci. USA 114, E6526–E6535 (2017).
McCann, J. A. & Berti, P. J. Transition state analysis of acid-catalyzed dAMP hydrolysis. J. Am. Chem. Soc. 129, 7055–7064 (2007).
Vocadlo, D. J., Davies, G. J., Laine, R. & Withers, S. G. Catalysis by hen egg-white lysozyme proceeds via a covalent intermediate. Nature 412, 835–838 (2001).
Messick, T. E. et al. Noncysteinyl coordination to the [4Fe-4S]2+ cluster of the DNA repair adenine glycosylase MutY introduced via site-directed mutagenesis. Structural characterization of an unusual histidinyl-coordinated cluster. Biochemistry 41, 3931–3942 (2002).
Engstrom, L. M., Partington, O. A. & David, S. S. An iron–sulfur cluster loop motif in the archaeoglobus fulgidus uracil-dna glycosylase mediates efficient uracil recognition and removal. Biochemistry 51, 5187–5197 (2012).
Cole, S. T. et al. Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature 393, 537–544 (1998).
Heewook, L., Ellen, P., Haixu, T. & Foster, P. L. Rate and molecular spectrum of spontaneous mutations in the bacterium Escherichia coli as determined by whole-genome sequencing. Proc. Natl Acad. Sci. USA 109, 16416–16417 (2012).
Kunkel, T. A. & Erie, D. A. DNA mismatch repair. Annu. Rev. Biochem. 74, 681–710 (2005).
Sassetti, C. M. & Rubin, E. J. Genetic requirements for mycobacterial survival during infection. Proc. Natl Acad. Sci. USA 100, 12989–12994 (2003).
Kucukyildirim, S. et al. The rate and spectrum of spontaneous mutations in Mycobacterium smegmatis, a bacterium naturally devoid of the postreplicative mismatch repair pathway. G3 6, 2157–2163 (2016).
Malshetty, V. S., Jain, R., Srinath, T., Kurthkoti, K. & Varshney, U. Synergistic effects of UdgB and Ung in mutation prevention and protection against commonly encountered DNA damaging agents in Mycobacterium smegmatis. Microbiology 156, 940–949 (2010).
Wanner, R. M. et al. The uracil DNA glycosylase UdgB of Mycobacterium smegmatis protects the organism from the mutagenic effects of cytosine and adenine deamination. J. Bacteriol. 191, 6312–6319 (2009).
Van der Veen, S. & Tang, C. M. The BER necessities: the repair of DNA damage in human-adapted bacterial pathogens. Nat. Rev. Microbiol. 13, 83–94 (2015).
Minor, W., Cymborowski, M., Otwinowski, Z. & Chruszcz, M. HKL-3000: the integration of data reduction and structure solution-from diffraction images to an initial model in minutes. Acta Crystallogr. D 62, 859–866 (2010).
Mccoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of COOT. Acta Crystallogr. D 66, 486–501 (2010).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D 68, 352–367 (2012).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Painter, J. & Merritt, E. A. Optimal description of a protein structure in terms of multiple groups undergoing TLS motion. Acta Crystallogr. D 62, 439–450 (2006).
Winn, M. D., Isupov, M. N. & Murshudov, G. N. Use of TLS parameters to model anisotropic displacements in macromolecular refinement. Acta Crystallogr. D 57, 122–133 (2001).
Baker, N. A., Sept, D., Joseph, S., Holst, M. J. & McCammon, J. A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl Acad. Sci. USA 98, 10037–10041 (2001).
Stivala, A., Wybrow, M., Wirth, A., Whisstock, J. C. & Stuckey, P. J. Automatic generation of protein structure cartoons with Pro-Origami. Bioinformatics 27, 3315–3316 (2011).
Acknowledgements
We thank J. Cox (UCSF) for providing the M. smegmatis strain, the BL19U1 beamline at Shanghai Synchrotron Radiation Facility for assistance during diffraction data collection, and Y. Ju, H. Zhou and Q. Jia for assistance with data collection and analyses in TSAs.
Author information
Authors and Affiliations
Contributions
W.X. conceived and designed the research. J.T., R.C. and Y.Y. performed the research. W.X. analyzed the data and wrote the paper. W.C. provided input into the data analysis and manuscript preparation. All authors reviewed the manuscript. Funding for this work was provided by the National Natural Science Foundation of China (grant nos. 31870782 and 31700657), the Foundation of State Key Laboratory for Biocontrol (grant no. SKLBC16KF02), the Natural Science Foundation of Guangdong province (grant no. 2018A030313313) and the NIH (grant no. GM121997).
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Tables 1–3, Supplementary Figures 1–17
Rights and permissions
About this article
Cite this article
Tu, J., Chen, R., Yang, Y. et al. Suicide inactivation of the uracil DNA glycosylase UdgX by covalent complex formation. Nat Chem Biol 15, 615–622 (2019). https://doi.org/10.1038/s41589-019-0290-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-019-0290-x
This article is cited by
-
Molecular Biology Applications of Psychrophilic Enzymes: Adaptations, Advantages, Expression, and Prospective
Applied Biochemistry and Biotechnology (2024)
-
HMGN1 enhances CRISPR-directed dual-function A-to-G and C-to-G base editing
Nature Communications (2023)
-
Bacterial DNA excision repair pathways
Nature Reviews Microbiology (2022)
-
Efficient C•G-to-G•C base editors developed using CRISPRi screens, target-library analysis, and machine learning
Nature Biotechnology (2021)