Abstract
Assessment of programmed cell death ligand 1 (PD-L1) immunohistochemical staining is used for decision on treatment with programmed cell death 1 and PD-L1 checkpoint inhibitors in lung adenocarcinomas and squamous cell carcinomas. This study aimed to compare the staining properties of tumor cells between the antibody clones 28-8, 22C3, SP142, and SP263 and investigate interrater variation between pathologists to see if these stainings can be safely evaluated in the clinical setting. Using consecutive sections from a tissue microarray with tumor tissue from 55 resected lung cancer cases, staining with five PD-L1 assays (28-8 from two different vendors, 22C3, SP142, and SP263) was performed. Seven pathologists individually evaluated the percentage of positive tumor cells, scoring each sample applying cutoff levels used in clinical studies: <1% positive tumor cells (score 0), 1–4% (score 1), 5–9% (score 2), 10–24% (score 3), 25–49% (score 4), and >50% positive tumor cells (score 5). Pairwise analysis of antibody clones showed weighted kappa values in the range of 0.45–0.91 with the highest values for comparisons with 22C3 and 28-8 and the lowest involving SP142. Excluding SP142 resulted in kappa 0.75–0.91. Weighted kappa for interobserver variation between pathologists was 0.71–0.96. Up to 20% of the cases were differently classified as positive or negative by any pathologist compared with consensus score using ≥1% positive tumor cells as cutoff. A significantly better agreement between pathologists was seen using ≥50% as cutoff (0–5% of cases). In conclusion, the concordance between the PD-L1 antibodies 22C3, 28-8 and SP263 is relatively good when evaluating lung cancers and suggests that any one of these assays may be sufficient as basis for decision on treatment with nivolumab, pembrolizumab, and durvalumab. The scoring of the pathologist presents an intrinsic source of error that should be considered especially at low PD-L1 scores.
Similar content being viewed by others
Main
Checkpoint inhibitors targeting programmed cell death 1 (PD-1) and programmed cell death ligand 1 (PD-L1) proteins are the latest addition to the treatment arsenal in lung cancer. Clinical trials have shown better survival for patients treated with PD-1/PD-L1 inhibitors compared with chemotherapy, especially for squamous cell carcinomas and for adenocarcinomas positive for PD-L1 assessed with immunohistochemical staining.1, 2, 3, 4, 5, 6, 7, 8, 9, 10
In the clinical studies, various antibodies for immunohistochemical staining for PD-L1 have been used for the different drugs; clone 28-8 for nivolumab, 22C3 for pembrolizumab, SP142 for atezolizumab and SP263 for durvalumab. So far, nivolumab, pembrolizumab and atezolizumab have all been approved by the US Food and Drug Administration for treatment of advanced lung cancer including squamous cell carcinomas and adenocarcinomas, while the 28-8 and 22C3 pharmDx/Dako and SP142 and SP263 Ventana laboratory tests have been approved for immunohistochemical testing. In addition to different antibody clones, two different staining platforms and several different cutoff levels (1, 5, 10, 25, and 50%) for positive staining have been used in the studies. Also, for SP142 lymphocytes are evaluated in addition to tumor cells.
In the recently published Blueprint PD-L1 Immunohistochemistry Assay Comparison Project, 39 lung cancer cases were stained with the four PD-L1 immunohistochemistry assays as used in the clinical trials. All cases were evaluated independently by three vendor-associated expert pathologists. The conclusion was that despite similarities (especially between 28-8, 22C3, and SP263), interchanging assays and cutoffs will lead to ‘misclassification’ of cases.11
However, the use of four different tests each linked to a specific treatment not only leads to increased cost and time spent on each case, but also to increased use of tissue that, since small biopsies and cytology is often the only specimen for lung cancer patients, may be needed for other treatment predictive molecular analyses.
Furthermore, evaluation of PD-L1 immunohistochemical staining may be difficult. Macrophages often exhibit membranous staining and may be misinterpreted as cancer cells in tissue samples from the lung. Also, unspecific cytoplasmic staining may occur, and weak partial membranous staining of tumors cells count as positive but may be difficult to detect. The mentioned difficulties are especially troublesome when a very limited proportion of tumor cells such as 1% is sufficient for a positive test, as is the case for some markers, and may lead to significant interobserver variation.
With this background, the aim of the present study was to compare the staining properties of tumor cells between the four antibody clones 28-8, 22C3, SP142, and SP263. Furthermore, the aim was to investigate interrater variation between several different pathologists to see if these stainings can be safely evaluated in the clinical setting. Staining of consecutive sections from tissue microarrays is ideal for such comparisons and has been used in the present study. A secondary aim was to investigate if a correlation could be seen between mRNA levels of the PD-L1 gene (CD274) and the protein expression measured with immunohistochemical staining.
Materials and methods
Study Material
Patient samples were selected from the Uppsala Lung Cancer Cohort, a consecutive tissue collection of lung cancer cases surgically treated at the Uppsala University between the years 2006 and 2010.12 Tumor areas from 58 blocks of formalin-fixed paraffin-embedded tissue were selected to compile a single tissue microarray. Each of the 58 cases was represented by two 1 mm cores. As three cases did not present sufficient tumor cells (<100) on the sections of the tissue microarray, they were excluded from the analysis. Patient characteristics are presented in Table 1. All cases were annotated in accordance with the 4th edition of the WHO classification system.13 The study was conducted in adherence to the Declaration of Helsinki and approved by the regional ethical review boards in Uppsala (Dnr 2012/532).
Immunohistochemical Staining and Evaluation
Consecutive 4 μm thick sections from the tissue microarray were pretreated and stained with the PD-L1 antibodies 28-8 and 22C3 from pharmDx on a Dako Autostainer PT Link 48 with EnVision DAB Detection System (Agilent/Dako, Santa Clara, CA, USA) and the SP142 and SP263 from Ventana on a Ventana Benchmark Ultra with OptiView Universal DAB Detection Kit (Ventana Medical Systems, Tucson, AZ, USA), respectively, in accordance with the manufacturers’ instructions. In addition, PD-L1 clone 28-8 from Abcam (Cambridge, UK), hereafter denoted 28-8A, was used for staining on the Ventana Benchmark Ultra with OptiView Universal DAB Detection Kit (dilution 1:100, incubation time/temperature 32 min/36 dgr, pretreatment with Ventana Cell Conditioning 1 pH 8, heating time/temperature 32 min/100 dgr). Tonsil tissue was included as positive control for low expression (macrophages in germinal centers) and high expression (crypt epithelium) for all assays. Controls provided with the pharmDx/Dako 28-8 and 22C3 assays (sections from cellblocks with PD-L1 positive and negative cell lines) were also included in each run for these assays, in accordance with the manufacturers’ instructions. For 28-8A, placental tissue was included as control in addition to tonsil.
All slides were independently evaluated by three board certified pathologists (MG, PM, and SWF) and three senior resident pathologists (AP, HB, and MM) working with thoracic pathology on a daily basis and also by one junior resident pathologist (AJ). Four of the participants were formally trained in evaluating the PD-L1 22C3 assay by Targos/Dako/NordiQC (AJ, HB, MG, and PM). All were blinded to the other participants’ results as well as to their own previous results. For parallel histomorphological evaluation a hematoxylin–eosin stained slide (consecutive section) was available. No therapeutic or outcome data was available during the study. Each case (average of both tissue microarray cores together) was separated into groups with <1% (score 0), 1–4% (score 1), 5–9% (score 2), 10–24% (score 3), 25–49% (score 4), or ≥50% (score 5) positive tumor cells. A cell with linear membranous staining, at least weak and partial, counted as positive.
RNA Sequencing and Data Analysis
For 35 of the 55 cases, RNA sequencing data (RNAseq) was available.14 In brief, RNA was extracted from fresh frozen tissue from the same tumor previous to formalin fixation. Samples were prepared for sequencing using the Illumina TruSeq RNA Sample Prep Kit v2, using polyA selection. The sequencing was performed multiplexed with 5 samples per lane on the Illumina HiSeq2500 machine (Illumina, San Diego, CA, USA) using the standard Illumina RNAseq protocol with a read length of 2 × 100 bases.
The raw sequencing data were mapped to the human reference genome (GRCh37) and the Ensembl version 73 gene annotation using TopHat version 2.0.8b.15, 16 Values for gene fragments per kilobase of transcript per million mapped reads were calculated from the generated alignments using Cufflinks version 2.1.1. Raw read counts were calculated using featureCounts from the Subread package version 1.4.0-p1.17
Statistical Analysis
Weighed kappa (linear weight) was used for interrater variability. For each case, a consensus score was calculated for each staining based on the score of the majority of observers, that is, four or more pathologists in agreement. If there was no majority (5% of the cases), the median score, here also equal to the mean in all these cases, was applied instead. Weighed kappa (linear weight) was then used to assess variability between assays. Furthermore, to assess differences in positive and negative cases between cutoffs, antibody clones and pathologists, Kruskal–Wallis test and Mann–Whitney U-test were used (based on a non-normal distribution). Correlations between mRNA levels and immunohistochemistry scores (consensus scores for each assay) were investigated using Spearman’s rank correlation coefficient. A P-value of <0.05 was considered statistically significant. The Spearman’s rank correlations were performed using the R version 3.2.1 (R Core Team, 2015). The other statistical analyses were performed using MedCalc for Windows, version 14.12.0 (MedCalc Software, Belgium).
Results
Staining Pattern in the Study Population
The study population included 55 lung cancer cases comprised of 29 adenocarcinomas, 23 squamous cell carcinomas, 2 large cell carcinomas, and 1 large cell neuroendocrine carcinoma. Patient characteristics are presented in Table 1 and were not significantly different compared with the whole Uppsala Lung Cancer Cohort. One squamous cell carcinoma case was not evaluable for 28-8A due to missing cores on the slide. All other cases were evaluable for all five assays.
For consensus score, four or more pathologists were in agreement in 261 of 274 (95%) of scored cases (55 cases, 5 assays, one case not evaluable for 28-8A). In the remaining 13 cases (5%), the median score was basis for the consensus score. Five or more pathologists were in agreement for 237 (86% of 274) of the cases.
Depending on assay, 16–44% of the cases showed a PD-L1 positivity of ≥1% tumor cells (based on consensus score), see Table 2. Comparative images of representative and discordant cases are found in Figure 1. Among non-tumor cells, membranous staining was commonly seen in macrophages, but cytoplasmic or no staining also occurred. Also, infrequent positive lymphocytes could be seen with all assays, but were not further evaluated in the present study (Figure 2).
Lung cancer cases stained with hematoxylin–eosin and different PD-L1 assays. Original magnifications × 200. (a–f) A case positive in all assays. (g–l,m–r) Two different cases with variation in staining properties between assays. (s–x) A case with unspecific cytoplasmic staining in 28-8, SP263, and 28-8A (Abcam) PD-L1 assays.
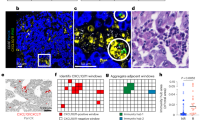
Staining of macrophages and lymphocytes. Original magnifications × 200. (a,b) A case with macrophages negative for PD-L1. (c,d) A case with macrophages with cytoplasmic PD-L1 staining. (e,f) A case with macrophages with membranous PD-L1 staining (the most common finding). (a,c,e) Stained with hematoxylin–eosin. (b,d,f) Stained with 22C3 and 28-8, respectively. (g,h) Lymphocytes positive for PD-L1, SP142, and 22C3, respectively.
Variation in Scoring Between Pathologists
Interobserver variation between pathologists, measured by weighted kappa in pairwise analyses, was in the range 0.71–0.96. See Table 3 for full data. The highest kappa values were seen for the SP142 assay (0.81–0.96), followed by 22C3 (0.71–0.95), 28-8A (0.80–0.95), 28-8 (0.80–0.93), and SP263 (0.75–0.91). With Kruskal–Wallis and pairwise Mann–Whitney U-tests, the kappa values were significantly higher for SP142 compared with SP263 and 28-8, while the kappa values for 22C3 and 28-8A were also significantly higher than those for SP263 (all P<0.013).
The individual pathologists’ scoring was compared with consensus. Using ≥1% stained tumor cells as cutoff for a positive test, 0–11 (0–20%) of the cases (median 3 cases; mean±s.d. 3.2±2.3) were differently classified as positive or negative by any one pathologist in comparison with consensus. The corresponding figure using ≥50% stained tumor cells as cutoff was 0–3 cases (0–5%; median 1 case; mean±s.d. 1.0±0.80). See Table 4 for details on each assay and pathologist. For the other cutoff levels, the figures were 0–6 cases (0–11%; median 1 case; mean±s.d. 1.5±1.4) for cutoff ≥5%, 0–6 cases (0–11%; median 1 case; mean±s.d. 1.4±1.6) for cutoff ≥10%, and 0–3 cases (0–5%; median 1 case; mean±s.d. 0.9±0.9) for cutoff ≥25%, respectively.
The variation in number of differently classified cases by any one pathologist compared with consensus was statistically significant between cutoffs (Kruskal–Wallis test; not separated for different assays). Using pairwise Mann–Whitney U-tests, the number of differently classified cases was significantly higher for the ≥1% cutoff than every other cutoff (all P<0.0001). No other statistically significant differences were seen.
The variation in number of differently classified cases by any one pathologist compared with consensus was also statistically significant between assays (Kruskal–Wallis test; not separated for different cutoffs). Using pairwise Mann–Whitney U-tests, the number of differently classified cases was significantly lower for the SP142 assay than for the SP263, 28-8, and 28-8A assays (all P<0.02). Also the 22C3 and 28-8A assays had a significantly lower number of differently classified cases than SP263 (both P<0.03). No other statistically significant differences were seen.
There was no obvious association between experience (specialist vs resident and formal PD-L1 training vs no training) and either high kappa value for interobserver variation or number of cases in agreement with consensus score.
Variation in Scoring Between Assays
In the next step, the variation between the different PD-L1 assays was assessed. The number of positive cases in the study population and depending on histological type, using ≥1% and ≥50% as cutoff for a positive test, is presented for each assay in Table 2.
Weighted kappa values between the five assays based on the consensus score for each case are presented in Table 5. As seen, the kappa values were in the range 0.45–0.91. The highest kappa values were seen for comparisons between 28-8, 22C3, and 28-8A (0.88–0.91), while the lowest values were seen for comparisons involving SP142 (0.45–0.63).
Using ≥1% stained tumor cells as cutoff for a positive test, 3–8 of the 55 cases (5–15%; median 4.5 cases) were differently classified as positive or negative when comparing any two assays excluding SP142. Agreement was higher using the ≥50% cutoff with only 1–3 of 55 cases (2–5%; median 2 cases) differently annotated. See Figure 3 for details. The number of differently classified cases was significantly lower using ≥50% cutoff than the ≥1% cutoff (Mann–Whitney U-test, P<0.01). Corresponding figures for both the ≥10% and ≥25% cutoffs were 0–4 cases (0–7%; median 2 and 1.5 cases, respectively), while the number of differently classified cases was 2–8 (4–15%; median 5 cases) for the ≥5% cutoff.
Correspondingly, when comparing two assays whereof one being SP142, 7–15 of the 55 cases (13–27%; median 11 cases) were differently classified as positive or negative using ≥1% stained tumor cells as cutoff for a positive test. The figure for the ≥50% cutoff was 7–10 cases (13–18%; median 8 cases), while the lowest number of differently classified cases (4–8 cases; 7–15%; median 7 cases) was seen for the ≥25% cutoff when comparing SP142 with any other assay.
Correlation of PD-L1 Scores with MRNA Levels
RNAseq data was available for 35 of the 55 cases represented on the tissue microarray. When the consensus score for each assay was used as reference, the correlation between protein and mRNA levels was significant for all assays using Spearman’s rank correlation (all P<0.0001). The highest rho values was seen for SP263 with rho 0.780 (95% CI 0.603–0.883) and 28-8 with rho 0.758 (0.568–0.871), followed by 28-8A rho 0.663 (0.423–0.816), 22C3 rho 0.651 (0.406–0.809) and SP142 rho 0.643 (0.394–0.804). Scatter plot diagrams demonstrating the distributions are presented in Figure 4.
Discussion
Checkpoint inhibitors targeting the immune regulatory molecules PD-1 and PD-L1 have now been approved for treatment of the main lung cancer types in advanced stage. For the majority of these novel drugs, the best predictor of response is PD-L1 expression in patients’ tumor tissue. In this study, we examined five different PD-L1 assays on 55 lung cancer cases that were independently annotated by seven pathologists. We demonstrated relatively good concordance using the clones 22C3, 28-8 and SP263 both as provided diagnostic tests as well as the in house protocol using clone 28-8 on the Ventana system. Not unexpected, the clone SP142 did show the highest deviation from the reference scores.11 However, SP142 is intended for evaluation of tumor associated immune cells in addition to tumor cells, which is a different analytical concept compared with the other tests, and the comparison of SP142 to other assays should be interpreted with caution.
Furthermore we demonstrated a relatively good interrater agreement for PD-L1 scoring between the pathologists, independent of training and professional education. Still, pathologists are a significant source of variability in the assessment of PD-L1 expression. Up to 20% of the cases were differently classified as positive or negative depending on the pathologist and used cutoff for a positive test. This is important to keep in mind if elaborate efforts to harmonize PD-L1 assays are to be undertaken.
One of the main strengths of the present study is its clinical applicability. It involved several pathologists working with thoracic pathology at different centers and with different levels of experience. All assays used in clinical trials were performed in the clinical routine within the local pathology departments. Also, all relevant cutoffs were evaluated with comprehensive data on interrater and interassay variation presented.
Tissue microarray is probably the best choice for method comparisons and investigation of interrater variation, but confirmation using biopsies and cytological specimen would also be preferable, as would an even larger study population. The number of cases was especially limited in the mRNA analysis, and these results must be interpreted with caution. Also, if interrater variation between pathologists created artefactual differences between the antibody clones, it could have affected the comparisons between assays. However, 95% of the consensus scores derived from the independent score of a majority of pathologists, which limits the risk. Furthermore, the experience of PD-L1 evaluation among the involved pathologists increased during the course of the study, but as all the pathologists were familiar with PD-L1 evaluation and did not evaluate the slides in the same order, it probably did not significantly influence the results.
Our data supports that the variation between the 22C3, 28-8, and SP263 assays is limited and comparable to differences between, for example, immunohistochemistry and fluorescence in situ hybridization for anaplastic lymphoma kinase, two assays today regarded as similar enough for either one to be used.18, 19, 20, 21 It is noteworthy that there was some variation between the assays concerning interrater agreement between pathologists. For SP142, the many obviously negative cases may have been an important factor for the high kappa values for this antibody clone (interrater agreement for scoring lymphocytes was not investigated). In our opinion, clone 28-8 exhibited slightly more cases with unspecific cytoplasmic staining, and the antibody from Abcam somewhat more so than the Dako assay (Figure 1). This may perhaps cause more interrater variation.
The highest interrater variability was observed for the ≥1% cutoff, with up to 20% of the cases classified differently between any one pathologist and consensus. In this perspective, it seems that immunohistochemistry for PD-L1 at the present may not be a solid test for treatment decision if the ≥1% cutoff is used for a positive test. All involved pathologists were well aware of macrophages being a pitfall in evaluation of PD-L1, but this may still have caused trouble. Also, different interpretation of very few or sometimes even single cells may lead to a case being classified as positive instead of negative or vice versa if using ≥1% as cutoff. According to our results, any other cutoff would be a better option with interrater agreement between pathologists in mind. A cutoff of 50% is well supported as clinically relevant in treatment studies.10
Only few other studies have compared the four PD-L1 clones used as diagnostic tests in clinical trials. The industrial-academic cooperation named Blueprint project published their first comparative study testing the four assays on 39 lung cancer cases with interpretation of results by three expert pathologists.11 The samples were in the majority of cases resected specimen and the staining was performed centrally at the vendor’s facilities. In contrast to our study, they compared the agreement based on assay-specific cutoffs, for example, SP263 positivity defined with ≥25% cutoff versus 28-8 ≥1% cutoff. Therefore the results are not exactly comparable, but the conclusion that the use of the three antibodies 28-8, 22C3, and SP263 results in relatively similar staining patterns is in line with our results.
The German harmonization study tested the four antibody clones on 15 resected specimens, and the evaluation was performed by nine pathologists.22 The two Dako assays with clone 28-8 and 22C3 showed similar scoring results. A larger study with 493 samples compared with the three antibody assays 28-8, 22C3, and SP263 has been presented.23 These samples were annotated by only one pathologist, but a subset of 200 cases was also reevaluated by an independent rater. Although interpretation of the data is difficult, the general conclusion that the three assays are interchangeable can be drawn also from this study. The latest comparison was a multi-institutional study including 90 resected cases evaluated by 13 pathologists.24 Staining included the 28-8, 22C3 and SP142 assays, as well as clone E1L3N (Cell Signaling), but not the Ventana SP263 assay. Again, the assays 28-8 and 22C3 showed comparable staining patterns, although with significantly but slightly lower scores for 22C3. While the concordance between pathologists was generally very good for scoring of tumor cells, it was limited for scoring of immune cells.
Based on the results of the German harmonization trial, the study by Rimm and co-workers and our study we believe that the major challenge is not the assay compatibility but the variability between pathologists’ assessment.22, 24 Although we did not see a significant difference due to the pathologists’ experience, our study with seven pathologist is possibly too small and not optimally structured to draw any conclusions whether training improves the intrinsic subjectivity of scoring immunohistochemical stainings. In our opinion, it is still preferable that pathologists experienced in thoracic pathology evaluate PD-L1 stainings in the clinical setting as familiarity with lung cancer and non-neoplastic cells in lung tissue is of obvious importance.
In our study, 16–44% of the cases showed any PD-L1 positivity, with SP263 exhibiting the highest number of positive cases and SP142 the lowest (though lymphocytes were not evaluated), in line with the German harmonization study.22 More positive cases can probably be expected from evaluations on whole slides of resected tumors, but the number of positive cases seen here is lower than those in the published clinical trials, with 61–72% positive cases with evaluation mainly on biopsies.4, 9, 25 The cause of this obvious difference is unclear. However, our results are in line with our clinical experience, with 40% positive cases of more than 100 unselective clinically tested lung adenocarcinomas at the Department of Pathology in Lund (28-8 Abcam on a Ventana Benchmark Ultra using ≥1% as cutoff).
Interestingly, the PD-L1 mRNA levels correlated with the immunohistochemistry scores for all assays. Although the overlap was not perfect, there was not a single distinct outlier showing strong mRNA levels but low PD-L1 protein levels or vice versa. This indicates that mRNA levels have also a potential as predictive biomarkers for PD-L1 immunotherapy. It would be possible to integrate PD-L1 mRNA levels in the newly developed multiplex gene expression assays applicable on small formalin-fixed paraffin-embedded tissue biopsies. For example, the NanoString technique is already today used to identify fusion genes in lung cancer samples and the PD-L1 assessment could be easily and cost-effectively included in this analysis. However, it is known that lung cancer not expressing PD-L1 (<1% positive cells) may respond to anti-PD-1 therapy,25 and correlation to benefit of treatment (not assessed in the present study) also needs to be evaluated for mRNA.
In conclusion, our study confirmed the overall good concordance between the PD-L1 assays including clone 28-8, 22C3, and SP263, indicating that one of these tests may presently be sufficient as basis for treatment decision for nivolumab, pembrolizumab and durvalumab. The improvement of interrater agreement remains an important challenge, and the ≥1% cutoff for a positive test seems to be the most problematic issue. This may hopefully be overcome by factors such as better training, longer assay experience and/or cutoff optimization.
References
Herbst RS, Soria JC, Kowanetz M et al. Predictive correlates of response to the anti-PD-L1 antibody MPDL3280A in cancer patients. Nature 2014;515:563–567.
Borghaei H, Paz-Ares L, Horn L et al. Nivolumab versus docetaxel in advanced nonsquamous non-small-cell lung cancer. N Engl J Med 2015;373:1627–1639.
Brahmer J, Reckamp KL, Baas P et al. Nivolumab versus docetaxel in advanced squamous-cell non-small-cell lung cancer. N Engl J Med 2015;373:123–135.
Garon EB, Rizvi NA, Hui R et al. Pembrolizumab for the treatment of non-small-cell lung cancer. N Engl J Med 2015;372:2018–2028.
Gettinger SN, Horn L, Gandhi L et al. Overall survival and long-term safety of nivolumab (anti-programmed death 1 antibody, BMS-936558, ONO-4538) in patients with previously treated advanced non-small-cell lung cancer. J Clin Oncol 2015;33:2004–2012.
Rizvi NA, Mazieres J, Planchard D et al. Activity and safety of nivolumab, an anti-PD-1 immune checkpoint inhibitor, for patients with advanced, refractory squamous non-small-cell lung cancer (CheckMate 063): a phase 2, single-arm trial. Lancet Oncol 2015;16:257–265.
Antonia S, Goldberg SB, Balmanoukian A et al. Safety and antitumour activity of durvalumab plus tremelimumab in non-small cell lung cancer: a multicentre, phase 1b study. Lancet Oncol 2016;17:299–308.
Fehrenbacher L, Spira A, Ballinger M et al. Atezolizumab versus docetaxel for patients with previously treated non-small-cell lung cancer (POPLAR): a multicentre, open-label, phase 2 randomised controlled trial. Lancet 2016;387:1837–1846.
Herbst RS, Baas P, Kim DW et al. Pembrolizumab versus docetaxel for previously treated, PD-L1-positive, advanced non-small-cell lung cancer (KEYNOTE-010): a randomised controlled trial. Lancet 2016;387:1540–1550.
Reck M, Rodríguez-Abreu D, Robinson AG et al. KEYNOTE-024 Investigators. Pembrolizumab versus chemotherapy for PD-L1-positive non-small-cell lungcancer. N Engl J Med 2016;375:1823–1833.
Hirsch FR, McElhinny A, Stanforth D et al. PD-L1 immunohistochemistry assays for lung cancer: results from phase 1 of the Blueprint PD-L1 IHC Assay Comparison Project. J Thorac Oncol 2017;12:208–222.
Micke P, Mattsson JS, Djureinovic D et al. The impact of the fourth edition of the WHO Classification of Lung Tumours on histological classification of resected pulmonary NSCCs. J Thorac Oncol 2016;11:862–872.
WHO working group. Tumours of the lung. In: Travis WD, Brambilla E, Burke AP, Marx A, Nicholson AG (eds). WHO Classification of tumours of the lung, pleura, thymus and heart 4th (ed). IARC Press: Lyon, France, 2015;9–151.
Djureinovic D, Hallström BM, Horie M et al. Profiling cancer testis antigens in non-small-cell lung cancer. JCI Insight 2016;1:e86837.
Flicek P, Amode MR, Barrell D et al. Ensembl 2012. Nucleic Acids Res 2012;40, (Database issue) D84–D90.
Trapnell C, Williams BA, Pertea G et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 2010;28:511–515.
Liao Y, Smyth GK, Shi W . featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 2014;30:923–930.
Wallander ML, Geiersbach KB, Tripp SR et al. Comparison of reverse transcription-polymerase chain reaction, immunohistochemistry, and fluorescence in situ hybridization methodologies for detection of echinoderm microtubule-associated proteinlike 4-anaplastic lymphoma kinase fusion-positive non-small cell lung carcinoma: implications for optimal clinical testing. Arch Pathol Lab Med 2012;136:796–803.
Sullivan HC, Fisher KE, Hoffa AL et al. The role of immunohistochemical analysis in the evaluation of EML4-ALK gene rearrangement in lung cancer. Appl Immunohistochem Mol Morphol 2015;23:239–244.
Mattsson JS, Brunnström H, Jabs V et al. Inconsistent results in the analysis of ALK rearrangements in non-small cell lung cancer. BMC Cancer 2016;16:603.
Marchetti A, Pace MV, Di Lorito A et al. Validation of a new algorithm for a quick and easy RT-PCR-based ALK test in a large series of lung adenocarcinomas: Comparison with FISH, immunohistochemistry and next generation sequencing assays. Lung Cancer 2016;99:11–16.
Scheel AH, Dietel M, Heukamp LC et al. Harmonized PD-L1 immunohistochemistry for pulmonary squamous-cell and adenocarcinomas. Mod Pathol 2016;29:1165–1172.
Ratcliffe MJ, Sharpe A, Midha A et al. Agreement between programmed cell death ligand-1 diagnostic assays across multiple protein expression cut-offs in non-small cell lung cancer. Clin Cancer Res 2017, (in press).
Rimm DL, Han G, Taube JM et al. A prospective, multi-institutional, pathologist-based assessment of 4 immunohistochemistry assays for PD-L1 expression in non-small cell lung cancer. JAMA Oncol 2017, in press.
Hellman MD, Rizvi NA, Goldman JW et al. Nivolumab plus ipilimumab as first-line treatment for advanced non-small-cell lung cancer (CheckMate 012): results of an open-label, phase 1, multicohort study. Lancet Oncol 2017;18:31–41.
Acknowledgements
We thank the immunohistochemistry sections at the Departments of Pathology in Uppsala, Lund and Umeå. This study was supported by the Regional Agreement on Medical Training and Clinical Research (ALF, Swedish government funding), the Swedish Cancer Society and the Lions Cancer Foundation, Uppsala, Sweden (non-profit organizations). The funders had no involvement in study design, data collection, analysis or interpretation, or writing the report etc.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
MG: consulting or advisory role for MSD and Roche Diagnostics. PM: stock and other ownership interests in Immunicum and Bioinvent. Consulting or advisory role for Roche Molecular Diagnostics, MSD Oncology, Bristol-Myers Squibb and HTG Molecular Diagnostics. Has received research funding from Roche Diagnostics. The remaining authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Brunnström, H., Johansson, A., Westbom-Fremer, S. et al. PD-L1 immunohistochemistry in clinical diagnostics of lung cancer: inter-pathologist variability is higher than assay variability. Mod Pathol 30, 1411–1421 (2017). https://doi.org/10.1038/modpathol.2017.59
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2017.59
This article is cited by
-
Association of artificial intelligence-powered and manual quantification of programmed death-ligand 1 (PD-L1) expression with outcomes in patients treated with nivolumab ± ipilimumab
Modern Pathology (2022)
-
Digital pathology and artificial intelligence in translational medicine and clinical practice
Modern Pathology (2022)
-
Artificial intelligence-assisted system for precision diagnosis of PD-L1 expression in non-small cell lung cancer
Modern Pathology (2022)
-
Choice of PD-L1 immunohistochemistry assay influences clinical eligibility for gastric cancer immunotherapy
Gastric Cancer (2022)
-
Automated tumor proportion scoring for PD-L1 expression based on multistage ensemble strategy in non-small cell lung cancer
Journal of Translational Medicine (2021)