Abstract
Exosomes are small membrane vesicles that have important roles in transporting a great variety of bioactive molecules between epithelial compartment and their microenvironment during tumor formation including colorectal adenoma–carcinoma sequence. We tested the mRNA expression of the top 25 exosome-related markers based on ExoCharta database in healthy (n=49), adenoma (n=49) and colorectal carcinoma (n=49) patients using Affymetrix HGU133 Plus2.0 microarrays. Most related genes showed significantly elevated expression including PGK1, PKM, ANXA5, ENO1, HSP90AB1 and MSN during adenoma–carcinoma sequence. Surprisingly, the expression of ALIX (ALG 2-interacting protein X), involved in multivesicular body (MVB) and exosome formation, was significantly reduced in normal vs adenoma (P=5.02 × 10−13) and in normal vs colorectal carcinoma comparisons (P=1.51 × 10−10). ALIX also showed significant reduction (P<0.05) at the in situ protein level in the epithelial compartment of adenoma (n=35) and colorectal carcinoma (n=37) patients compared with 27 healthy individuals. Furthermore, significantly reduced ALIX protein levels were accompanied by their gradual transition from diffuse cytoplasmic expression to granular signals, which fell into the 0.6–2 μm diameter size range of MVBs. These ALIX-positive particles were seen in the tumor nests, including tumor–stroma border, which suggest their exosome function. MVB-like structures were also detected in tumor microenvironment including α-smooth muscle actin-positive stromal cells, budding off cancer cells in the tumor front as well as in cancer cells entrapped within lymphoid vessels. In conclusion, we determined the top aberrantly expressed exosome-associated markers and revealed the transition of diffuse ALIX protein signals into a MVB-like pattern during adenoma–carcinoma sequence. These tumor-associated particles seen both in the carcinoma and the surrounding microenvironment can potentially mediate epithelial–stromal interactions involved in the regulation of tumor growth, metastatic invasion and therapy response.
Similar content being viewed by others
Main
Exosomes are small (~30–100 nm in diameter), membrane vesicle-like structures, which transport complex information between cells mainly associated with pathological conditions including human carcinomas.1, 2 These powerful intercellular regulators originate from endosomal, internal vesicles of large multivesicular bodies (MVBs).3, 4 Although several details of exosome genesis remains unknown, its key molecular complexes were identified, such as endosomal sorting complex required for transport (ESCRT) machinery, which is involved in MVB biogenesis and sorting.5, 6 After fusion with the plasma membrane, MVBs release their content into the extracellular compartment as exosomes.3, 4 Exosome release is mainly studied in tumor cell cultures. Regarding to colorectal carcinoma, the vast majority of examined in vitro models (eg, HT-29, SW480 and HCT-116 cultures)7, 8, 9 contain heterogeneous population of partially differentiated cells (ie, low expression of CD133 and CD44 stem cell markers).10, 11, 12, 13 The role of colorectal carcinoma cells with stem cell characteristics, for example, high CD133, CD44 and Musashi1 expression,14 in exosome releasing has not been analyzed in colorectal carcinoma cell cultures and particularly in situ at tissue level.
As paracrine mediators, the tumor cell-released exosomes may influence the behavior of adjacent cells by delivering oncogenic proteins (eg, KRAS) and receptors (eg, epidermal growth factor receptor variant III), as well as receptor ligands (eg, transforming growth factor-β) and epigenetic regulators, such as microRNAs.15, 16 This process also has an important role in developing tumor chemoresistance and interfering with the tumor microenvironment by modifying stromal cell functions, such as neovascularization, immunosuppression, tumor cell invasion (eg, epithelial to mesenchymal transition) and transition of stromal cells to carcinoma-associated fibroblasts (CAFs).1, 6, 17, 18 CAFs as key factors of abnormal epithelial–stromal interaction and their powerful tumor-supportive effect (eg, increased regulator ligand and extracellular matrix molecule expression) are well known compared with normal stromal cells with similar morphology, that is, subepithelial myofibroblasts.18, 19, 20 Tumor cell-released exosomes are also detectable in the circulation where the plasma level of their specific proteins (eg, CD63 and tumor susceptibility gene 101/TSG101) and miRNA content may be useful as diagnostic and/or prognostic markers in different tumor types including colorectal carcinomas.21, 22, 23, 24
Our study was designed to analyze the top exosome-specific markers based on change of their tissue mRNA level during colorectal adenoma–carcinoma sequence. For testing exosomes at the in situ protein level, we selected the ALG 2-interacting protein X (ALIX; also known as programmed cell death 6-interacting protein (PDCD6IP)), a multifunctional protein, which is involved in biogenesis of MVBs (as ESCRT I–III binding protein), lysosomal degradation and accumulates in exosomes.3, 25, 26, 27 ALIX has been tested widely in exosome studies of colorectal carcinoma cell cultures9, 28 and has shown diagnostic and prognostic significance in different tumors.29, 30, 31 Besides testing the transcript levels, here we determined ALIX protein expression in situ in the epithelial and stromal compartment including α-smooth muscle actin (SMA)-positive cells in low- and high-grade dysplastic adenomas as well as non-metastatic and metastatic (both lymph node and distant) colorectal carcinomas with particular focus on individual Musashi1-positive cancer stem cells at the tumor front.
Materials and methods
In Silico mRNA Expression Analysis of Exosome Markers
Biopsy samples taken for mRNA expression microarray experiments were stabilized in RNALater Reagent (Qiagen, Germantown, MD, USA) and stored at −80 °C until use. Parallel surgical and biopsy samples were routinely fixed in 4% formaldehyde and embedded in paraffin wax for histopathology. Diagnoses were based on the WHO criteria32 using H&E-stained slides, which were also used for selecting representative areas into tissue microarray blocks. mRNA expression of the top 25 exosome markers (according to the Exocarta database; http://exocarta.org/exosome_markers) was analyzed using Affymetrix HGU133 Plus2.0 whole transcriptome data of 147 colorectal biopsy samples (containing both epithelial and stromal compartments) from 49 healthy, 49 adenoma, including 25 low-grade (mean age at diagnosis: 66±13 years, 12f/13m) and 24 high-grade adenoma (mean age: 70±12 years, 13f/11m) and 49 colorectal carcinoma (including 24 non-metastatic (Dukes A and B, mean age: 70±10 years, 15f/9m) and 25 metastatic (Dukes C and D, mean age: 66±12 years, 12f/13m)) patients (according to Astler-Coller-modified Dukes’ classification) previously hybridized by our research group (GEO serial accession numbers: GSE37364,33 GSE1071434 and GSE4183;35 Supplementary Table 1). Detailed patients data were described previously.33, 34, 35 For confirmation of exosome marker mRNA expression results revealed from our microarray data originate from all stages of colorectal adenoma–carcinoma sequence, four additional GEO data sets were also involved in the analysis, three of them includes HGU133 Plus2.0 microarray data of colorectal carcinoma (n=99) and normal/normal adjacent tissue (NAT) (n=39) tissue samples (GSE18105,36 GSE410737 and GSE934838) and one of them with HGU133 Plus2.0 microarray data of adenoma (n=32) and normal/NAT (n=32) biopsy samples (GSE8671)39 (Supplementary Table 1).
Immunohistochemistry
Immunohistochemistry was performed on healthy (n=27), adenoma (n=35), including 13 low-grade (mean age at histology examination: 68±14; 7f/6m) and 22 high-grade (mean age: 61±12 years; 14f/8m) dysplasia (according to the description of Fleming et al40) as well as non-metastatic (n=12; including 3 Dukes A and 9 Dukes B, mean age: 61±5 years; 2f/10m) and metastatic (n=25; included 14 Dukes C and 11 Dukes D, mean age: 62±7 years; 9f/16m) colorectal carcinoma samples. Written informed consent was provided by all patients. The study was approved by the local ethics committee (Semmelweis University Regional and Institutional Committee of Science and Research Ethics; Nr.: ETT TUKEB 23970/2011 and 8-23/2009-1018EKU(ad.60/PI/09)). We used anti-ALIX (HPA011905, Sigma-Aldrich, St Louis, USA; 1:400) monoclonal antibody (labeled with Alexa Fluor 546, A11035, Invitrogen, Eugene, CA, USA) for immunohistochemical detection of exosomes. Anti-cytokeratin (CK; AE1/AE3, Dako, Glostrup, Denmark; 1:100) and anti-Musashi1 (EP1302, Abcam, Cambridge, USA; 1:100, both antibodies were labeled with Alexa Fluor 488, Invitrogen) were used for identification of epithelial and stem cells, respectively. Antibody for Ki-67 (MIB-1, M7240, Dako; 1:150, labeled with Alexa Fluor 488) was used to examine the cell proliferation. We identified lymphatic vessel cells using antibody to podoplanin (PDPN; D2-40, M3619, Dako; 1:200, labeled with Alexa Fluor 488) and the stromal myofibroblast and CAFs using anti-α-SMA (1A4, Dako, CA, USA; 1:1, labeled with Alexa Fluor 488). H&E and immunohistochemistry slides were digitally archived using Pannoramic 250 Flash II (with Zeiss Plan-Apochromat 20x objective; 3DHISTECH, Hungary, Budapest) and Pannoramic Confocal (with Zeiss C-Apochromat 63x objective; 3DHISTECH) digital scanners. Owing to the discrete Musashi1 expression, in the case of ALIX/Musashi1 double staining, we used two step staining and double digitalization as described previously.41 The digital slides were analyzed with a Pannoramic Viewer (v: 1.15.3, 3DHISTECH) digital microscope. Areas of interest were selected on the basis of morphology analyses of H&E-stained tissue microarray slides, then immunohistochemistry results were assessed on parallel slides. The percentage of cells with granular ALIX expression (PCGE) was determined by counting 800–1300 cells both in the epithelial and stromal compartments (including at least 80 α-SMA-positive cells in subepithelial region) in five representative core/groups with the Marker Counter module of the Pannoramic Viewer program. ALIX expression was validated in epithelial/tumor and in stromal cells with the modified Q-score method, that is, multiplying the percentage of the area of positive particles (P) in the cytoplasm of epithelial/carcinoma and stromal cells by the intensity (I; 0,+1,+2,+3); Formula: Q=P x I; maximum: 300 (+3 x 100).42 Three-dimensional (3D) reconstruction was created based on the combination of nine Z-axial confocal layers (0.4 μm intervals) applying 3DView software (v: 2.2.0; 3DHISTECH).
Statistical Analysis
Preprocessing of in silico gene expression microarray analysis was performed using Guanine Cytosine Robust Multi-array Average (GCRMA) method (including quantile normalization). In case of pairwise comparisons, the differentially expressed genes were determined using paired Student's t-test with Benjamini and Hochberg correction. For LogFC calculation, differences between the group averages were considered. When more than two sample groups were compared, ANOVA and Tukey's honest significant difference post tests were applied for statistical analysis.
In case of protein expression—because of their non-normal distribution—Kruskal–Wallis test and post-analysis test were applied where pairwise comparison of the different subgroups were performed according to Conover.43 Statistically significant states were considered at P<0.05. Boxplots were constructed for visualization. Statistical analyses were performed under R 3.2.1. environment.44
Results
Diminished ALIX Exosome Marker mRNA Expression in Pre-Neoplastic and Cancerous Colorectal Biopsy Samples
Twenty from the studied top 25 exosome markers (according to the ExoCarta data) showed significantly differential mRNA expression in adenoma and colorectal carcinoma tissue compared with normal biopsy samples (P<0.05), seven with absolute value of logFC higher than 0.5.
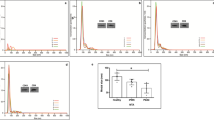
Most increased expression of PGK1, PKM, ANXA5, ENO1, HSP90AB1 and MSN mRNAs was detected in colorectal carcinoma compared with healthy normal samples, whereas PKM, HSP90AB1, ANXA2, MSN, EEF1A1 and ENO1 were found to be upregulated in adenoma vs normal comparison (Table 1). ALIX (PDCD6IP) showed the most significant downregulation in low- and high-grade dysplastic adenomas, as well as non-metastatic and metastatic colorectal carcinoma samples according to both Affymetrix IDs (217746_s_at and 222394_at) representing ALIX transcripts (Figure 1).
ALIX mRNA expression during the colorectal adenoma–carcinoma sequence according to the normalized HGU133 Plus2.0 microarray expression values of 217746_s_at (a) and 222394_at (b) Affymetrix IDs representing ALIX transcripts. (The * denotes statistically significant differences (P<0.01) between groups). HgAD, high-grade dysplastic adenoma; LgAD, low-grade dysplastic adenoma; MetCRC, metastatic colorectal carcinoma; N, normal; NmetCRC, non-metastatic colorectal carcinoma.
Beside significant upregulation of PGK1, PKM, ENO1, HSP90AB1 and MSN exosome marker genes, downregulation of ALIX (PDCD6IP) mRNA in colorectal carcinoma samples compared with normal tissue was confirmed on HGU133 Plus2.0 microarray data sets of others as well.36, 37, 38, 39 In adenoma vs normal comparison, results of GSE8671 data set completely supported the exosome marker mRNA level alterations revealed in analysis of our microarray data sets (Supplementary Table 2).
Altered ALIX Protein Expression in Epithelial Compartment During Colorectal Adenoma–Carcinoma Sequence
Based on mRNA expression results, we examined the protein expression pattern of ALIX in all histological groups of the adenoma–carcinoma sequence in the cytokeratin (CK)-positive epithelial and carcinoma compartment (white arrowheads in Figure 2). Strong ALIX expression (Q-score: 232.22±16.01; Figure 2b) with diffuse cytoplasmic expression pattern (percentage of cells with granular expression: 2.18±0.41) was observed in normal epithelium. Low-grade adenomas with mild dysplasia (ie, polarized, moderately enlarged and elongated nuclei, single-cell layer or mild pseudostratification, retained cytoplasmic mucin) showed a similar pattern, but decreased (mainly moderate) protein expression compared with normal cells (Figure 2e). Our definite low-grade (ie, prominent pseudostratification and mucin depletion) and high-grade adenomas (Figure 2h) showed mild diffuse and moderate/strong granular, MVB-like epithelial ALIX expression (Q-scores: 142.85±16.37 and 143.63±14.65; percentages of cells with granular expression: 43.68±31.68 and 56.78±13.40, respectively). In adenoma cases, ALIX expression showed a heterogeneous pattern in neighboring glands (Supplementary Figure 1/A) and even in the epithelium of the same gland (Supplementary Figure 1/B). ALIX expression showed strong granular pattern in the carcinoma compartment of non-metastatic (Q-score: 145.86±21.08; percentage of cells with granular expression: 89.98±10.73; Figure 2k) and metastatic (Q-score: 143.20±15.73; percentage of cells with granular expression: 99.52±0.59; Figure 2n) colorectal carcinoma samples. The Q-scores and percentage of cells with granular expression values of epithelial compartment are indicated in Figures 3a and b.
ALIX (red fluorescent staining) co-expression with CK and Ki-67 (both molecules were labeled with green fluorescent staining) in the normal (b, c), adenoma with low-grade (LgAD; e, f) and high-grade (HgAD; h, i) dysplasia, as well as in non-metastatic (NmetCRC; k, l) and metastatic colorectal cancers (MetCRC: n, o). The white arrowheads show the epithelial, whereas the yellow arrowheads show the stromal compartments. The correspondent H&E staining of b, e, h, k and n figures are depicted in a, d, g, j and m pictures. x40 magnification; scale bar: 50 μm. Digital microscopic images.
Expression change (Q-score) of ALIX exosome marker and percentage of cells with granular expression (PCGE) in epithelial (a, b) and stromal (c, d) component in different histological stages of colorectal adenoma–carcinoma sequence (The * denotes statistically significant differences (P<0.05) between groups).
Owing to challenging exosome detection in small-sized intercellular compartment in colonic tissues, we identified ALIX-positive particles at the cancer cell–stroma border where these structures mainly belong to the cancer cells (Figure 4a). They showed partial overlap with cytoplasmic CK staining (Figure 4b) from which some ALIX particles were budding off in 3D reconstruction (Figure 4c), others were also seen in adjacent or in stromal cells (Figures 4d and e).
Confocal microscopy images from ALIX-positive (red fluorescent staining) MVB-like structures (white arrows) in carcinoma cell–stromal border ((a, b); x360 magnification, scale bar: 5 μm, (c): 3D reconstruction of this area; CK: green fluorescent staining) and adjacent area to carcinoma cells ((d, e); x190 magnification, scale bar: 10 μm).
Stromal ALIX Protein Expression in Adenomas and Colorectal Carcinomas
We observed significant (P<0.05) differences in stromal ALIX expression among the normal, adenoma and colorectal carcinoma samples (Figure 3c). We found a low degree of diffuse stromal ALIX expression in normal samples (Q-score: 43.70±12.75; percentage of cells with granular expression: 7.36±1.49; Figure 2b), primarily in the cytoplasm of immune cells. Higher ALIX expression with an increased appearance of granular expression pattern was detected in the stroma of low-grade (Q-score: 66.00±23.23; percentage of cells with granular expression: 19.50±10.93; Figure 2e) and high-grade adenomas (Q-score: 64.09±19.18; percentage of cells with granular expression: 23.00±11.31; Figure 2h). Stroma of both non-metastatic (Q-score: 119.16±28.10; percentage of cells with granular expression: 61.96±20.98; Figure 2k) and metastatic (Q-score: 122.40±40.13; percentage of cells with granular expression: 79.48±15.6; Figure 2n) carcinomas showed strong granular exosome marker expression.
Among stromal cells, normal myofibroblasts showed low, diffuse ALIX expression (Figure 5a), granular protein expression was not detectable in these cells. In adenomas, α-SMA-positive cells typically showed low diffuse protein expression (Figure 5b), but granular pattern was detectable in some myofibroblasts (percentages of cells with granular expression were 8.08±5.63 in LgAD and 8.32±3.69 in HgAD). High percentage of CAFs were detected with low diffuse and strong granular ALIX expression in non-metastatic (percentage of cells with granular expression: 89.76±5.11) and metastatic (percentage of cells with granular expression: 90.54±4.98) colorectal carcinomas (Figure 5c).
(a, b) Low diffuse ALIX (red fluorescent staining) expression in myofibroblasts (yellow arrows) of normal and adenoma samples. (c) Granular, MVB-like protein expression (white arrows) in CAFs (yellow arrow; α-SMA: green fluorescent staining, x170 magnification, scale bar: 10 μm). (d) ALIX expression in individual, CK-positive (epithelial-origin; green fluorescent staining) budding tumor cells. (e) Granular ALIX (white arrowheads) and discrete cytoplasmic Musashi1-positivity (green fluorescent staining; yellow arrowheads) in individual cells (potential cancer stem cells) of in tumor stroma ((a, b) x200 magnification, scale bar: 10 μm). (f) Granular ALIX expression of tumor cell cluster in the lumen of podoplanin-positive (green fluorescent staining) lymphatic vessel (white arrows; immunohistochemistry and H&E staining, x15 magnification, scale bar: 50 μm). Digital microscopic images.
The ALIX-positive particles were bigger (approximately 0.6–2 μm in diameter) than the size of exosomes (30–100 nm in diameter)1 both in carcinoma and stromal cells (Supplementary Figures 1/C and D). Q-scores and percentage of cells with granular expression (PCGE) values for stromal compartment of normal, adenoma and carcinoma tissues were indicated in Figures 3c and d.
The Origin and Localization of ALIX-Positive Cells
Neither diffuse nor granular ALIX expression was limited to the Ki-67-positive cells; these also occurred in the non-proliferative cells during colorectal adenoma–carcinoma sequence (Figures 2c, f, i, l and o). Similarly to the observation of Nishimura et al,45 we found both nuclear and cytoplasmic Musashi1 expression in epithelial and epithelial-origin colorectal carcinoma cells. In the tumor front of metastatic colorectal carcinoma samples, granular ALIX expression was detected in the cytoplasm of individual, CK-positive (Figure 5d) and Musashi1-positive (Figure 5e) cancer cells or/and cancer stem cells. ALIX-positive particles were also detected in cancer cells in the lumen of larger podoplanin-positive lymphatic vessels (Figure 5f).
Discussion
Abnormal intercellular communication between tumor cells and their supportive microenvironment has a fundamental role in the formation and growth of different tumor types including colorectal carcinomas.46, 47, 48, 49 As an important part of this process, we found the steady elevation of most of the top exosome-associated genes during colorectal adenoma–carcinoma sequence, some of which had been examined before as secreted proteins (ie, HSP90, PGK1 and ENO1) in colorectal carcinoma cell cultures.50, 51 Few of the tested markers showed decreasing transcript levels including ALIX, which was one of the most widely examined exosome markers in cancer development.29, 30, 52, 53 Our gene expression data correlated well with those gained from independent databases,36, 37, 38, 39 showing similarly reduced ALIX mRNA levels during adenoma–carcinoma sequence, which was also validated at the in situ protein level. Similar alterations between whole biopsy mRNA and protein level in epithelial compartment suggest that the majority of ALIX originated from these components. Reduced global ALIX expression in colorectal carcinoma compared with normal colon epithelia was accompanied by the gradual transition of the protein expression pattern from diffuse to granular and may be associated with epigenetic suppression of gene activity within 3p22 chromosome region.54 Strong, diffuse expression of ALIX in the normal colonic epithelium (especially in the goblet cells) may also contribute to other epithelial functions besides its regulatory function in exosome biogenesis. In pre-neoplastic lesions, the heterogeneous appearance of diffuse to granular transition potentially reflects early signs of tumor development. In colorectal carcinoma samples, reduced diffuse and increased granular ALIX expression levels were detected both in tumor and stromal compartment. We found morphological and size similarity between the expression pattern of ALIX (in our adenoma and colorectal carcinoma samples) with TSG101 (involved in MVB and exosome formation, an interaction partner of ALIX) and CD63 molecules in in vitro and in vivo models of others.55, 56, 57, 58 Based on the relatively large (approximately 0.6–2 μm) diameter (knowing the inaccuracy deriving from fluorescent measurement)59 and molecular composition of these particles they may designate MVBs including intracytoplasmic clusters of exosomes. These structures were also detectable on cancer–stroma border and adjacent carcinoma cells, which suggest that they have a role in cell–cell communication via elevated exosomal function. Similarly, granular ALIX expression pattern and release of ALIX-positive exosomes were described in prostate and breast cancer stem cell cultures.60 Frequent appearance of granular ALIX expression in tumor microenvironment involving CAFs may refer to an increased cell–cell communication with potential clinicopathological importance)15, 61, 62, 63, 64 and may result the increased exosome level in plasma of colorectal carcinoma patients compared with normal persons.21 Excessive ALIX expression of individual cells in the stromal components and lumen of lymphatic vessels may become important indicator of the metastatic potential as these cells are essential in establishing a local and/or distant pre-metastatic microenvironment.8, 21, 65
In conclusion, here we identified the differential mRNA expression of the top exosome-related markers during colorectal carcinoma formation. Furthermore, we visualized in situ the potential exosome formation through detecting ALIX-positive MVB-like structures both in the epithelial and stromal compartments during adenoma–carcinoma sequence. These structures can be the substrates of exosome-based communication progressively developing from pre-neoplastic lesions to colorectal carcinoma transition. Our results may serve useful clues to understanding interactions between cancer and the surrounding microenvironment that may affect the regulation of tumor growth, metastatic invasion, as well as therapy response.
References
Yang C, Robbins PD . The roles of tumor-derived exosomes in cancer pathogenesis. Clin Dev Immunol 2011;2011:842849.
Braicu C, Tomuleasa C, Monroig P et al. Exosomes as divine messengers: are they the Hermes of modern molecular oncology? Cell Death Differ 2015;22:34–45.
Baietti MF, Zhang Z, Mortier E et al. Syndecan-syntenin-ALIX regulates the biogenesis of exosomes. Nat Cell Biol 2012;14:677–685.
Fevrier B, Raposo G . Exosomes: endosomal-derived vesicles shipping extracellular messages. Curr Opin Cell Biol 2004;16:415–421.
Hurley JH, Hanson PI . Membrane budding and scission by the ESCRT machinery: it’s all in the neck. Nat Rev Mol Cell Biol 2010;11:556–566.
Roma-Rodrigues C, Fernandes AR, Baptista PV . Exosome in tumour microenvironment: overview of the crosstalk between normal and cancer cells. Biomed Res Int 2014;2014:179486.
Ragusa M, Statello L, Maugeri M et al. Highly skewed distribution of miRNAs and proteins between colorectal cancer cells and their exosomes following Cetuximab treatment: biomolecular, genetic and translational implications. Oncoscience 2014;1:132–157.
Wang X, Ding X, Nan L et al. Investigation of the roles of exosomes in colorectal cancer liver metastasis. Oncol Rep 2015;33:2445–2453.
Ji H, Greening DW, Barnes TW et al. Proteome profiling of exosomes derived from human primary and metastatic colorectal cancer cells reveal differential expression of key metastatic factors and signal transduction components. Proteomics 2013;13:1672–1686.
Fan X, Ouyang N, Teng H et al. Isolation and characterization of spheroid cells from the HT29 colon cancer cell line. Int J Colorectal Dis 2011;26:1279–1285.
Feng HL, Liu YQ, Yang LJ et al. Expression of CD133 correlates with differentiation of human colon cancer cells. Cancer Biol Ther 2010;9:216–223.
Kai K, Nagano O, Sugihara E et al. Maintenance of HCT116 colon cancer cell line conforms to a stochastic model but not a cancer stem cell model. Cancer Sci 2009;100:2275–2282.
Lugli A, Iezzi G, Hostettler I et al. Prognostic impact of the expression of putative cancer stem cell markers CD133, CD166, CD44s, EpCAM, and ALDH1 in colorectal cancer. Br J Cancer 2010;103:382–390.
Vaiopoulos AG, Kostakis ID, Koutsilieris M et al. Colorectal cancer stem cells. Stem cells 2012;30:363–371.
Kahlert C, Kalluri R . Exosomes in tumor microenvironment influence cancer progression and metastasis. J Mol Med 2013;91:431–437.
Gangoda L, Boukouris S, Liem M et al. Extracellular vesicles including exosomes are mediators of signal transduction: are they protective or pathogenic? Proteomics 2015;15:260–271.
Grange C, Tapparo M, Collino F et al. Microvesicles released from human renal cancer stem cells stimulate angiogenesis and formation of lung premetastatic niche. Cancer Res 2011;71:5346–5356.
Valcz G, Sipos F, Tulassay Z et al. Importance of carcinoma-associated fibroblast-derived proteins in clinical oncology. J Clin Pathol 2014;67:1026–1031.
Räsänen K, Vaheri A . Activation of fibroblasts in cancer stroma. Exp Cell Res 2010;316:2713–2722.
Powell DW, Mifflin RC, Valentich JD et al. Myofibroblasts. II. Intestinal subepithelial myofibroblasts. Am J Physiol 1999;27:183–201.
Silva J, Garcia V, Rodriguez M et al. Analysis of exosome release and its prognostic value in human colorectal cancer. Genes Chromosomes Cancer 2012;51:409–418.
Taylor DD, Gercel-Taylor C . MicroRNA signatures of tumor-derived exosomes as diagnostic biomarkers of ovarian cancer. Gynecol Oncol 2008;110:13–21.
Rabinowits G, Gercel-Taylor C, Day JM et al. Exosomal microRNA: a diagnostic marker for lung cancer. Clin Lung Cancer 2009;10:42–46.
Jakobsen KR, Paulsen BS, Baek R et al. Exosomal proteins as potential diagnostic markers in advanced non-small cell lung carcinoma. J Extracell Vesicles 2015;4:26659.
Hurley JH, Odorizzi G . Get on the exosome bus with ALIX. Nat Cell Biol 2012;14:654–655.
Bissig C, Gruenberg J . ALIX and the multivesicular endosome: ALIX in Wonderland. Trends Cell Biol 2014;24:19–25.
Yi X, Bouley R, Lin HY et al. Alix (AIP1) is a vasopressin receptor (V2R)-interacting protein that increases lysosomal degradation of the V2R. Am J Physiol Renal Physiol 2007;292:1303–1313.
Tauro BJ, Greening DW, Mathias RA et al. Comparison of ultracentrifugation, density gradient separation, and immunoaffinity capture methods for isolating human colon cancer cell line LIM1863-derived exosomes. Methods 2012;56:293–304.
Johnson IR, Parkinson-Lawrence EJ, Keegan H et al. Endosomal gene expression: a new indicator for prostate cancer patient prognosis? Oncotarget 2015;6:37919–37929.
Duijvesz D, Burnum-Johnson KE, Gritsenko MA et al. Proteomic profiling of exosomes leads to the identification of novel biomarkers for prostate cancer. PloS One 2013;8:e82589.
Husi H, Skipworth RJ, Cronshaw A et al. Programmed cell death 6 interacting protein (PDCD6IP) and Rabenosyn-5 (ZFYVE20) are potential urinary biomarkers for upper gastrointestinal cancer. Proteomics Clin Appl 2015;9:586–596.
Bosman FT, Carneiro F, Hruban RH, Theise ND, WHO classification of tumours of the digestive system. International Agency for Research on Cancer, 4th edn 2010.
Galamb O, Wichmann B, Sipos F et al. Dysplasia-carcinoma transition specific transcripts in colonic biopsy samples. PloS One 2012;7:e48547.
Galamb O, Sipos F, Solymosi N et al. Diagnostic mRNA expression patterns of inflamed, benign, and malignant colorectal biopsy specimen and their correlation with peripheral blood results. Cancer Epidemiol Biomarkers Prev 2008;17:2835–2845.
Galamb O, Gyorffy B, Sipos F et al. Inflammation, adenoma and cancer: objective classification of colon biopsy specimens with gene expression signature. Dis Markers 2008;25:1–16.
Matsuyama T, Ishikawa T, Mogushi K et al. MUC12 mRNA expression is an independent marker of prognosis in stage II and stage III colorectal cancer. Int J Cancer 2010;127:2292–2299.
Hong Y, Ho KS, Eu KW et al. A susceptibility gene set for early onset colorectal cancer that integrates diverse signaling pathways: implication for tumorigenesis. Clin Cancer Res 2007;13:1107–1114.
Hong Y, Downey T, Eu KW et al. A ‘metastasis-prone’ signature for early-stage mismatch-repair proficient sporadic colorectal cancer patients and its implications for possible therapeutics. Clin Exp Metastasis 2010;27:83–90.
Sabates-Bellver J, Van der Flier LG, de Palo M et al. Transcriptome profile of human colorectal adenomas. Mol Cancer Res 2007;5:1263–1275.
Fleming M, Ravula S, Tatishchev SF et al. Colorectal carcinoma: pathologic aspects. J Gastrointest Oncol 2012;3:153–173.
Valcz G, Krenacs T, Sipos F et al. Lymphoid aggregates may contribute to the migration and epithelial commitment of bone marrow-derived cells in colonic mucosa. J Clin Pathol 2011;64:771–775.
Millard M, Pathania D, Shabaik Y et al. Preclinical evaluation of novel triphenylphosphonium salts with broad-spectrum activity. PloS One 2010;5:e13131.
Conover WJ . Practical Nonparametric Statistics, 3rd edn. John Wiley & Sons: New York, USA, 1999.
Team RC R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing: Vienna, Austria, 2015. Available at: http://www.R-project.org/.
Nishimura S, Wakabayashi N, Toyoda K et al. Expression of Musashi-1 in human normal colon crypt cells: a possible stem cell marker of human colon epithelium. Dig Dis Sci 2003;48:1523–1529.
Liotta LA, Kohn EC . The microenvironment of the tumour-host interface. Nature 2001;411:375–379.
Mueller MM, Fusenig NE . Friends or foes—bipolar effects of the tumour stroma in cancer. Nat Rev 2004;4:839–849.
Bosman FT, de Bruine A, Flohil C et al. Epithelial-stromal interactions in colon cancer. Int J Dev Biol 1993;37:203–211.
Valcz G, Patai AV, Kalmar A et al. Myofibroblast-derived SFRP1 as potential inhibitor of colorectal carcinoma field effect. PloS One 2014;9:e106143.
Hong BS, Cho JH, Kim H et al. Colorectal cancer cell-derived microvesicles are enriched in cell cycle-related mRNAs that promote proliferation of endothelial cells. BMC Genomics 2009;10:556.
Diehl HC, Stuhler K, Klein-Scory S et al. A catalogue of proteins released by colorectal cancer cells in vitro as an alternative source for biomarker discovery. Proteomics Clin Appl 2007;1:47–61.
Paggetti J, Haderk F, Seiffert M et al. Exosomes released by chronic lymphocytic leukemia cells induce the transition of stromal cells into cancer-associated fibroblasts. Blood 2015;126:1106–1117.
Bhattacharya S, Pal K, Sharma AK et al. GAIP interacting protein C-terminus regulates autophagy and exosome biogenesis of pancreatic cancer through metabolic pathways. PloS One 2014;9:e114409.
Hitchins MP, Lin VA, Buckle A et al. Epigenetic inactivation of a cluster of genes flanking MLH1 in microsatellite-unstable colorectal cancer. Cancer Res 2007;67:9107–9116.
Razi M, Futter CE . Distinct roles for Tsg101 and Hrs in multivesicular body formation and inward vesiculation. Mol Biol Cell 2006;17:3469–3483.
Bache KG, Brech A, Mehlum A et al. Hrs regulates multivesicular body formation via ESCRT recruitment to endosomes. J Cell Biol 2003;162:435–442.
Odorizzi G . The multiple personalities of Alix. J Cell Sci 2006;119:3025–3032.
Suetsugu A, Honma K, Saji S et al. Imaging exosome transfer from breast cancer cells to stroma at metastatic sites in orthotopic nude-mouse models. Adv Drug Deliv Rev 2013;65:383–390.
Shaw PJ, Rawlins DJ . The point-spread function of a confocal microscope: its measurement and use in deconvolution of 3-D data. J Microsc 1991;163:151–165.
Kumar D, Gupta D, Shankar S et al. Biomolecular characterization of exosomes released from cancer stem cells: possible implications for biomarker and treatment of cancer. Oncotarget 2015;6:3280–3291.
Azmi AS, Bao B, Sarkar FH . Exosomes in cancer development, metastasis, and drug resistance: a comprehensive review. Cancer Metastasis Rev 2013;32:623–642.
Brinton LT, Sloane HS, Kester M et al. Formation and role of exosomes in cancer. Cell Mol Life Sci 2015;72:659–671.
Corcoran C, Rani S, O’Brien K et al. Docetaxel-resistance in prostate cancer: evaluating associated phenotypic changes and potential for resistance transfer via exosomes. PloS One 2012;7:e50999.
Hu Y, Yan C, Mu L et al. Fibroblast-derived exosomes contribute to chemoresistance through priming cancer stem cells in colorectal cancer. PloS One 2015;10:e0125625.
Lee TH, Chennakrishnaiah S, Audemard E et al. Oncogenic ras-driven cancer cell vesiculation leads to emission of double-stranded DNA capable of interacting with target cells. Biochem Biophys Res Commun 2014;451:295–301.
Acknowledgements
This study was funded by the Research and Technology Innovation Fund, Hungary, KMR_12-1-2012-0216 and Hungarian Scientific Research Fund (OTKA-K111743 grant). We would like to thank to Marcell Szász for his help in sample collection. We also thank our assistants Gabriella Kónyáné Farkas (Cell Analysis Laboratory, 2nd Department of Medicine, Semmelweis University) and Zita Bratu (1st Department of Pathology and Experimental Cancer Research, Semmelweis University) for their hard work in this field. We also thank Zoltán Szállási and Theo deVos for language revision.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Modern Pathology website
Rights and permissions
About this article
Cite this article
Valcz, G., Galamb, O., Krenács, T. et al. Exosomes in colorectal carcinoma formation: ALIX under the magnifying glass. Mod Pathol 29, 928–938 (2016). https://doi.org/10.1038/modpathol.2016.72
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2016.72
This article is cited by
-
Light-emitting diode photomodulation of uterine adenocarcinoma cells inhibited angiogenesis capacity via the regulation of exosome biogenesis
Lasers in Medical Science (2022)
-
Global DNA hypomethylation of colorectal tumours detected in tissue and liquid biopsies may be related to decreased methyl-donor content
BMC Cancer (2022)
-
Focus on the morphogenesis, fate and the role in tumor progression of multivesicular bodies
Cell Communication and Signaling (2020)
-
Perspective: bidirectional exosomal transport between cancer stem cells and their fibroblast-rich microenvironment during metastasis formation
npj Breast Cancer (2018)