Abstract
The clinical course of prostate cancer is highly variable. Current prognostic variables, stage, and Gleason score have limitations in assessing treatment regimens for individual patients, especially in the intermediate-risk group of Gleason score 7. ERG:TMPRSS2 fusion and loss of PTEN are some of the most common genetic alterations in prostate cancer. Immunohistochemistry of PTEN and ERG has generated interest as a promising method for more precise outcome prediction but requires further validation in population-based cohorts. We studied the predictive value of ERG and PTEN expression by immunohistochemistry in two large radical prostatectomy cohorts comprising 815 patients with extensive follow-up information. Clinical end points were initiation of secondary therapy, overall survival, and disease-specific survival. Predictions of clinical outcomes were also assessed according to androgen receptor (AR) activity. PTEN loss, especially in ERG-negative cancers, predicted initiation of secondary treatments and shortened disease-specific survival time, as well as stratifying Gleason score 7 patients into different prognostic groups with regard to secondary treatments and disease-specific survival. High AR immunoreactivity in ERG-negative cancers with PTEN loss predicted worse disease-specific survival. We also observed that in Gleason score 7 ERG-negative cases with PTEN loss and high AR expression have significantly shorter disease-specific survival time compared with ERG-positive cases. Our conclusion is that loss of PTEN is a strong determining factor for shorter disease-specific survival time and initiation of secondary therapies after radical prostatectomy. The predictive value of PTEN immunoreactivity is further accentuated in ERG-negative cancers with high AR expression. Negative PTEN expression, accompanied by ERG status, can be used to stratify patients with Gleason score 7 into different survival groups. Assessment of PTEN and ERG status could provide an additional tool for initial diagnostics when determining the prognosis and subsequent follow-up regimen for patients treated by radical prostatectomy.
Similar content being viewed by others
Main
Prostate cancer is the most common malignant disease among men in the western world.1 Aside from its high incidence and prevalence, it is biologically a highly variable disease. A considerable proportion of cancers are clinically insignificant and do not cause any serious morbidity during the lifetime of an individual. However, some prostate cancers are highly aggressive, quickly progress to metastatic disease, and ultimately become treatment resistant, leading to cancer-specific death. Gleason grade and tumor stage are still the most widely accepted and clinically valuable tools for predicting disease-specific survival in prostate cancer. However, Gleason grading, even after having gone through modifications,2, 3 needs to be supplemented with, or accompanied by, new markers to differentiate between low-risk and high-risk diseases. This is especially important for Gleason score 7 disease, where prediction of clinical outcome is uncertain. An optimal predictive biomarker would stratify patients by treatment response and disease-specific mortality while also providing biologically relevant information for applicable therapy. Although knowledge of genomic markers in prostate cancer has greatly improved in recent years, there is still a lack of validated, reliable markers for use in clinical practice.4
Among the most prevalent genetic alterations in prostate cancer are the fusion of the androgen-regulated serine protease TMPRSS2 with the ets family transcription factor ERG and the inactivating rearrangements in the tumor suppressor gene PTEN (phosphatase and tensin homolog).5 TMPRSS2:ERG fusions occur frequently, presenting in 36–78% of patients, and appear to occur early in carcinogenesis.6, 7, 8 PTEN loss occurs in 16–47% of radical prostatectomy or biopsy specimens and has been shown to associate with adverse pathological findings and impaired survival, as well as the development of castration resistance and metastasis.7, 9, 10, 11 PTEN loss is a less prevalent, but potentially clinically more relevant genetic rearrangement than TMPRSS2:ERG fusions in prostate cancer. Immunohistochemical methods have proven reliable and useful in detecting the alterations in the expression of ERG protein, a surrogate marker for TMPRSS2:ERG fusion, and PTEN.8, 12, 13, 14, 15
ERG overexpression alone does not determine disease progression or patient outcome.12, 13, 16, 17 It may, however, indicate the predisposing effect of the gene fusion upon other cancer cell regulatory pathways such as loss of PTEN,7, 18, 19, 20, 21 and thus, may reflect disease aggressiveness. PTEN loss alone seems to22, 23 show some value in predicting patient outcomes.9, 15, 24, 25 However, the combined function of these genetic alterations— TMPRSS2:ERG fusion and PTEN deletion—apparently has more clinical significance than either one alone.18, 22, 23, 26 Reports of ERG fusion-positive–PTEN-negative prostate cancer suggest that this combination may lead to unfavorable disease outcomes such as earlier biochemical recurrence10, 21, 27 and cancer progression.28, 29 The opposite may also be true22, 23 and, to date, there are few studies with disease-specific survival as the prediction end point.
Prostate cancer is highly dependent upon androgen receptor (AR) pathway activity. The controversy in the findings of ERG regulating the AR pathway30, 31, 32 and the lack of PTEN and AR comparison in large human-derived clinical cohorts warrants further study. Here we sought to validate the prognostic significance of the ERG and PTEN expression in patients treated with radical prostatectomy and to study the association of AR expression with ERG and PTEN expressions in a large cohort with long follow-up time and disease-specific survival as the major end point.
Materials and methods
Study Population
As a substudy of FinnProstate IX, a national tissue microarray initiative, we utilized a clinical database and accompanying tissue microarray consisting of all patients treated with radical prostatectomy between 1983 and 1998 at Helsinki University Hospital and between 2000 and 2005 at Turku University Hospital. Limited pelvic lymph node dissection accompanied most of the prostatectomies in both cohorts. The Helsinki database with clinical preoperative and follow-up information was gathered originally in 2005 and the Turku database in 2010. The clinical databases were combined and updated in 2015 with the Finnish Cancer Registry's data on patients’ all-cause and disease-specific mortality, resulting in a median follow-up time of 15.7 years for the Helsinki cohort (range 0.7–28.6) and 9.5 years (range 0.2–14.0) for the Turku cohort. The Helsinki cohort comprised 478 patients of whom 452 had formalin- and paraffin-embedded prostatectomy blocks available for tissue microarray construction in 2010. The Turku cohort comprised 532 patients, all of whom had prostatectomy blocks available for tissue microarray construction in 2010. To avoid possible interference on marker status, only patients who had not received neoadjuvant therapies prior to surgery were selected for analysis, which led to 401 and 503 patients in the Helsinki and Turku cohorts, respectively. Because of incomplete tissue material or staining failure, the final analysis comprised a total of 358 patients in the Helsinki cohort and 457 patients in the Turku cohort. Postoperative follow-up was conducted by clinical examinations and prostate-specific antigen (PSA) measurements at least three times during the first year after surgery and at least once a year during the following years. Patient characteristics of both study populations are presented in Table 1. One hundred and thirty-one and 145 patients received secondary therapies after radical prostatectomy in Helsinki and Turku, respectively, according to clinical practices at that time. For the Helsinki cohort, full information on the commencement dates of secondary therapies was also available (Supplementary Table 1). As no significant differences were found between the cohorts in terms of the conventional clinical predictive markers, the cohorts were combined for survival analysis with regard to ERG and PTEN expressions.
The use of tissue material and corresponding clinicopathological data was approved by the ethics committees of the corresponding hospital districts (Hospital District of Helsinki and Uusimaa and Hospital District of Southwest Finland). Approval for the use of tissue materials was also obtained from the National Authority for Welfare and Health in Finland, according to national legislation, and use of the Finnish Cancer Registry’s data was approved by Finland’s National Institute for Health and Welfare. Patient's personal information was de-identified prior to analyses.
Tissue Microarray Construction
The prostatectomy samples were fixed in neutral buffered formalin (10%) prior to tissue processing and paraffin embedding. For construction of the tissue microarray blocks, histopathology was re-evaluated and Gleason scores assigned according to updated criteria2 from corresponding hematoxylin and eosin or herovici slides by experienced uropathologists (TM, SN, and AB in Helsinki and TM in Turku). The grading was further assessed according to the proposed new Gleason grade grouping.3 The pathologists also selected representative areas for the tissue microarray cores. The Helsinki tissue microarray blocks were constructed as previously described33 and consisted of four samples from each patient: two tissue cores from the area containing the most dominant Gleason grade pattern, one core out of the area with the secondary Gleason grade pattern and one core from an adjacent benign glandular area. The Turku tissue microarray blocks were constructed as previously described.34 Three to twelve (median three) adjacent cores 1 mm in diameter were drilled and transferred to the recipient block from the index carcinoma lesion (based on the Gleason grade pattern, the volume of cancer lesion, and/or extra-prostatic extension). In addition, one core from histologically benign prostate tissue was obtained from the same patient.
Immunohistochemistry
For the Helsinki cohort, freshly cut 4 μm thick tissue microarray sections, mounted on electrically charged glass slides (SuperFrost Plus, Menzel-Gläser, Braunschweig, Germany), were stained using Dako Autostainer (Dako Denmark A/S, Glostrup, Denmark) for ERG and PTEN. The immunohistochemistry was performed after heat-induced epitope retrieval on consecutive slides with 1:300 dilution for rabbit monoclonal ERG antibody (clone EPR 3864, Abcam PLC, Cambridge, UK) and with 1:100 dilution for rabbit monoclonal PTEN antibody (clone D4.3 XP, Cell Signaling Technology, Danvers, MA, USA). Directly after each staining, to prevent any effect of fading, the slides were digitized with an automated whole-slide scanner (Mirax Scan, Zeiss, Oberkochen, Germany) using a 20 × objective (numerical aperture 0.75) and a Sony DFW-X710 camera with a 1024 × 768 pixel CCD sensor (Sony Corporation, Tokyo, Japan). The pixel resolution was 0.26 μm. The images were compressed to a wavelet file format (Enhanced Compressed Wavelet, ECW, ER Mapper, Erdas, Norcross, GA, USA) with a compression ratio of 1:5. The compressed virtual slides were uploaded to a web-based scoring tool (WebMicroscope, Helsinki, Finland) running image server software (Image Web Server, Erdas, Norcross, GA, USA). Our immunohistochemistry method for AR detection has been reported earlier.33
The same antibodies and dilutions were used to evaluate ERG and PTEN expressions for the Turku cohort. Stainings were carried out using a Ventana automated staining machine (Ventana Medical Systems, Tucson, AZ, USA) and the primary antibodies were detected with Vectastain anti-rabbit HRP-conjugated secondary antibodies. The slides were counterstained with hematoxylin and analyzed directly after staining with Olympus BX60 microscope (Olympus Optical, Tokyo, Japan). Immunohistochemistry for AR expression status was not performed for the Turku cohort.
Marker Scoring
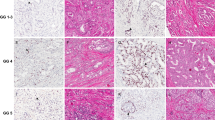
The expressions of ERG and PTEN in tissue cores were evaluated independently by investigators (KL and TM in Helsinki and IS and PT in Turku) blinded to the clinical and other pathological data. Internal staining controls were endothelium for ERG and benign prostatic epithelium for PTEN. Nuclear ERG expression was considered negative, weak, moderate, or strong (and later dichotomized as either negative or positive for statistical analyses), and cytoplasmic PTEN expression as either negative or positive. No clear cut off values were defined, but ERG was considered negative or weak if the staining was absent or markedly lower than in the endothelial cell nuclei. PTEN expression in an individual tissue microarray core was considered lost if the intensity of cytoplasmic staining was markedly lower than in the surrounding benign glands. On the patient level, ERG and PTEN scores were assessed separately for each core and the final analysis was made by comparing the scores pairwise similarly to a recent analysis by Ahearn et al.22 Complete or partial PTEN loss was assessed overall in all of the cores of an individual patient. PTEN loss was determined as either complete (all cores negative) or partial (any core negative). AR expression scores for the Helsinki cohort were adapted from our previous work with the same tissue microarrays, and were dichotomized to be either low or high.33 Representative sequential sections showing ERG, PTEN, and AR immunohistochemistry are shown in Figure 1.
Immunohistochemical staining on the consecutive tissue microarray sections for ERG, PTEN, and AR proteins. (a) Prostate cancer with negative PTEN staining. (b) Sequential section representing nuclear ERG immunostaining in cancerous glands. (c) Sequential section showing high nuclear AR expression in the cancerous glands. (d) Another example of PTEN loss in prostate cancer glandular structures. (e) Subsequent section for the sample in figure (d) lacking ERG nuclear expression. (f) Same spot as in figures (d and e) showing high AR expression.
Statistical Analysis
χ2 and Fisher's exact tests were employed, when appropriate, to investigate the correlations between ERG and PTEN expressions and clinical variables. Kaplan–Meier survival analyses with Mantel-Cox log-rank statistics were conducted to compare different marker expressions for their effect on disease-specific survival, overall survival, and for the Helsinki cohort, secondary therapy-free survival. Uni- and multivariate Cox regression analyses were performed to determine the effects of marker status (ERG positivity, complete loss of PTEN, and strong AR expression) and standard variables (age at operation, preoperative PSA, pT stage, Gleason score, and lymph node status) on disease-specific mortality, overall mortality, and receiving secondary therapies. ERG expression was used as a stratification variable, because it was non-proportional when checking for proportional hazards. All statistical analyses were performed with IBM SPSS Statistics 23 (IBM, Chicago, IL, USA). Statistical significance was deemed at P<0.05 in two-sided tests.
Results
During follow-up, there were 33 (9.2%) prostate cancer-related deaths in the Helsinki cohort and 19 (4.2%) in the Turku cohort among the patients with full ERG and PTEN status available. All-cause mortality in the combined data was 30.0% (48.0% in Helsinki and 16.0% in Turku) and disease-specific mortality was 6.4% during a median follow-up of 11.9 years. The most common Gleason score in the prostatectomy specimens in both cohorts was 7 (49.6%). For detailed patient characteristics and follow-up information, see Table 1.
In the crosstabs analyses, PTEN loss was significantly associated with higher preoperative PSA, higher Gleason score, higher pT stage, positive lymph nodes, poorer overall survival and disease-specific survival, and increased likelihood for receiving secondary treatments. On the other hand, positive ERG status correlated only with lower preoperative PSA and increased likelihood for receiving secondary therapies but not with disease-specific survival or overall survival (Supplementary Table 2). Kaplan–Meier analysis demonstrated that ERG status alone was not associated with overall survival, disease-specific survival, or time to secondary therapy, nor were there any significant associations between PTEN status alone and overall survival (data not shown). PTEN loss was statistically significantly associated with shorter disease-specific survival time (Figures 2a and b). Complete loss of PTEN expression, compared with intact or partially lost PTEN, raised the risk of prostate cancer death with a hazard ratio (HR) of 2.156 (95% CI 1.169–3.976, P=0.014) in univariate Cox regression analysis, although not in multivariate analysis (Supplementary Table 3). PTEN loss alone did not associate with shorter disease-specific survival time in the individual grade groups or Gleason scores, nor in the higher grade groups of 3–5 combined (data not shown).
Considering the recent findings of ERG expression-related differences in prediction of lethal cancer by Ahearn et al,22 we tested the association of combined ERG/PTEN expression status with the likelihood of receiving secondary treatment after radical prostatectomy. The original treatment decisions for these patients had been made by the treating urologists, reflecting clinical practice at that time (Supplementary Table 1). Loss of PTEN was associated with shorter secondary therapy-free survival in both ERG-positive and ERG-negative patients (Figure 3a). In logistic regression analyses, the HR for commencing secondary treatment was 2.782 (95% CI 1.846–4.193, P<0.001) for complete PTEN loss in univariate analysis and 2.290 (95% CI 1.314–3.989, P=0.003) in multivariate analysis, when compared with intact or partially lost PTEN (Supplementary Table 3). Complete PTEN loss in ERG-negative patients was significantly associated with shorter disease-specific survival time when compared with ERG-positive patients with PTEN intact or only partially lost (P=0.024, Figure 3b). Any PTEN loss differed from intact PTEN in terms of disease-specific survival, but only in ERG-negative patients (P=0.021).
In order to assess the utility of ERG/PTEN status in stratifying patients with intermediate grade prostate cancer (Gleason score 7 or grade group 3), we tested complete or partial PTEN loss in ERG-negative and -positive cases. Kaplan–Meier survival analysis showed that ERG-negative patients with complete PTEN loss had the poorest disease-specific survival in the Gleason score 7 (P=0.008, Figure 4a) and grade group 3 (P=0.013, Figure 4b) subgroups.
In univariate Cox regression analysis strong AR expression raised the risk of prostate cancer death with a HR of 2.375 (95% CI 1.008–5.598, P=0.048) (Supplementary Table 3). As recent studies provide evidence that ERG-negative prostate cancer with PTEN loss retains androgen activity, we wanted to test the effect of AR expression in ERG-positive and ERG-negative cancers with PTEN loss. Half of the ERG-negative patients had high AR expression (92/177, 52.0%) compared with 76.8% (139/181) of the ERG-positive patients. In Kaplan–Meier analysis, ERG-negative patients with high AR expression had significantly shorter disease-specific survival time than ERG-negative patients with low AR expression (Figure 5a, P=0.036). This shorter disease-specific survival time among ERG-negative patients with high AR was further accentuated by complete loss of PTEN (Figure 5b, P=0.012) and a similar effect was observed in the subanalysis of Gleason score 7 patients (Figure 5c, P=0.036). For ERG-positive patients, AR status did not determine survival differences (P=0.485), nor were there differences among patients with complete PTEN loss in terms of AR status-determined disease-specific survival (P=0.226).
(a) AR expression defines disease-specific survival time after radical prostatectomy in ERG-negative patients (n=177). (b) In the patients with high AR expression ERG/PTEN status determines different survival probabilities (n=133). (c) In the subanalysis of Gleason score 7 patients AR expression status combined with ERG/PTEN status determines different survival probabilities (n=81).
Discussion
Promising predictive biomarkers that would complement histopathological evaluation in prostate cancer are regularly discovered, but thus far, only with conflicting results in terms of clinical relevance. TMPRSS2:ERG is the most prevalent known fusion gene in the development of prostate cancer. Fusion occurs either through insertional chromosomal arrangement or intrachromosomal deletion. A recent animal model study by Linn et al35 found that in the background of PTEN deficiency, ERG fusion through deletion drives prostate cancer to poor differentiation and epithelial–mesenchymal transition, a much debated phenomenon in human prostate cancer. They also found that exclusively intrachromosomal deletion was associated with lethal features in clinical cancer through the loss of tumor suppressor genes. These tumors in the animal model seem to retain AR activity but lose the protein expression of ERG, contrary to what is seen in human tumors with known ERG fusion. In this study we validated the effects of PTEN loss, combined with ERG and AR status, in the lethal progression of prostate cancer in a clinical patient cohort with sufficient follow-up time.
Earlier, we found that FOXA1 regulates AR-related progression of prostate cancer through cistrome reprogramming.33 Linn et al35 also speculated, based on their findings, that AR function is increased through molecular rearrangements or cistrome reprogramming. Our current results show high AR expression in ERG-negative prostate cancer with PTEN loss. Furthermore, these patients have a poorer prognosis compared with ERG-negative-PTEN intact- or ERG-positive–PTEN-negative patients. This suggests that AR-regulated pathways are maintained, or even promoted, in highly malignant prostate cancer, regardless of ERG fusion status. Our results also concur with earlier findings suggesting poorer response to conventional androgen-deprivation therapy or novel antiandrogen therapy in men with ERG-negative lesions when compared with men with ERG-positive disease.36, 37 These results raise the question of whether FOXA1 and other AR regulators are drivers of progression within ERG-negative–PTEN-negative prostate cancer with high AR activity. The functional mechanisms by which ERG fusion-negative tumors with PTEN loss promote cancer lethality remain unsolved at this time. Possibly other competing mechanisms of AR-signaling activation have a stronger influence during later stages of disease progression, as ERG fusion is a relatively early event in prostate cancer pathogenesis.
Notably, our ERG and AR antibodies are the same as used in the studies by Ahearn et al22 and Linn et al.35 However, neither our study nor Ahearn's22 was able to show whether negative ERG protein expression in fact reflects fusion by intrachromosomal deletion, which may lead to AR overexpression through mechanisms yet poorly characterized. In addition, type of ERG activation and possible different prognostic correlations were not assessed in our study. The antibody used by us and by Ahearn et al22 should detect the ERGα subtype that is generated by both types of fusion. It is evident that this kind of subanalysis for the ERG fusion-negative cases with impaired prognosis will ultimately be needed and the recent findings from animal models warrant validation in large-scale human cohorts.
Our results strongly support the notion that TMPRSS2:ERG fusion is an early event in carcinogenesis and does not, as such, correlate with lethal prostate cancer. While we were conducting this study, Ahearn et al22 published prediction results similar to ours in their surgically treated cohort. The main difference between the studies is that our survival analysis for lethal disease stringently included only patients who had died of prostate cancer, whereas in Ahearn's22 study a lethal event was defined as cancer death or metastasis. The fact that our results simultaneously confirm their findings of ERG-negative–PTEN-negative patients having the poorest prognosis is relevant and raises the question whether the same predictive power also applies for diagnostic samples. The shorter time until secondary therapies after radical prostatectomy associated with PTEN loss, irrespective of other pathological parameters, suggests that PTEN loss is a strong driver of disease progression and warrants a future study in diagnostic biopsies coupled to different treatment regimens. Based on our data and earlier reports, it is becoming evident that PTEN loss is a more important driver of progression than ERG fusion. Our study is, to our knowledge, the first to show that high AR activity in PTEN-negative prostate cancer determines shorter disease-specific survival time, especially in ERG fusion-negative cases. A possible explanation is that ERG actually downregulates AR activity and, when the fusion is not present, other factors, especially ETV1, promote AR transcription in patients with PTEN loss.30
The current evidence of ERG and PTEN protein expression in prostate cancer may help clinicians in assessing the need for postoperative adjuvant therapies after radical prostatectomy. This may be especially helpful in patients with Gleason score 7 disease, a group of patients who often poses a challenge in clinical decision-making. In our study cohorts, as in most contemporary radical prostatectomy cohorts, Gleason score 7 patients are the most prevalent group. Our results confirm the utility of immunohistochemistry for detecting ERG overexpression and PTEN loss as a determinant of lethal disease. We also found that in the intermediate group of Gleason score 7 it is relevant to consider stratification by ERG and PTEN status. However, further studies in cohorts with information on second-line treatment regimens and large cohorts of prostatectomy-coupled biopsies are needed to test the value of these markers in clinical practice and decision-making.
Although our study is retrospective in nature and utilizes a historical cohort, its strengths are the comprehensive mortality data and long follow-up time necessary for clinically relevant end points such as secondary treatments and mortality. The Helsinki cohort comprises patients diagnosed before PSA screening was introduced and who, thus, tend to have had more advanced disease at the time of surgery than the Turku patients. This difference is reflected in the Helsinki cohort’s higher preoperative PSA levels and Gleason scores and is possibly why PTEN expression was found more often to be intact for Turku patients. However, in terms of pT stage and lymph node status, perhaps the strongest conventional determinants of disease outcome, the two cohorts are remarkably similar. Complete loss of PTEN expression showed to be significantly predictive of disease-specific survival in both cohorts independently, so we found it appropriate to combine these two large patient cohorts for more robust analyses.
Further studies assessing the value of an ERG/PTEN-double marker panel combined with AR expression level as a tool for planning possible adjuvant therapies and tailored follow-up are warranted. Furthermore, an additional approach could be to study the ERG/PTEN status in diagnostic prostate biopsies to better identify, eg, whether patients are suitable for active surveillance due to higher risk of disease progression.
In conclusion, our results suggest that prostate cancer patients with PTEN loss might be candidates for adjuvant therapies after radical prostatectomy. It is probable that regulators of the AR pathway other than ERG fusion determine dedifferentiation and lethal progression in prostate cancer with PTEN loss. The relatively simple and accurate immunohistochemical detection of ERG, PTEN, and AR expressions holds strong promise in stratifying patients into relevant risk groups of lethal disease.
References
Center MM, Jemal A, Lortet-Tieulent J et al.International variation in prostate cancer incidence and mortality rates. Eur Urol 2012;61:1079–1092.
Epstein JI . An update of the Gleason grading system. J Urol 2010;183:433–440.
Epstein JI, Egevad L, Amin MB et al.The 2014 International Society of Urological Pathology (ISUP) Consensus Conference on Gleason Grading of Prostatic Carcinoma: definition of Grading Patterns and Proposal for a New Grading System. Am J Surg Pathol 2016;40:244–252.
Bostrom PJ, Bjartell AS, Catto JW et al.Genomic predictors of outcome in prostate cancer. Eur Urol 2015;68:1033–1044.
Zhao L, Yu N, Guo T et al.Tissue biomarkers for prognosis of prostate cancer: a systematic review and meta-analysis. Cancer Epidemiol Biomarkers Prev 2014;23:1047–1054.
Tomlins SA, Rhodes DR, Perner S et al.Recurrent fusion of TMPRSS2 and ETS transcription factor genes in prostate cancer. Science 2005;310:644–648.
Krohn A, Freudenthaler F, Harasimowicz S et al.Heterogeneity and chronology of PTEN deletion and ERG fusion in prostate cancer. Mod Pathol 2014;27:1612–1620.
Shah RB, Bentley J, Jeffery Z et al.Heterogeneity of PTEN and ERG expression in prostate cancer on core needle biopsies: implications for cancer risk stratification and biomarker sampling. Hum Pathol 2015;46:698–706.
Krohn A, Diedler T, Burkhardt L et al.Genomic deletion of PTEN is associated with tumor progression and early PSA recurrence in ERG fusion-positive and fusion-negative prostate cancer. Am J Pathol 2012;181:401–412.
Fontugne J, Lee D, Cantaloni C et al.Recurrent prostate cancer genomic alterations predict response to brachytherapy treatment. Cancer Epidemiol Biomarkers Prev 2014;23:594–600.
Mithal P, Allott E, Gerber L et al.PTEN loss in biopsy tissue predicts poor clinical outcomes in prostate cancer. Int J Urol 2014;21:1209–1214.
Minner S, Enodien M, Sirma H et al.ERG status is unrelated to PSA recurrence in radically operated prostate cancer in the absence of antihormonal therapy. Clin Cancer Res 2011;17:5878–5888.
Pettersson A, Graff RE, Bauer SR et al.The TMPRSS2:ERG rearrangement, ERG expression, and prostate cancer outcomes: a cohort study and meta-analysis. Cancer Epidemiol Biomarkers Prev 2012;21:1497–1509.
Lotan TL, Gurel B, Sutcliffe S et al.PTEN protein loss by immunostaining: analytic validation and prognostic indicator for a high risk surgical cohort of prostate cancer patients. Clin Cancer Res 2011;17:6563–6573.
Chaux A, Peskoe SB, Gonzalez-Roibon N et al.Loss of PTEN expression is associated with increased risk of recurrence after prostatectomy for clinically localized prostate cancer. Mod Pathol 2012;25:1543–1549.
Xu B, Chevarie-Davis M, Chevalier S et al.The prognostic role of ERG immunopositivity in prostatic acinar adenocarcinoma: a study including 454 cases and review of the literature. Hum Pathol 2014;45:488–497.
Hoogland AM, Jenster G, van Weerden WM et al.ERG immunohistochemistry is not predictive for PSA recurrence, local recurrence or overall survival after radical prostatectomy for prostate cancer. Mod Pathol 2012;25:471–479.
Carver BS, Tran J, Gopalan A et al.Aberrant ERG expression cooperates with loss of PTEN to promote cancer progression in the prostate. Nat Genet 2009;41:619–624.
Gumuskaya B, Gurel B, Fedor H et al.Assessing the order of critical alterations in prostate cancer development and progression by IHC: further evidence that PTEN loss occurs subsequent to ERG gene fusion. Prostate Cancer Prostatic Dis 2013;16:209–215.
Leinonen KA, Saramaki OR, Furusato B et al.Loss of PTEN is associated with aggressive behavior in ERG-positive prostate cancer. Cancer Epidemiol Biomarkers Prev 2013;22:2333–2344.
Yoshimoto M, Joshua AM, Cunha IW et al.Absence of TMPRSS2:ERG fusions and PTEN losses in prostate cancer is associated with a favorable outcome. Mod Pathol 2008;21:1451–1460.
Ahearn TU, Pettersson A, Ebot EM et al.A prospective investigation of PTEN loss and ERG expression in lethal prostate cancer. J Natl Cancer Inst 2016;108.
Reid AH, Attard G, Ambroisine L et al.Molecular characterisation of ERG, ETV1 and PTEN gene loci identifies patients at low and high risk of death from prostate cancer. Br J Cancer 2010;102:678–684.
Barnett CM, Heinrich MC, Lim J et al.Genetic profiling to determine risk of relapse-free survival in high-risk localized prostate cancer. Clin Cancer Res 2014;20:1306–1312.
Cuzick J, Yang ZH, Fisher G et al.Prognostic value of PTEN loss in men with conservatively managed localised prostate cancer. Br J Cancer 2013;108:2582–2589.
Chen Y, Chi P, Rockowitz S et al.ETS factors reprogram the androgen receptor cistrome and prime prostate tumorigenesis in response to PTEN loss. Nat Med 2013;19:1023–1029.
Grupp K, Kohl S, Sirma H et al.Cysteine-rich secretory protein 3 overexpression is linked to a subset of PTEN-deleted ERG fusion-positive prostate cancers with early biochemical recurrence. Mod Pathol 2013;26:733–742.
Han B, Mehra R, Lonigro RJ et al.Fluorescence in situ hybridization study shows association of PTEN deletion with ERG rearrangement during prostate cancer progression. Mod Pathol 2009;22:1083–1093.
Sowalsky AG, Ye H, Bubley GJ et al.Clonal progression of prostate cancers from Gleason grade 3 to grade 4. Cancer Res 2013;73:1050–1055.
Baena E, Shao Z, Linn DE et al.ETV1 directs androgen metabolism and confers aggressive prostate cancer in targeted mice and patients. Genes Dev 2013;27:683–698.
Huang KC, Alshalalfa M, Hegazy SA et al.The prognostic significance of combined ERG and androgen receptor expression in patients with prostate cancer managed by androgen deprivation therapy. Cancer Biol Ther 2014;15:1120–1128.
Qu X, Jeldres C, Glaskova L et al.Identification of combinatorial genomic abnormalities associated with prostate cancer early recurrence. J Mol Diagn 2016;18:215–224.
Sahu B, Laakso M, Ovaska K et al.Dual role of FoxA1 in androgen receptor binding to chromatin, androgen signalling and prostate cancer. EMBO J 2011;30:3962–3976.
Mirtti T, Leiby BE, Abdulghani J et al.Nuclear Stat5a/b predicts early recurrence and prostate cancer-specific death in patients treated by radical prostatectomy. Hum Pathol 2013;44:310–319.
Linn DE, Penney KL, Bronson RT et al.Deletion of interstitial genes between TMPRSS2 and ERG promotes prostate cancer progression. Cancer Res 2016;76:1869–1881.
Attard G, de Bono JS, Logothetis CJ et al.Improvements in radiographic progression-free survival stratified by ERG gene status in metastatic castration-resistant prostate cancer patients treated with abiraterone acetate. Clin Cancer Res 2015;21:1621–1627.
Graff RE, Pettersson A, Lis RT et al.The TMPRSS2:ERG fusion and response to androgen deprivation therapy for prostate cancer. Prostate 2015;75:897–906.
Acknowledgements
We thank Tiina Vesterinen and Sinikka Collanus for valuable technical and administrative assistance and Kristina Mirtti for language-editing assistance. This study was supported by the Päivikki and Sakari Sohlberg Foundation, the Finnish Urological Association, Cancer Society of Finland, and Academy of Finland (grant #268531).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Modern Pathology website
Rights and permissions
About this article
Cite this article
Lahdensuo, K., Erickson, A., Saarinen, I. et al. Loss of PTEN expression in ERG-negative prostate cancer predicts secondary therapies and leads to shorter disease-specific survival time after radical prostatectomy. Mod Pathol 29, 1565–1574 (2016). https://doi.org/10.1038/modpathol.2016.154
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2016.154
This article is cited by
-
Poly(ADP-Ribose) Polymerase Inhibitors in Prostate Cancer: Molecular Mechanisms, and Preclinical and Clinical Data
Targeted Oncology (2020)
-
Clinical implications of PTEN loss in prostate cancer
Nature Reviews Urology (2018)
-
Association of ERG/PTEN status with biochemical recurrence after radical prostatectomy for clinically localized prostate cancer
Medical Oncology (2018)
-
ITGB1-dependent upregulation of Caveolin-1 switches TGFβ signalling from tumour-suppressive to oncogenic in prostate cancer
Scientific Reports (2018)