Abstract
TMPRSS2–ERG gene rearrangement is seen in about half of clinically localized prostate cancers, yet controversy exists with regard to its prognostic implications. Similarly, the relationship of TMPRSS2–ERG fusion to Gleason score and morphology remains uncertain. We assigned Gleason scores and recorded morphological features for 521 clinically localized prostate cancers sampled in triplicate and arrayed in eight tissue microarray blocks. Fluorescence in situ hybridization was performed to delineate TMPRSS2–ERG aberrations. Using maximum Gleason score, based on three core evaluation, and overall Gleason score, based on prostatectomy sections, Fisher's exact test was performed for tumors with TMPRSS2–ERG translocation/deletion, copy number increase (≥3) of the TMPRSS2–ERG region without translocation/deletion, and copy number increase and concomitant translocation/deletion. In all, 217 (42%) translocation/deletion and 30 (5.9%) copy number increase-alone cases were detected. Among 217 translocation/deletion cases, 32 had translocation/deletion with copy number increase. In all, 237, 200, and 75 cancers had maximum core-specific Gleason score of 6, 7, and 8–10, respectively. Tumors with translocation/deletion tended toward lower Gleason scores than those without (P=0.002) with similar results for overall Gleason score (P=0.02); copy number increase cases tended toward higher Gleason scores than those without (P<0.001). Gleason score of 8–10 tumors demonstrated lower odds of translocation/deletion (odds ratio (OR) 0.38; 95% CI 0.21–0.68) and higher odds of copy number increase alone (OR 7.33; 95% CI 2.65–20.31) or copy number increase+translocation/deletion (OR 3.03; 95% CI 1.12–8.15) relative to Gleason score of <7 tumors. No significant difference in TMPRSS2–ERG incidence was observed between patients with and without cribriform glands, glomerulations, signet-ring cells, or intraductal cancer (P=0.821, 0.095, 0.132, 0.375). TMPRSS2–ERG gene fusion is associated with lower core-specific and overall Gleason scores and not with high-grade morphologies. Conversely, TMPRSS2–ERG copy number increase, with or without rearrangement, is associated with higher Gleason score. These findings indicate that translocation/deletion of TMPRSS2–ERG is not associated with histological features of aggressive prostate cancer.
Similar content being viewed by others
Main
In the nearly 5 years since Tomlins et al1 reported the discovery of a recurrent translocation involving TMPRSS2 and ERG on chromosome 21 in prostate cancer, numerous groups have examined these findings in various patient populations,2, 3, 4 disease states,4, 5, 6, 7, 8, 9 and morphological contexts.10, 11, 12, 13 Although various frequencies of TMPRSS2–ERG gene fusion, ranging from 15–30% in conservatively managed European-based cohorts2, 3 to 70–80% in initial small selected cohorts obtained using PCR-based detection1, 14 have been reported, some consistency has emerged in clinically localized prostate cancer, with most groups finding ∼40–55% of cancers with this genetic change.5, 8, 13, 15, 16, 17, 18, 19 Frustratingly however, little consensus has been achieved with regard to the clinical significance of TMPRSS2–ERG rearrangement. Earlier studies reported associations with high stage, metastasis, and prostate cancer-specific death,3, 4, 8, 17 whereas more recent reports have found no association with outcome,5 an association with favorable outcome,9 or a similar percentage of TMPRSS2–ERG gene fusion in minute and nonminute adenocarcinomas,15 all suggesting its lack of value as a marker of aggressive prostate cancer.
In an analogous manner, examination of the relationship between TMPRSS2–ERG fusion and morphological manifestations of prostate cancer has produced varying results.2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 16, 18, 20, 21 Specifically, the majority of authors have found no statistically significant association between TMPRSS2–ERG rearrangement and Gleason score, the primary grading parameter in prostate cancer,4, 6, 8, 9, 12, 13, 20, 21 while some have demonstrated an association with either higher2, 3, 18 or lower Gleason scores.5, 16 Although fewer in number, studies of the association of TMPRSS2–ERG and specific prostate cancer morphological features have generally shown a correlation with high-grade histologies.7, 12, 13 Given the ongoing lack of consensus with regard to these findings, we investigated the relationship of TMPRSS2–ERG rearrangement with both tissue microarray core-specific and overall radical prostatectomy Gleason score, as well as a group of familiar prostate cancer morphologies.
Materials and methods
Case Selection
The patient cohort was comprised of prostate cancer tissue from 544 patients who underwent radical prostatectomy for clinically localized disease without previous androgen deprivation or radiation therapy at our institution between 1985 and 2002 and in whom extensive clinical follow-up was available. These cases had previously been sampled in triplicate and arrayed in eight tissue blocks. The cores represented in the tissue microarrays were from three different sites within the index tumor focus in the majority of cases.
Among these 544 cancers, 23 were not evaluable by fluorescence in situ hybridization (FISH) due to the absence of the tissue core on the section, no carcinoma present in the core, or a failed FISH assay, leaving 521 patients for analysis. For evaluation of maximum (core-specific) Gleason score only, an additional nine patients had either no tissue core present on the section or no carcinoma present in the core, resulting in only 512 patients' data being applicable in the related analyses.
FISH for Detection of Gene Fusion Status
This cohort of cases were previously studied5 using a three-color break-apart FISH probe set prepared by combining two BAC clones at 3′ ERG (RP11-315E22 and RP11-720N21) labeled with SpectrumOrange, with two BAC clones each at 5′ TMPRSS2 (RP11-35C4 and RP-891L10) labeled with SpectrumGreen, and 3′ TMPRSS2 (RP11-825A8 and RP11-120C17) labeled with SpectrumRed (Vysis, Abbott Molecular, Des Plaines, IL, USA). For each slide, 50–100 ng of each labeled clone was ethanol precipitated with 2–3 μg Cot-1 DNA and resuspended in 15 μl hybridization buffer.22
Hybridization, washing, and fluorescence detection were performed according to standard procedures. Briefly, sections were deparaffinized in xylenes, microwaved in 10 mM sodium citrate (pH 6.0) solution for 5–10 min, cooled to room temperature, rinsed, and then treated with pepsin–HCl for ∼5 min at 37°C before being rinsed and dehydrated. The prewarmed probe mixture was applied to the slides, and a coverslip was sealed in place with rubber cement. The slides were then denatured at 83°C for 4–6 min on a HYBrite automated hybridizer (Vysis, Abbott Molecular) and then incubated overnight at 37°C. After standard post-hybridization washes, the slides were stained with 4′,6-diamidino-2-phenylindole and mounted in antifade (Vectashield, Vector Laboratories).
Image Analysis
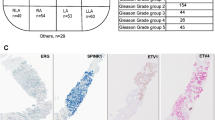
Samples were analyzed using an automated imaging system (MetaSystems, Altlussheim, Germany) and Isis 5.0 scanning and imaging software. Slides were scanned at × 5, and the resulting composite was segmented using the Metafer microarray tool. Segmentation generated a position list corresponding to each available core, linking slide location to subsequent high-resolution FISH images. Evaluation and analysis of the cases was carried out by a pathologist (AG) and a molecular cytogeneticist (MAL). A minimum of 100 cancer cells were evaluated for each case, whenever possible. If there was only one core that was positive, then the rearrangement status was recorded as positive for that case. This three-color experiment, incorporating 3′ ERG and 3′ and 5′ TMPRSS2 clones was used to facilitate the screening of a large sample size. Four major patterns were observed with FISH: (a) Wild type, with two sets of triplet orange (3′ ERG), red (3′ TMPRSS2), and green (5′ TMPRSS2) signals in each cell; (b) Deletion, with one orange-green doublet and loss of corresponding red signal in one allele and a wild-type second allele; (c) Translocation, with an orange-green doublet in one allele and retention, yet separation of one red (3′ TMPRSS2) signal; the other allele is wild type; (d) Copy number increase, with >2 wild-type signals in each cell (Figures 1a–d).
Four major FISH patterns: (a) Wild type, two sets of triplet orange (3′ ERG), red (3′ TMPRSS2), and green (5′ TMPRSS2) signals in each cell; (b) Deletion, one orange-green doublet and loss of corresponding red signal in one allele and a wild-type second allele; (c) Translocation, an orange-green doublet in one allele and retention, yet separation of one red (3′ TMPRSS2) signal; the other allele is wild type; (d) Copy number increase, greater than two wild-type signals in each cell.
Gleason Score and Morphological Assessment
A Gleason score (primary+secondary grade) was assigned for each evaluable core on eight tissue microarrays (core-specific Gleason score) by a single urological pathologist (SWF). The Gleason score assigned to the radical prostatectomy specimen, based on review of tissue sections (overall Gleason score), was also recorded. In addition, the presence of seven specific morphological features were recorded for each tissue microarray core, including cribriform glands, glomerulations, signet-ring cells, intraductal carcinoma, mucinous fibroplasia (collagenous micronodules), mucinous features (characterized by extravasated mucin) (Figures 2a–f), and foamy gland histology.
Statistical Analysis
Although a single FISH result was previously reported for each tumor,5 three core-specific Gleason scores were available for each patient. For analysis, Gleason score was grouped into clinically relevant categories (Gleason score <7, 7, and >7) and the maximum Gleason score from the three cores was used. Fisher's exact test was used to evaluate the significance of the relationship between the maximum Gleason score from the three cores and TMPRSS2–ERG aberration status. Odds ratio (OR) and 95% CI were reported using Gleason score <7 as the baseline for presence/absence of translocation/deletion, presence/absence of copy number increase alone, and presence/absence of copy number increase+translocation/deletion. For assessment of morphological features, any histological element observed was analyzed, such that if a case displayed both cribriform glands and glomerulations, it was counted independently in the analysis for each of these features. Similarly, Fisher's exact test was used to evaluate the significance of the relationship between morphology and the presence of translocation/deletion. Owing to the exploratory nature of these analyses, tests with P-values <0.05 were declared to be statistically significant without pursuing any adjustment for multiple comparisons. This study was approved by the Memorial Sloan-Kettering Cancer Center Institutional Review Board.
Results
The results of FISH analysis and both maximum and overall Gleason score are detailed in Tables 1 and 2. In total, 217 of 521 (42%) prostate cancers had TMPRSS2–ERG alterations (translocation/deletion), with 128 of these 217 (59%) displaying intergenic deletion. Copy number increase of TMPRSS2–ERG was present in 62 of 521 (12%) cases—concomitantly with rearrangement (copy number increase+translocation/deletion) in 32 (6.1%) cases, and without rearrangement (copy number increase alone) in the remaining 30. The maximum core-specific Gleason score was 6 for 237 patients, 7 for 200 patients, and 8–10 for 75 patients (Gleason score 8, 63; 9, 11; 10, 1).
Association Between FISH Results and Gleason Score
The relationships between (a) translocation/deletion, (b) copy number increase alone, (c) copy number increase+translocation/deletion and Gleason score were evaluated using Fisher's exact test and are detailed in Table 1.
Translocation/deletion
In all, 108 (51%), 89 (41%), and 18 (8%) of the 215 cases with translocation/deletion had core-specific Gleason score <7, 7, and >7, respectively, compared with 129 (43%), 111 (37%), and 57 (19%) of the 297 cases without translocation/deletion, indicating that tumors with TMPRSS2–ERG rearrangement tended to have lower Gleason scores than those without (P=0.002). A similar result was seen when analyzing for overall Gleason score (P=0.02) (Table 2).5
Copy number increase
Tumors with copy number increase alone tended to have significantly higher Gleason scores than those without (P<0.001).
Copy number increase+translocation/deletion
Tumors with copy number increase+translocation/deletion did not exhibit a linear trend in terms of Gleason score, likely because of the small number of tumors exhibiting this combination of FISH results. In particular, 28% of tumors with copy number increase+translocation/deletion have Gleason score <7, 47% Gleason score 7, and 25% Gleason score >7. The difference in Gleason score between tumors having copy number increase+translocation/deletion and those without was marginally significant (P=0.052).
The OR and 95% CI of presence or absence of (a) translocation/deletion, (b) copy number increase, and (c) copy number increase+translocation/deletion are summarized in Table 3.
Translocation/deletion
The odds of having TMPRSS2–ERG rearrangement among tumors with Gleason score 7 relative to the odds of having translocation/deletion among Gleason score 6 tumors was 0.96. In contrast, the OR of having TMPRSS2–ERG rearrangement among tumors with Gleason score 8–10 relative to Gleason score 6 cancers was 0.38, which is considerably less likely.
Copy number increase
The odds of having copy number increase alone among tumors with Gleason score 8–10 compared with Gleason score 6 tumors was 7.33, which is considerably more likely.
Copy number increase+translocation/deletion
Akin to copy number increase alone, the OR of having copy number increase+translocation/deletion in Gleason score 8–10 cancers relative to those with Gleason score 6 was 3.03.
Association Between FISH Results and Prostate Cancer Morphologies
The relationship between TMPRSS2–ERG rearrangement and specific morphological features was evaluated using Fisher's exact test and are detailed in Table 4. No significant difference was observed in the proportion of translocation/deletion between patients with and without cribriform glands, glomerulations, signet-ring cells, and intraductal carcinoma (P=0.821, 0.095, 0.132, and 0.375, respectively), all are considered histologically high grade. Conversely, statistically significant differences were seen in the proportion of TMPRSS2–ERG rearrangement between patients with and without mucinous fibroplasia and mucinous features, such that there was a positive association between translocation/deletion and these findings (P=0.001 and 0.018, respectively). A single case with foamy gland histology was observed with TMPRSS2–ERG rearrangement.
Discussion
Over the past four decades, use of the Gleason grading system has become almost universal, in large part because of its ability to predict outcome in prostate cancer.23, 24, 25, 26, 27, 28, 29 Indeed, it is a key component of modern statistical models for biochemical/clinical recurrence and prostate cancer-related mortality.30, 31 In the context of TMPRSS2–ERG rearrangement, therefore, it is somewhat perplexing that many of the reports tying gene fusion to adverse clinical and pathological outcomes simultaneously found no association with Gleason score.4, 6, 8, 12, 20 Moreover, although a small number of studies have described an association of TMPRSS2–ERG gene fusion with higher overall Gleason score, it is difficult to extrapolate those findings to most men diagnosed with prostate cancer in the United States. Specifically, the findings of DeMichelis et al3 and Attard et al2 were derived from unscreened, conservatively managed population-based European cohorts that revealed a relatively low incidence (15% and 30%, respectively) of gene fusion and are not truly comparable to cohorts of PSA-screened and actively treated men, in which a higher incidence of translocation/deletion has been detected.4, 5, 13, 15, 20 Furthermore, the exceedingly small number of rearrangement events (17 of 111 patients) observed by DeMichelis et al3 likely precludes meaningful evaluation of Gleason score between TMPRSS2–ERG rearranged and nonrearranged cases.
Discordant results have also been seen among other studies, which used tissue microarray core-specific grading. In particular, Rajput et al18 report an association between TMPRSS2–ERG rearrangements and moderately to poorly differentiated prostate cancer. However, these authors only assigned one core-specific Gleason pattern for 106 patients, which included 15 cases with Gleason pattern 2, a curious finding because of the difficulties in evaluating architectural circumscription (a key feature of Gleason pattern 2) on tissue microarray cores. Although they conclude that translocation/deletion was more common among Gleason patterns 3–5 than in Gleason pattern 2, the unusual composition of this cohort complicates comparison with other analyses. Conversely, Darnel et al16 found no difference in mean overall Gleason score among 163 patients with and without translocation/deletion, yet a statistically significant association was found between gene fusion and tissue microarray core-specific Gleason pattern 3, as opposed to Gleason pattern 4. However, Gleason pattern 5 was underrepresented in this cohort and only a single core-specific Gleason grade was assigned, such that their findings are difficult to interpret in a clinical context, in which a cancer with primary Gleason pattern 3 may result in a Gleason score of 6, 7, or 8.
This study was undertaken in the background of these discrepant findings, as an attempt to clarify the relationship of TMPRSS2–ERG gene fusion and Gleason score. In a large cohort of PSA-screened US men with surgically treated, clinically localized prostate cancer that displays a broad and representative range of Gleason scores, we have found similar rates of rearrangement to other comparable populations,4, 8, 13, 17, 19, 20 but herein demonstrate that prostate cancers with TMPRSS2–ERG rearrangement are associated with lower core-specific and overall Gleason scores than tumors without this genetic change. As it is well established that TMPRSS2–ERG rearrangement stimulates overexpression of ERG,1 our data compare well with the early work of Petrovics et al32 who observed an inverse relationship between ERG expression and tumor aggressiveness, specifically that higher ERG mRNA levels were significantly associated with low Gleason grade. Likewise, the current core-specific Gleason score data are consonant with our previous report on the relationship between TMPRSS2–ERG gene fusion and clinical outcomes, which demonstrated no association with pathological stage, biochemical recurrence, metastasis, or prostate cancer-related death.5
Attard et al2 were the first to note the potential biological significance of increased copy number of the TMPRSS2–ERG gene fusion and specifically the poor prostate cancer-specific survival of patients with two or more copies of the TMPRSS2–ERG gene fusion. At present, we found an association between cases with copy number increase and higher Gleason scores and a significantly greater likelihood of detecting copy number increase with or without translocation/deletion in Gleason score of 8–10 cancers than in those with Gleason score <7. This relationship with higher Gleason score and similar associations with adverse clinicopathological features/poor outcomes in our previous report5 raised the possibility that either the biological effects of copy number increase in this specific region of chromosome 21 contribute to aggressive disease or that more generalized copy number changes, including those on chromosome 21, are more frequent in prostate cancer with aggressive histological features. The discovery that the majority of copy number increase cases with or without TMPRSS2–ERG rearrangement were nondiploid and showed copy number increase of chromosome 9 (an independent chromosome not typically altered in prostate cancer) by FISH has led to the conclusion that most unfavorable outcomes in prostate cancers with copy number increase are associated with more generalized chromosomal instability.5
Few reports have attempted to correlate TMPRSS2–ERG gene fusion with specific morphological features of prostate cancer. To date, the most comprehensive analyses are those of Mosquera et al7, 12 who found an association between TMPRSS2–ERG rearrangement and a number of histologies, including cribriform glands, signet-ring cells, and intraductal carcinoma. However, the same authors observed no association between rearrangement and Gleason score, a surprising result given that cribriforming (Gleason pattern 4) and signet-ring cells (Gleason pattern 5) are widely considered features of high Gleason grade,23, 24, 33, 34 while intraductal carcinoma is overwhelmingly seen in large volume35 and high Gleason score36 disease. In a similar vein, Tu et al13 report an increased frequency of cribriform glands for TMPRSS2–ERG rearranged vs nonrearranged cases, yet this finding did not achieve statistical significance possibly due to small case numbers.
In this study, we also evaluated these morphological features in a core-specific manner and found no difference in the proportion of translocation/deletion between patients with and without cribriform glands, signet-ring cells, and intraductal carcinoma. In addition, we observed no relationship with glomerulations, a pathognomonic feature of prostate cancer,37 which displays an overwhelming association with Gleason pattern 4 or higher carcinoma.38 In a related finding, Lotan et al11 have recently reported on a series of 38 prostatic ductal adenocarcinomas, a phenotype generally recognized as high grade,33 and demonstrated a low incidence of TMPRSS2–ERG rearrangement when compared with usual acinar carcinoma. These results correspond well with our Gleason score data and support the notion that TMPRSS2 –ERG gene fusion is not linked to morphological elements of aggressive disease.
It is important to note that some of the disparity between studies correlating TMPRSS2–ERG rearrangement with Gleason score may reflect varying sample sizes, differences in clinical and demographic features among cohorts and/or divergence between overall Gleason score, as seen in radical prostatectomy specimens, and evaluation of tissue microarray core-specific Gleason score. In the realm of morphology, the relatively small number of cases of an individual histology (eg, signet-ring cells—seen in only 11 cases in this study and that of Mosquera et al) may lead to dissimilar results. However, it is also curious that intraductal carcinoma, a finding typically observed in high-grade disease, was seen by Mosquera et al12 in 35% (87 of 253) of their cohort, but in only 2% (11 of 521) of cases in our study. This degree of discrepancy suggests substantial variation in defining this morphology.
An interesting pattern that has emerged from multiple studies is the association of TMPRSS2–ERG gene fusions and mucin-related features of prostatic carcinoma. In 2007, both Mosquera et al12 and Tu et al13 reported significant differences in the occurrence of intraluminal blue mucin between TMPRSS2–ERG rearranged and nonrearranged tumors. We initially found this puzzling, as wispy blue acidic mucin is an intraluminal content typically used as a minor diagnostic criterion for prostate cancer.39, 40, 41, 42 Remarkably, these observations have subsequently been extended to include significant associations with mucinous fibroplasia (collagenous micronodules),7 another pathognomonic feature of prostate cancer,37 and mucinous (colloid) carcinoma.10 Most notably, Han et al10 found TMPRSS2–ERG rearrangement in 15 of 18 (83%) mucinous carcinoma specimens drawn from cases that showed definitive mucinous carcinoma on radical prostatectomy. Osunkoya et al43 have highlighted a correlation between the latter two features, with two-thirds of 47 mucinous carcinomas in their series also exhibiting mucinous fibroplasia. This study, which demonstrates translocation/deletion in 77% (17 of 22) of cases with mucinous fibroplasia and 67% (16 of 24) of cases with mucinous features (P=0.001 and 0.018, respectively), confirms these findings. Although all these mucin-related results are derived from relatively small numbers of cases,7, 10, 12, 13 the recurrence of these findings in multiple patient cohorts may signify a link between gene fusion and alteration of molecular pathways favoring a secretory phenotype,12 a phenomenon that should be further explored.
In summary, we have shown that tumors with TMPRSS2–ERG gene fusion are associated with lower core-specific and overall Gleason scores than those without. In a similar manner, we highlight the lack of relationship between translocation/deletion and high-grade morphological features such as cribriform glands, glomerulations, signet-ring cells, and intraductal carcinoma. Conversely, it appears that TMPRSS2–ERG copy number increase, with translocation/deletion, is associated with higher Gleason score. These collective findings suggest that TMPRSS2–ERG rearrangement is not associated with histological features of aggressive prostate cancer.
References
Tomlins SA, Rhodes DR, Perner S, et al. Recurrent fusion of TMPRSS2 and ETS transcription factor genes in prostate cancer. Science 2005;310:644–648.
Attard G, Clark J, Ambroisine L, et al. Duplication of the fusion of TMPRSS2 to ERG sequences identifies fatal human prostate cancer. Oncogene 2007;27:253–263.
Demichelis F, Fall K, Perner S, et al. TMPRSS2:ERG gene fusion associated with lethal prostate cancer in a watchful waiting cohort. Oncogene 2007;26:4596–4599.
Perner S, Mosquera JM, Demichelis F, et al. TMPRSS2-ERG fusion prostate cancer: an early molecular event associated with invasion. Am J Surg Pathol 2007;31:882–888.
Gopalan A, Leversha M, Satagopan J, et al. TMPRSS2-ERG gene fusion is not associated with outcome in patients treated by prostatectomy. Cancer Res 2009;69:1400–1406.
Lapointe J, Kim YH, Miller MA, et al. A variant TMPRSS2 isoform and ERG fusion product in prostate cancer with implications for molecular diagnosis. Mod Pathol 2007;20:467–473.
Mosquera JM, Mehra R, Regan MM, et al. Prevalence of TMPRSS2-ERG fusion prostate cancer among men undergoing prostate biopsy in the United States. Clin Cancer Res 2009;15:4706–4711.
Perner S, Demichelis F, Beroukhim R, et al. TMPRSS2:ERG fusion-associated deletions provide insight into the heterogeneity of prostate cancer. Cancer Res 2006;66:8337–8341.
Saramaki OR, Harjula AE, Martikainen PM, et al. TMPRSS2:ERG fusion identifies a subgroup of prostate cancers with a favorable prognosis. Clin Cancer Res 2008;14:3395–3400.
Han B, Mehra R, Suleman K, et al. Characterization of ETS gene aberrations in select histologic variants of prostate carcinoma. Mod Pathol 2009;22:1176–1185.
Lotan TL, Toubaji A, Albadine R, et al. TMPRSS2-ERG gene fusions are infrequent in prostatic ductal adenocarcinomas. Mod Pathol 2009;22:359–365.
Mosquera JM, Perner S, Demichelis F, et al. Morphological features of TMPRSS2-ERG gene fusion prostate cancer. J Pathol 2007;212:91–101.
Tu JJ, Rohan S, Kao J, et al. Gene fusion between TMPRSS2 and ETS family genes in prostate cancer: frequency and transcript variant analysis by RT-PCR and FISH on paraffin-embedded tissue. Mod Pathol 2007;20:921–928.
Soller MJ, Isaksson M, Elfving P, et al. Confirmation of the high frequency of TMPRSS2/ERG fusion gene in prostate cancer. Genes Chromosomes Cancer 2006;45:717–719.
Albadine R, Latour M, Toubaji A, et al. TMPRSS2-ERG gene fusion in minute (minimal) prostatic adenocarcinoma. Mod Pathol 2009;22:1415–1422.
Darnel AD, LaFargue CJ, Vollmer RT, et al. TMPRSS2-ERG fusion is frequently observed in Gleason pattern 3 prostate cancer in a Canadian cohort. Cancer Bio Ther 2009;8:125–130.
Nam RK, Sugar L, Yang W, et al. Expression of TMPRSS2:ERG fusion gene predicts cancer recurrence after surgery for localised prostate cancer. Br J Cancer 2007;97:1690–1695.
Rajput AB, Miller MA, De Luca A, et al. Frequency of the TMPRSS2-ERG fusion is increased in moderate to poorly differentiated prostate cancers. J Clin Pathol 2007;60:1238–1243.
Yoshimoto M, Joshua AM, Cunha IW, et al. Absence of TMPRSS2:ERG fusions and PTEN losses in prostate cancer is associated with a favorable outcome. Mod Pathol 2008;21:1451–1460.
Mehra R, Han B, Tomlins SA, et al. Heterogeneity of TMPRSS2 gene rearrangements in multifocal prostatic adenocarcinoma: molecular evidence for an independent group of diseases. Cancer Res 2007;67:7991–7995.
Wang J, Cai Y, Ren C, et al. Expression of variant TMPRSS2/ERG fusion messenger RNAs is associated with aggressive prostate cancer. Cancer Res 2006;66:8347–8351.
Leversha MA . Mapping of genomic clones by fluorescence in situ hybridization. Methods Mol Biol 2001;175:109–127.
Gleason DF, Mellinger GT . Prediction of prognosis for prostatic adenocarcinoma by combined histologic grading and clinical staging. J Urol 1974;111:58–64.
Gleason DF . Histologic grading and staging of prostatic carcinoma In: Tannenbaum M (ed). Urologic Pathology: The Prostate. Lea and Febiger: Philadelphia, 1977, pp 171–198.
Epstein JI, Partin AW, Sauvageot J, et al. Prediction of progression following radical prostatectomy: a multi-variate analysis of 721 men with long-term follow-up. Am J Surg Pathol 1996;20:286–292.
Pound CR, Partin AW, Eisenberger MA, et al. Natural history of progression after PSA elevation following radical prostatectomy. JAMA 1999;281:1642–1645.
Bostwick DG, Grignon DJ, Hammond ME, et al. Prognostic factors in prostate cancer: College of American Pathologists consensus statement 1999. Arch Pathol Lab Med 2000;124:995–1000.
Epstein JI, Amin A, Boccon-Gibod L, et al. Prognostic factors and reporting of prostate carcinoma in radical prostatectomy and pelvic lymphadenectomy specimens. Scand J Urol Nephrol Suppl 2005;216:34–63.
Guimaraes MS, Quintal MM, Meirelles LR, et al. Gleason score as predictor of clinicopathologic findings and biochemical (PSA) progression following radical prostatectomy. Int Braz J Urol 2008;34:23–29.
Cooperberg MR, Pasta DJ, Elkin EP, et al. The University of California, San Francisco cancer of the prostate risk assessment score: a straightforward and reliable preoperative predictor of disease recurrence after radical prostatectomy. J Urol 2005;173:1938–1942.
Stephenson AJ, Scardino PT, Eastham JA, et al. Postoperative nomogram predicting the 10-year probability of prostate cancer recurrence after radical prostatectomy. J Clin Oncol 2005;23:7005–7012.
Petrovics G, Liu A, Shaheduzzaman S, et al. Frequent expression of ETS-related gene-1 (ERG1) in prostate cancer transcriptome. Oncogene 2005;24:3847–3852.
Epstein JI, Allsbrook Jr WC, Amin MB, et al. The 2005 International Society of Urologic Pathology (ISUP) consensus conference on Gleason grading of prostatic carcinoma. Am J Surg Pathol 2005;29:1228–1242.
Latour M, Amin MB, Billis A, et al. Grading of invasive cribriform carcinoma on prostate needle biopsy: an interobserver study among experts in genitourinary pathology. Am J Surg Pathol 2008;32:1532–1539.
McNeal JE, Yemoto CEM . Spread of adenocarcinoma within prostatic ducts and acini. Am J Surg Pathol 1996;20:802–814.
Guo CC, Epstein JI . Intraductal carcinoma of the prostate on needle biopsy: histologic features and clinical significance. Mod Pathol 2006;19:1528–1535.
Baisden BL, Kahane H, Epstein JI . Perineural invasion, mucinous fibroplasia, and glomerulations: diagnostic features of limited cancer on prostate needle biopsy. Am J Surg Pathol 1999;23:918–924.
Lotan TL, Epstein JI . Gleason grading of prostatic adenocarcinoma with glomeruloid features on needle biopsy. Hum Pathol 2009;40:471–477.
Iczkowski KA, Bostwick DG . Criteria for biopsy diagnosis of minimal volume prostatic adenocarcinoma: analytic comparison with nondiagnostic but suspicious atypical small acinar proliferation. Arch Pathol Lab Med 2000;124:98–107.
Varma M, Lee MW, Tamboli P, et al. Morphologic criteria for the diagnosis of prostatic adenocarcinoma in needle biopsy specimens: a study of 250 consecutive cases in a routine surgical pathology practice. Arch Pathol Lab Med 2002;126:554–561.
Epstein JI . Diagnosis and reporting of limited adenocarcinoma of the prostate on needle biopsy. Mod Pathol 2004;17:307–315.
Humphrey PA . Diagnosis of adenocarcinoma in prostate needle biopsy tissue. J Clin Pathol 2007;60:35–42.
Osunkoya AO, Nielsen ME, Epstein JI . Prognosis of mucinous adenocarcinoma of the prostate treated by radical prostatectomy: a study of 47 cases. Am J Surg Pathol 2008;32:468–472.
Acknowledgements
This work was supported in part by a Specialized Program of Research Excellence (SPORE) Grant (2P50-CA92629-06) from the National Cancer Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Fine, S., Gopalan, A., Leversha, M. et al. TMPRSS2–ERG gene fusion is associated with low Gleason scores and not with high-grade morphological features. Mod Pathol 23, 1325–1333 (2010). https://doi.org/10.1038/modpathol.2010.120
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2010.120
Keywords
This article is cited by
-
Relationship between Proclarix and the Aggressiveness of Prostate Cancer
Molecular Diagnosis & Therapy (2023)
-
Historical and contemporary perspectives on cribriform morphology in prostate cancer
Nature Reviews Urology (2018)
-
Prostate cancer in Jordanian-Arab population: ERG status and relationship with clinicopathologic characteristics
Virchows Archiv (2017)
-
Micro-RNA-204 Participates in TMPRSS2/ERG Regulation and Androgen Receptor Reprogramming in Prostate Cancer
Hormones and Cancer (2017)
-
The oncogene ERG: a key factor in prostate cancer
Oncogene (2016)