Abstract
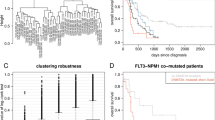
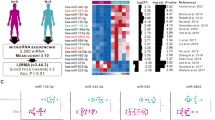
Recently, the p53-miR-34a network has been identified to have an important role in tumorigenesis. As in acute myeloid leukemia with complex karyotype (CK-AML) TP53 alterations are the most common known molecular lesion, we further analyzed the p53-miR-34a axis in a large cohort of CK-AML with known TP53 status (TP53altered, n=57; TP53unaltered, n=31; altered indicates loss and/or mutation of TP53). Profiling microRNA (miRNA) expression delineated TP53 alteration-associated miRNA profiles, and identified miR-34a and miR-100 as the most significantly down- and upregulated miRNA, respectively. Moreover, we found a distinct miR-34a expression-linked gene expression profile enriched for genes belonging to p53-associated pathways, and implicated in cell cycle progression or apoptosis. Clinically, low miR-34a expression and TP53 alterations predicted for chemotherapy resistance and inferior outcome. Notably, in TP53unaltered CK-AML, high miR-34a expression predicted for inferior overall survival (OS), whereas in TP53biallelic altered CK-AML, high miR-34a expression pointed to better OS. Thus, detailed molecular profiling links impaired p53 to decreased miR-34a expression, but also identifies p53-independent miR-34a induction mechanisms as shown in TP53biallelic altered cell lines treated with 15-deoxy-Δ12,14-prostaglandin. An improved understanding of this mechanism might provide novel therapeutic options to restore miR-34a function and thereby induce cell cycle arrest and apoptosis in TP53altered CK-AML.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Dohner H, Estey EH, Amadori S, Appelbaum FR, Buchner T, Burnett AK et al. Diagnosis and management of acute myeloid leukemia in adults: recommendations from an international expert panel, on behalf of the European LeukemiaNet. Blood 2010; 115: 453–474.
Frohling S, Scholl C, Gilliland DG, Levine RL . Genetics of myeloid malignancies: pathogenetic and clinical implications. J Clin Oncol 2005; 23: 6285–6295.
Marcucci G, Mrozek K, Radmacher MD, Garzon R, Bloomfield CD . The prognostic and functional role of microRNAs in acute myeloid leukemia. Blood 2011; 117: 1121–1129.
Bartel DP . MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004; 116: 281–297.
Chen J, Odenike O, Rowley JD . Leukaemogenesis: more than mutant genes. Nat Rev Cancer 2010; 10: 23–36.
Garzon R, Volinia S, Liu CG, Fernandez-Cymering C, Palumbo T, Pichiorri F et al. MicroRNA signatures associated with cytogenetics and prognosis in acute myeloid leukemia. Blood 2008; 111: 3183–3189.
Schwind S, Maharry K, Radmacher MD, Mrozek K, Holland KB, Margeson D et al. Prognostic significance of expression of a single microRNA, miR-181a, in cytogenetically normal acute myeloid leukemia: a Cancer and Leukemia Group B study. J Clin Oncol 2010; 28: 5257–5264.
Hermeking H . The miR-34 family in cancer and apoptosis. Cell Death Differ 2010; 17: 193–199.
Cole KA, Attiyeh EF, Mosse YP, Laquaglia MJ, Diskin SJ, Brodeur GM et al. A functional screen identifies miR-34a as a candidate neuroblastoma tumor suppressor gene. Mol Cancer Res 2008; 6: 735–742.
Zenz T, Mohr J, Eldering E, Kater AP, Buhler A, Kienle D et al. miR-34a as part of the resistance network in chronic lymphocytic leukemia. Blood 2009; 113: 3801–3808.
Swerdlow SH, Campo E, Harris NL, Jaffe ES, Pileri SA, Stein H et al WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues. IARC Press: Lyon, France, 2008.
Rücker FG, Schlenk RF, Bullinger L, Kett H, Becker A, Kugler CM et al. In acute myeloid leukemia with complex karyotype TP53 alterations are associated with specific genomic aberrations and predict inferior survival. Blood (ASH Meeting Abstracts) 2009; 114 abstract 2632.
Bennett JM, Catovsky D, Daniel MT, Flandrin G, Galton DA, Gralnick HR et al. Proposed revised criteria for the classification of acute myeloid leukemia. A report of the French-American-British Cooperative Group. Ann Intern Med 1985; 103: 620–625.
Jaffe ES, Harris NL, Stein H, Vardiman JD eds. World Health Organization Classification of Tumours. Pathology and Genetics of Tumours of Haematopoietic and Lymphoid Tissues. 3rd edn. (IARC Press: Lyon, France, 2001).
Schlenk RF, Dohner K, Mack S, Stoppel M, Kiraly F, Gotze K et al. Prospective evaluation of allogeneic hematopoietic stem-cell transplantation from matched related and matched unrelated donors in younger adults with high-risk acute myeloid leukemia: German-Austrian trial AMLHD98A. J Clin Oncol 2010; 28: 4642–4648.
Schlenk RF, Frohling S, Hartmann F, Fischer JT, Glasmacher A, Del Valle F et al. Intensive consolidation versus oral maintenance therapy in patients 61 years or older with acute myeloid leukemia in first remission: results of second randomization of the AML HD98-B treatment Trial. Leukemia 2006; 20: 748–750.
Breems DA, Van Putten WL, De Greef GE, Van Zelderen-Bhola SL, Gerssen-Schoorl KB, Mellink CH et al. Monosomal karyotype in acute myeloid leukemia: a better indicator of poor prognosis than a complex karyotype. J Clin Oncol 2008; 26: 4791–4797.
Rucker FG, Schlenk RF, Bullinger L, Kayser S, Teleanu V, Kett H et al. TP53 alterations in acute myeloid leukemia with complex karyotype correlate with specific copy number alterations, monosomal karyotype, and dismal outcome. Blood 2012; 119: 2114–2121.
Sander S, Bullinger L, Klapproth K, Fiedler K, Kestler HA, Barth TF et al. MYC stimulates EZH2 expression by repression of its negative regulator miR-26a. Blood 2008; 112: 4202–4212.
Bullinger L, Dohner K, Bair E, Frohling S, Schlenk RF, Tibshirani R et al. Use of gene-expression profiling to identify prognostic subclasses in adult acute myeloid leukemia. N Engl J Med 2004; 350: 1605–1616.
Kharas MG, Lengner CJ, Al-Shahrour F, Bullinger L, Ball B, Zaidi S et al. Musashi-2 regulates normal hematopoiesis and promotes aggressive myeloid leukemia. Nat Med 2010; 16: 903–908.
Luck SC, Russ AC, Du J, Gaidzik V, Schlenk RF, Pollack JR et al. KIT mutations confer a distinct gene expression signature in core binding factor leukaemia. Br J Haematol 2010; 148: 925–937.
Liberzon A, Subramanian A, Pinchback R, Thorvaldsdottir H, Tamayo P, Mesirov JP . Molecular signatures database (MSigDB) 3.0. Bioinformatics 2011; 27: 1739–1740.
Kohlmann A, Bullinger L, Thiede C, Schaich M, Schnittger S, Dohner K et al. Gene expression profiling in AML with normal karyotype can predict mutations for molecular markers and allows novel insights into perturbed biological pathways. Leukemia 2010; 24: 1216–1220.
Rucker FG, Sander S, Dohner K, Dohner H, Pollack JR, Bullinger L . Molecular profiling reveals myeloid leukemia cell lines to be faithful model systems characterized by distinct genomic aberrations. Leukemia 2006; 20: 994–1001.
Rucker FG, Bullinger L, Schwaenen C, Lipka DB, Wessendorf S, Frohling S et al. Disclosure of candidate genes in acute myeloid leukemia with complex karyotypes using microarray-based molecular characterization. J Clin Oncol 2006; 24: 3887–3894.
Lamb J, Crawford ED, Peck D, Modell JW, Blat IC, Wrobel MJ et al. The Connectivity Map: using gene-expression signatures to connect small molecules, genes, and disease. Science 2006; 313: 1929–1935.
Guo H, Ingolia NT, Weissman JS, Bartel DP . Mammalian microRNAs predominantly act to decrease target mRNA levels. Nature 2010; 466: 835–840.
Griffiths-Jones S, Grocock RJ, van Dongen S, Bateman A, Enright AJ . miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res 2006: (34 (Database issue)): D140–D144.
Dweep H, Sticht C, Pandey P, Gretz N . miRWalk—Database: prediction of possible miRNA binding sites by "walking" the genes of three genomes. J Biomed Inform 2011; 44: 839–847.
Maragkakis M, Reczko M, Simossis VA, Alexiou P, Papadopoulos GL, Dalamagas T et al. DIANA-microT web server: elucidating microRNA functions through target prediction. Nucleic Acids Res 2009: (37 (Web Server issue)): W273–W276.
Grimson A, Farh KK, Johnston WK, Garrett-Engele P, Lim LP, Bartel DP . MicroRNA targeting specificity in mammals: determinants beyond seed pairing. Mol Cell 2007; 27: 91–105.
Krek A, Grun D, Poy MN, Wolf R, Rosenberg L, Epstein EJ et al. Combinatorial microRNA target predictions. Nat Genet 2005; 37: 495–500.
Liu C, Kelnar K, Liu B, Chen X, Calhoun-Davis T, Li H et al. The microRNA miR-34a inhibits prostate cancer stem cells and metastasis by directly repressing CD44. Nat Med 2011; 17: 211–215.
Sun F, Fu H, Liu Q, Tie Y, Zhu J, Xing R et al. Downregulation of CCND1 and CDK6 by miR-34a induces cell cycle arrest. FEBS Lett 2008; 582: 1564–1568.
Li WQ, Chen C, Xu MD, Guo J, Li YM, Xia QM et al. The rno-miR-34 family is upregulated and targets ACSL1 in dimethylnitrosamine-induced hepatic fibrosis in rats. Febs J 2011; 278: 1522–1532.
Welch C, Chen Y, Stallings RL . MicroRNA-34a functions as a potential tumor suppressor by inducing apoptosis in neuroblastoma cells. Oncogene 2007; 26: 5017–5022.
Tamura H, Dan K, Tamada K, Nakamura K, Shioi Y, Hyodo H et al. Expression of functional B7-H2 and B7.2 costimulatory molecules and their prognostic implications in de novo acute myeloid leukemia. Clin Cancer Res 2005; 11: 5708–5717.
Zou L, Wu Y, Pei L, Zhong D, Gen M, Zhao T et al. Identification of leukemia-associated antigens in chronic myeloid leukemia by proteomic analysis. Leuk Res 2005; 29: 1387–1391.
Schotte D, De Menezes RX, Moqadam FA, Khankahdani LM, Lange-Turenhout E, Chen C et al. MicroRNA characterize genetic diversity and drug resistance in pediatric acute lymphoblastic leukemia. Haematologica 2011; 96: 703–711.
Doghman M, El Wakil A, Cardinaud B, Thomas E, Wang J, Zhao W et al. Regulation of insulin-like growth factor-mammalian target of rapamycin signaling by microRNA in childhood adrenocortical tumors. Cancer Res 2010; 70: 4666–4675.
Zheng YS, Zhang H, Zhang XJ, Feng DD, Luo XQ, Zeng CW et al. MiR-100 regulates cell differentiation and survival by targeting RBSP3, a phosphatase-like tumor suppressor in acute myeloid leukemia. Oncogene 2011; 31: 80–92.
Christoffersen NR, Shalgi R, Frankel LB, Leucci E, Lees M, Klausen M et al. p53-independent upregulation of miR-34a during oncogene-induced senescence represses MYC. Cell Death Differ 2010; 17: 236–245.
Pulikkan JA, Peramangalam PS, Dengler V, Ho PA, Preudhomme C, Meshinchi S et al. C/EBPalpha regulated microRNA-34a targets E2F3 during granulopoiesis and is down-regulated in AML with CEBPA mutations. Blood 2010; 116: 5638–5649.
Marcucci G, Maharry K, Radmacher MD, Mrozek K, Vukosavljevic T, Paschka P et al. Prognostic significance of, and gene and microRNA expression signatures associated with, CEBPA mutations in cytogenetically normal acute myeloid leukemia with high-risk molecular features: a Cancer and Leukemia Group B Study. J Clin Oncol 2008; 26: 5078–5087.
Zhao M, Discipio RG, Wimmer AG, Schraufstatter IU . Regulation of CXCR4-mediated nuclear translocation of extracellular signal-related kinases 1 and 2. Mol Pharmacol 2006; 69: 66–75.
Chim CS, Wong KY, Qi Y, Loong F, Lam WL, Wong LG et al. Epigenetic inactivation of the miR-34a in hematological malignancies. Carcinogenesis 2010; 31: 745–750.
Kaller M, Liffers ST, Oeljeklaus S, Kuhlmann K, Roh S, Hoffmann R et al. Genome-wide characterization of miR-34a induced changes in protein and mRNA expression by a combined pulsed SILAC and micro-array analysis. Mol Cell Proteomics 2011; 10: M111.010462.
Lewis JD, Payton LA, Whitford JG, Byrne JA, Smith DI, Yang L et al. Induction of tumorigenesis and metastasis by the murine orthologue of tumor protein D52. Mol Cancer Res 2007; 5: 133–144.
Acknowledgements
Analyses were performed using BRB-ArrayTools developed by Dr Richard Simon and BRB-ArrayTools Development Team. We thank all members of the German-Austrian AML Study Group (AMLSG) for their participation in this study and for providing patient samples. This study was supported in part by the Bundesministerium für Bildung und Forschung (NGFNplus Grant No. 01GS0871), by the Deutsche José Carreras Stiftung e.V. (DJCLS R 06/41v), and LB was supported in part by the Deutsche Forschungsgemeinschaft (Heisenberg-Stipendium BU 1339/3-1).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Leukemia website
Supplementary information
Rights and permissions
About this article
Cite this article
Rücker, F., Russ, A., Cocciardi, S. et al. Altered miRNA and gene expression in acute myeloid leukemia with complex karyotype identify networks of prognostic relevance. Leukemia 27, 353–361 (2013). https://doi.org/10.1038/leu.2012.208
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2012.208