Abstract
In Epstein–Barr virus (EBV) negative Hodgkin's cell lines and classical EBV-negative Hodgkin's lymphoma (HL), Reed–Sternberg cells (RS cells) represent end-stage tumor cells, in which further nuclear division becomes impossible because of sustained telomere loss, shortening and aggregation. However, the three-dimensional (3D) telomere organization in latent membrane protein 1 (LMP1)-expressing RS cells of EBV-associated HL is not known. We performed a 3D telomere analysis after quantitative fluorescent in situ hybridization on 5 μm tissue sections on two LMP1-expressing HL cases and showed highly significant telomere shortening (P<0.0001) and formation of telomere aggregates in RS cells (P<0.0001), when compared with the mononuclear precursor Hodgkin cells (H cells). Telomere-poor or telomere-free ‘ghost’ nuclei were a regular finding in these RS cells. These nuclei and their telomere content strongly contrasted with the corona of surrounding lymphocytes showing numerous midsized telomere hybridization signals. Both H cells and RS cells of two EBV-negative HL cases analyzed in parallel showed 3D telomere patterns identical to those of LMP1-expressing cases. As a major advance, our 3D nuclear imaging approach allows the visualization of hitherto unknown profound changes in the 3D nuclear telomere organization associated with the transition from LMP1-positive H cells to LMP1-positive RS cells. We conclude that RS cells irrespective of LMP1 expression are end-stage tumor cells in which the extent of their inability to divide further is proportional to the increase of very short telomeres, telomere loss, aggregate formation and the generation of ‘ghost’ nuclei.
Similar content being viewed by others
Main
Binuclear or multinuclear Reed–Sternberg cells (RS cells) are the key elements for the diagnosis of Hodgkin's lymphoma (HL).1, 2 In about 40–50% of HL cases, Hodgkin cells (H cells) and RS cells express the Epstein–Barr virus (EBV)-encoded latent membrane protein 1 (LMP1) oncoprotein or its deletion variants,3, 4 and the risk of developing LMP1-expressing HL within a median incubation time of 4 years after symptomatic EBV infection is significantly (fourfold) increased.5
Mononuclear H cells are the precursors of multinuclear RS cells,6, 7, 8, 9 and endomitotic multinucleation is associated with disturbed cytokinesis and jumping translocations10, 11 pointing to a severe telomere dysfunction.12, 13 Telomeres are the nucleoprotein complexes at the ends of chromosomes in which a number of specific proteins, either binding telomere proteins directly or a protein complex, termed ‘shelterin,’ is directly associated with telomeric DNA.14, 15, 16
Profound changes in the three-dimensional (3D) nuclear organization of telomeres are the hallmark of the transition from mononuclear H cells to multinuclear RS cells in EBV-negative Hodgkin cell lines and in classical EBV-negative HL. We recently showed that EBV-negative RS cells represent end-stage tumor cells, in which further nuclear division becomes impossible because of sustained telomere loss, shortening and aggregation.1 However, nothing is known about the 3D telomere organization in LMP1-expressing H and RS cells of EBV-associated Hodgkin's disease (HD). In this study, we document 3D telomere dynamics in LMP1-expressing H and RS cells and show conformity with those observed in EBV-negative HD analyzed in parallel.
MATERIALS AND METHODS
Ethics
The study design has been approved by the Institutional Ethics Committee of the Université de Sherbrooke and the Institutional Review Board of the University of Manitoba.
Informed consent was obtained from each patient, and the investigations were conducted according to the Declaration of Helsinki.
Tissue Slides
Archival formalin-fixed, paraffin-embedded tissue slides (serial sections of 5 μm) from three patients of EBV-associated and from three patients of EBV-negative classical HD were deparaffinized twice for 15 min at room temperature in xylene and placed in 100% ethanol. Slides were subsequently rehydrated in a descending gradient of ethanol–water to 30% ethanol, transferred to PBS and used for hematoxylin–eosin staining (serial section no. 1), immunostaining (CD30, serial section no. 2; LMP1, serial section no. 3) and quantitative fluorescent in situ hybridization (Q-FISH, serial section no. 4).
Identification of LMP1-expressing H and RS cells on serial sections with combined 3D DAPI/Cy-3 telomere Q-FISH nuclear staining were as follows: On serial section no. 3, lymph node regions with LMP1-expressing H and RS cells were identified at a × 200 magnification and individual cells were further confirmed at a × 630 magnification. Subsequently, on serial section no. 4, the corresponding region was identified at a × 200 magnification, and the corresponding H and RS cells (hence LMP1 expressing) were further analyzed for 3D nuclear telomere organization at a × 630 magnification.
Immunohistochemistry
Immunostaining was performed by standard indirect immunoperoxidase technique using primary monoclonal mouse antibodies anti-CD30 (Ber-H2) and anti-LMP1 (clones CS1-CS4) from Dako (Glostrup, Denmark), at a dilution of 1:40 and 1:50, respectively. Photomicrographs were performed using a Zeiss Axioskop 2 microscope with a Polaroid C11806TV camera and Polaroid DMC2 v2.01.software.
Telomere Q-FISH
The telomere FISH protocol was performed17, 18, 19 using Cy3-labeled peptide nucleic acid probes (DAKO). Imaging of interphases after telomere FISH was performed using Zeiss AxioImager Z1 with a cooled AxioCam HR B&W, DAPI, Cy3 filters in combination with a Planapo 63 × /1.4 oil objective lens. Images were acquired using AXIOVISION4.6 (Zeiss) in a multichannel mode, followed by constraint iterative deconvolution as specified below.
3D Image Acquisition
At least 30 H-cell interphase nuclei and at least 30 RS-cell interphase polycaria were analyzed in each of the two cases of EBV-associated and EBV-negative HD. One case each of EBV-associated and EBV-negative HL with <30 RS cells were not included in the statistical analysis. However, these RS cells showed the same telomere organization as did the statistically analyzed cases. AXIOVISION 4.6 with deconvolution module and rendering module were used. For every fluorochrome, the 3D image consists of a stack of 40 images with a sampling distance of 200 nm along the z direction and 107 nm in the xy direction. The constrained iterative algorithm option was used for deconvolution.20
3D Image Analysis for Telomeres
Telomere measurements were performed using TeloView18, 21 By choosing a simple threshold for the telomeres, a binary image was found. On the basis of that, the center of gravity of intensities was calculated for every object resulting in a set of coordinates (x, y, z) denoted by crosses on the screen. The integrated intensity of each telomere was calculated because it is proportional to the telomere length.22
Segmental Nuclear Volume
Nuclear volume within one 5 μm thin nuclear section of H or RS cell is calculated according to the 3D nuclear DAPI staining as described previously.23 Contrary to whole-cell preparations (cells or cell lines), in which the nuclei can be visualized with their entire volumes and z-stack analysis along the z direction over 15 μm allows the calculation of the entire nuclear volume, in tissue sections the nuclear volume analysis is limited to 5 μm nuclear segments (used as a standard for histopathological diagnosis) along the z direction.
Deparaffinized tissue slides of 10 and 15 μm thickness are technically unsatisfactory for Q-FISH analysis. Thus, the segmental nuclear volume represents about 30–50% of the total nuclear volume of H cells (nuclear diameter of about 10–15 μm) and about 15–25% of the total nuclear volume of RS cells (diameter of two up to several nuclei about 20–40 μm).
Segmental Telomere Number
Segmental telomere number is the sum of all short, midsized and large telomeres and aggregates identified within one 5 μm thin nuclear section of an H or RS cell.
Segmental Telomere Intensity
Segmental telomere intensity is the sum of intensities of all short, midsized and large telomeres and aggregates identified within one 5 μm thin nuclear section of an H or RS cell (viz ∑2 × 15 000 Units >∑7 × 4000 Units).
Mean Telomere Intensity
Indicates mean telomere relative fluorescent intensity (length) of all telomeres within a given segmental volume.
Telomere Length
Telomeres with a relative fluorescent intensity (y axis) ranging from 0 to 5000 Units are classified as very short, with an intensity ranging from 5000 to 15 000 Units as short, with an intensity from 15 000 to 30 000 Units as midsized and with an intensity >30 000 Units as large.
Telomere Aggregates
Telomere aggregates are defined as clusters of telomeres that are found in close association and cannot be further resolved as separate entities at an optical resolution limit of 200 nm.18
Statistical Analysis
For each case, normally distributed parameters are compared between the two types of cells using nested ANOVA or two-way ANOVA. Multiple comparisons using the least square means tests followed, in which interaction effects between two factors were found to be significant. Other parameters that were not normally distributed were compared using a nonparametric Wilcoxon rank sum test. Significance level was set at P=0.05. Analyses were carried out using SAS v9.1 programs.
RESULTS
In EBV-associated HL, mononuclear H cells and multinuclear (at least binuclear) RS cells were identified by LMP1 and CD30 expression, whereas in EBV-negative HL, CD30 had to be present (Figures 1a and b).
Mononuclear Hodgkin and multinuclear Reed–Sternberg cells in classical Hodgkin's lymphoma. (a) Female, 29 years, classical HD, nodular sclerosis subtype, EBV-positive, supra-clavicular lymph node. Two LMP1-expressing Hodgkin cells (brown staining, bottom) and two LMP1-expressing Reed–Sternberg cells (brown staining, middle and top) are shown. (b) Male, 34 years, classical HD, nodular sclerosis subtype, EBV-negative, cervical lymph node. One multinucleated Reed–Sternberg cell (top) and several mononuclear Hodgkin cells, some of them with twisted nuclei (middle), show strong CD30 staining (brown).
LMP1-Expressing HL
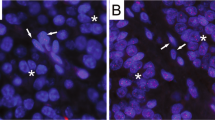
RS cells of both cases were characterized by a significant shift from midsized and short telomeres to very short telomeres (P<0.0001) when compared with mononuclear H cells as quantified in Figure 2. This is further illustrated in representative images of Figures 3a–c; the large RS cell in the center of the image showed few, unevenly distributed small-sized telomeric signals, whereas the surrounding lymphocytes (labeled as 1, 2 and 3) displayed bright telomeric signals. Loss of midsized telomeres was also prominent in large twisted H cells (see arrows in Figure 4), and in many RS cells, telomere-poor or telomere-free ‘ghost’ nuclei (Figure 5a and b, nucleus no. 2) were adjacent to telomere-rich micronuclei with aggregates (Figure 5a and b, nucleus no. 3). Overall telomere characteristics of LMP1-expressing H and RS cells of an EBV-associated HL are shown in Table 1a. As expected, the segmental nuclear volume of RS cells is about twice that of H cells and the segmental telomere number is also higher. However, the segmental telomere intensity, ie, the total DNA mass of telomeres is nearly the same, indicating that no additional and significant telomere elongation takes place. On the contrary, telomeres have to get much shorter during the transition from H to RS cells, as demonstrated by a decrease in the mean telomere intensity, especially through a highly significant increase of very short telomeres. Telomere shortening is accompanied by a doubling of number of telomere aggregates.
Telomere distribution according to size in mononuclear Hodgkin (blue) and at least binuclear Reed–Sternberg cells (red) in EBV-positive Hodgkin's disease. Results are based on 3D analysis of at least 30 Hodgkin and 30 Reed–Sternberg cells. Same patient as shown in Figure 1a. Frequency (x axis) and relative fluorescent intensity, ie, size of telomeres (y axis) in a diagnostic 5 μm thin supra-clavicular lymph node section. There is a highly significant shift (P<0.0001) from midsized and short telomeres to very short telomeres of 0–5000 relative fluorescence units.
3D nuclear organization of telomeres (red) and total nuclear DNA (blue) in EBV-positive, LMP1-expressing Sternberg–Reed cells of classical Hodgkin's lymphoma. Same patient as shown in Figure 1a. (a) Multinuclear outré Reed–Sternberg cell showing unequal nuclear distribution of mainly short telomeres when compared with surrounding lymphocytes (1, 2, 3), which contain midsized telomeres. (b) DAPI staining (gray scale image, for better contrast) of the same Reed–Sternberg cell and surrounding lymphocytes is shown. On representative 2D z-stack image no. 20 out of 40 multinuclearity and endomitosis with incomplete nuclear separation is evident. (c) Mononuclear Hodgkin cell (to scale) shows mainly midsized telomeres.
3D identification of telomere shortening and loss in a large, twisted LMP1-expressing Hodgkin cell. Female, 77 years, axillary lymph node, classical Hodgkin's lymphoma, nodular sclerosis subtype, EBV positive. (a) In a large Hodgkin cell, some very short (arrows) telomeres (red) are identified, whereas the surrounding lymphocytes contain numerous midsized telomeres. (b) DAPI staining (blue) of the same Hodgkin cell reveals twisted nuclear structure in contrast to surrounding lymphocytes with homogenous nuclear structure. (c) Combined 3D nuclear staining confirms telomere loss and shortening (arrows) in a large twisted Hodgkin cell, whereas surrounding lymphocytes show normal telomere distribution.
3D identification of a completely disturbed nuclear telomere organization in a trinuclear LMP1-expressing Reed–Sternberg cell. Same patient as shown in Figure 4. (a) DAPI staining (gray scale image, for better contrast) identifies a trinuclear (1, 2, 3) Reed–Sternberg cell and surrounding lymphocytes. (b) Combined 3D nuclear staining demonstrates few remnant telomeres (red) in nucleus 1 (blue), identifies nucleus 2 as telomere-free ‘ghost’ nucleus and nucleus 3 as micronucleus with several midsized telomeres or aggregates.
EBV-Negative HL
Both cases showed a significant shift from midsized and short telomeres to very short telomeres (P<0.0001) when compared with mononuclear H cells (Figure 6), and lacunar ‘ghost’ RS cells were frequently observed (Figure 7). Overall telomere characteristics of H and RS cells of an EBV-negative HL are shown in Table 1b (identical experimental conditions, as the LMP1-expressing case of Table 1a). Telomere dynamics show the same pattern as in the LMP1-expressing cases. In particular, characteristics earlier identified for EBV-negative Hodgkin cell lines and EBV-negative HL are still highly significant in the setting of LMP1-expressing HL and inclusion of additional EBV-negative HL (Table 2).
Telomere distribution according to size in mononuclear Hodgkin (blue) and at least binuclear Reed–Sternberg cells (red) in EBV-negative Hodgkin's disease. Results are based on 3D analysis of at least 30 Hodgkin and 30 Reed–Sternberg cells. Frequency (x axis) and relative fluorescent intensity, ie, size of telomeres (y axis) in a diagnostic 5 μm thin cervical lymph node section is shown. There is a highly significant shift (P<0.0001) from midsized and short telomeres to very short telomeres of 0–5000 relative fluorescence units.
Same patient as shown in Figure 1b. 3D combined telomere FISH (red) and nuclear DAPI staining (blue) of a diagnostic 5 μm thin cervical lymph node section. A huge telomere-free end-stage multinuclear ‘ghost’ Reed–Sternberg cell is surrounded by multiple reactive lymphocytes containing midsized telomeres. Arrows point to the nuclei. Inset (upper right) confirms telomere-free ‘ghost’ nuclei.
In summary, LMP1-expressing H and RS cells display nearly congruent 3D telomere configurations when compared with their EBV-negative counterparts. RS cells, irrespective of their LMP1 expression, are characterized by a highly significant loss of their telomere mass (quantity of telomere ADN repeats in relation to nuclear volume), a highly significant increase of telomere aggregates and a highly significant increase of very short telomeres, indicating a real telomere erosion during the passage from H to RS cells.
DISCUSSION
EBV-associated HL was first described by Poppema et al24 in 1984 and was confirmed at a large scale a few years later by EBER in situ hybridization and LMP1 immunohistochemistry.25, 26 The presentation, clinical course and response to chemotherapy of EBV-associated HL are very similar to EBV-negative HL,27 but the LMP1-expressing nodular sclerosis type may have a less favorable long-term prognosis.28, 29 Relevant differences in EBV association are observed according to socioeconomic status,30 and the risk to develop EBV-associated HL after symptomatic infectious mononucleosis is significantly increased.5
In EBV-associated HL, H and RS cells display a latency type II pattern with expression of EBER, EBNA1, LMP2A, B and LMP1, a multifunctional oncoprotein acting through the activation of the NF-κ-B, JNK and the JAK3-STAT pathway.31, 32 Transient expression of the LMP1 oncoprotein in the EBV-negative Hodgkin's cell lines L-428 and HD-MyZ results in a significant increase of LMP1-expressing RS cells, thus identifying LMP1 as a potent inducer of multinuclearity.31, 33 LMP1 also activates the hTERT promoter in germinal center-derived BJAB Burkitt cells enhancing telomerase activity at the transcriptional level.34 Moreover, EBNA1, also expressed in EBV-associated HL, promotes genomic instability in EBV-negative Burkitt cell lines BJAB and DG75 cells by the induction of reactive oxygen species, heralded by compensatory upregulation of γH2AX.35 Thus, experimental LMP1 and EBNA1 upregulation in a germinal center-derived B-cell setting—Burkitt and RS cells are both germinal center derived 2—mimics three in vivo and in vitro characteristics of EBV-negative RS cells, namely multinuclearity, high telomerase activity and genomic instability.1, 2, 36, 37, 38
Our results of 3D nuclear telomere dynamics associated with the transition of LMP1-expressing H to LMP1-expressing RS cells show complete congruity with our recently published findings in Hodgkin's cell lines and classical EBV-negative HL.1 In this study, three LMP1-expressing HL cases and three additional EBV-negative HL cases, analyzed under identical experimental conditions, show largely identical telomere characteristics. To strengthen this point, we even identify the characteristic 3D telomeric features when data from both HL groups are pooled (Table 2). LMP1-expressing RS cells are—similar to LMP1-negative RS cells—characterized by a high number of very short telomeres, a loss of total telomere mass and a significant increase of aggregates when compared with mononuclear H cells. These very short telomeres termed ‘t-stumps’ were recently identified and represent a common hallmark of tumor cells.39 These ‘t-stumps’ remain short even when telomerase activity is high.39 There is growing evidence that the integrity of nuclear chromosome territories is essential for successful mitosis.40, 41 With that in mind, it will be hard to imagine that trinuclear LMP1-positive RS cells (as shown in Figure 5) composed of a first nucleus with a few short telomeres, a second telomere-free ‘ghost’ nucleus and a third micronucleus with several midsized telomeres or aggregates, will be able to get through the next round of successful karyokinesis and thus may represent true end-stage tumor cells. Thus, our findings in LMP1-expressing HL are also in accordance with the recent identification of circulating clonotypic B cells in classical HL,42 which give rise to H cells that may progress to real end-stage RS cells. As a major advance, our 3D nuclear imaging approach allowed the visualization of hitherto unknown profound changes in the 3D nuclear telomere organization associated with the transition from LMP1-positive H to LMP1-positive RS cells.
References
Knecht H, Sawan B, Lichtensztejn D, et al. The 3D nuclear organization of telomeres marks the transition from Hodgkin to Reed-Sternberg cells. Leukemia 2009;23:565–573.
Küppers R . The biology of Hodgkin's lymphoma. Nat Rev Cancer 2009;9:15–27.
Glaser SL, Lin RJ, Stewart SL, et al. Epstein-Barr virus-associated Hodgkin's disease: epidemiologic characteristics in international data. Int J Cancer 1997;70:375–382.
Knecht H, Bachmann E, Brousset P, et al. Deletions within the LMP1 oncogene of Epstein-Barr virus are clustered in Hodgkin's disease and identical to those observed in nasopharyngeal carcinoma. Blood 1993;82:2937–2942.
Hjalgrim H, Askling J, Rostgaard K, et al. Characteristics of Hodgkin's lymphoma after infectious mononucleosis. N Engl J Med 2003;349:1324–1332.
Newcom SR, Kadin ME, Phillips C . L-428 Reed-Sternberg cells and mononuclear Hodgkin's cells arise from a single cloned mononuclear cell. Int J Cell Cloning 1993;6:417–431.
Hsu SM, Zhao X, Chakraborty S, et al. Reed-Sternberg cells in Hodgkin's cell lines HDLM, L-428, and KM-H2 are not actively replicating: lack of bromodeoxyuridine uptake by multinuclear cells in culture. Blood 1988;71:1382–1389.
Drexler HG, Gignac SM, Hoffbrand AV, et al. Formation of multinucleated cells in a Hodgkin's-disease-derived cell line. Int J Cancer 1989;43:1083–1090.
Wolf J, Kapp U, Bohlen H, et al. Peripheral blood mononuclear cells of a patient with advanced Hodgkin's lymphoma give rise to permanently growing Hodgkin-Reed Sternberg cells. Blood 1996;87:3418–3428.
Pihan GA, Purohit A, Wallace J, et al. Centrosome defects and genetic instability in malignant tumors. Cancer Res 1998;58:3974–3985.
MacLeod RA, Spitzer D, Bar-Am I, et al. Karyotypic dissection of Hodgkin's disease cell lines reveals ectopic subtelomeres and ribosomal DNA at sites of multiple jumping translocations and genomic amplification. Leukemia 2000;14:1803–1814.
Bailey SM, Murnane JP . Telomeres, chromosome instability and cancer. Nucleic Acid Res 2006;34:2408–2417.
Mai S, Garini Y . The significance of telomeric aggregates in the interphase nuclei of tumor cells. J Cell Biochem 2006;97:904–915.
LeBel C, Wellinger RJ . Telomeres: what's new at your end? J Cell Sci 2005;118:2787–2788.
De Lange T . Shelterin: the protein complex that shapes and safeguards human telomeres. Gene Dev 2005;19:2100–2110.
Hug N, Lingner J . Telomere length homeostasis. Chromosoma 2006;115:413–425.
Figueroa R, Lindenmaier H, Hergenhan M, et al. Telomere erosion varies during in vitro aging of normal human fibroblasts from young and adult donors. Cancer Res 2000;60:2770–2774.
Chuang TC, Moshir S, Garini Y, et al. The three-dimensional organization of telomeres in the nucleus of mammalian cells. BMC Biol 2004;2:12.
Louis SF, Vermolen BJ, Garini Y, et al. C-Myc induces chromosomal rearrangements through telomere and chromosome remodeling in the interphase nucleus. Proc Natl Acad Sci USA 2005;102:9613–9618.
Schaefer LH, Schuster D, Herz H . Generalized approach for accelerated maximum likelihood based image restoration applied to three-dimensional fluorescence microscopy. J Microsc 2001;204:99–107.
Vermolen BJ, Garini Y, Mai S, et al. Characterizing the three-dimensional organization of telomeres. Cytometry A 2005;67:144–150.
Poon SS, Martens UM, Ward RK, et al. Telomere length measurements using digital fluorescence microscopy. Cytometry 1999;36:267–278.
Sarkar R, Guffei A, Vermolen BJ, et al. Alterations of centromere positions in nuclei of immortalized and malignant mouse lymphocytes. Cytometry A 2007;71:386–392.
Poppema S, van Imhoff G, Torensma R, et al. Lymphadenopathy morphologically consistent with Hodgkin's disease associated with Epstein-Barr virus infection. Am J Clin Pathol 1984;84:385–390.
Weiss LM, Movahed LA, Warnke RA, et al. Detection of Epstein-Barr viral genomes in Reed-Sternberg cells of Hodgkin's disease. N Engl J Med 1989;320:502–506.
Pallesen G, Hamilton-Dutoit SJ, Rowe M, et al. Expression of Epstein-Barr virus latent gene products in tumor cells of Hodgkin's disease. Lancet 1991;337:320–322.
Murray PG, Billingham LJ, Hassan HT, et al. Effect of Epstein-Barr virus infection on responses to chemotherapy and survival in Hodgkin's disease. Blood 1999;94:442–447.
Claviez A, Tiemann M, Lüders H, et al. Impact of latent Epstein-Barr virus infection on outcome in children and adolescents with Hodgkin's lymphoma. J Clin Oncol 2005;23:4048–4056.
Keegan TH, Glaser SL, Clarke CA, et al. Epstein-Barr virus as a marker of survival after Hodgkin's lymphoma: a population based study. J Clin Oncol 2005;23:7604–7613.
Dinand V, Arya LS . Epidemiology of childhood Hodgkin's disease: is it different in developing countries? Indian Pediatr 2006;43:141–147.
Knecht H, Berger C, Rothenberger S, et al. The role of Epstein-Barr virus in neoplastic transformation. Oncology 2001;60:289–302.
Dolcetti R, Masucci MG . Epstein-Barr virus: induction and control of cell transformation. J Cell Physiol 2001;196:207–218.
Knecht H, McQuain C, Rothenberger S, et al. Expression of the LMP1 oncoprotein in the EBV negative Hodgkin's disease cell line L-428 is associated with Reed-Sternberg cell morphology. Oncogene 1996;13:947–953.
Terrin L, Dal Col J, Rampazzo E, et al. Latent membrane protein 1 of Epstein-Barr virus activates the hTERT promoter and enhances telomerase activity in B lymphocytes. J Virol 2008;82:10175–10187.
Gruhne B, Sompallae R, Marescotti D, et al. The Epstein-Barr virus nuclear antigen-1 promotes genomic instability via induction of reactive oxygen species. Proc Natl Acad Sci USA 2009;106:2313–2318.
Norrback KF, Enblad G, Erlanson M, et al. Telomerase activity in Hodgkin's disease. Blood 1998;92:567–573.
Heine B, Hummel M, Demel G, et al. Hodgkin and Reed-Sternberg cells of classical Hodgkin's disease overexpress the telomerase RNA template (hTR). J Pathol 1999;188:139–145.
Martin-Subero JI, Knippschild U, Harder L, et al. Segmental chromosomal aberrations and centrosome amplifications: pathogenetic mechanisms in Hodgkin and Reed-Sternberg cells of classical Hodgkin's lymphoma. Leukemia 2003;17:2214–2219.
Xu L, Blackburn EH . Human cancer cells harbour T-stumps, a distinct class of extremely short telomeres. Mol Cell 2007;28:315–327.
Cremer T, Cremer M, Dietzel S, et al. Chromosome territories—a functional nuclear landscape. Curr Opin Cell Biol 2006;18:307–316.
De Vos WH, Hoebe RA, Joss GH, et al. Controlled light exposure microscopy reveals dynamic telomere microterritories throughout the cell cycle. Cytometry A 2009;75:428–439.
Jones RJ, Gocke CD, Kasamon JL, et al. Circulating clonotypic B cells in classical Hodgkin lymphoma. Blood 2009;113:5920–5926.
Acknowledgements
We thank Mary Cheang, PhD, University of Manitoba Biostatistics Unit, for statistical analysis of data. We are grateful to receive support from the Canadian Institutes of Health Research (SM), and the Centre de Recherche Clinique du CHUS (Grant no. PAFI 90914 to HK).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Knecht, H., Sawan, B., Lichtensztejn, Z. et al. 3D Telomere FISH defines LMP1-expressing Reed–Sternberg cells as end-stage cells with telomere-poor ‘ghost’ nuclei and very short telomeres. Lab Invest 90, 611–619 (2010). https://doi.org/10.1038/labinvest.2010.2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.2010.2
Keywords
This article is cited by
-
Three-dimensional nuclear telomere architecture and differential expression of aurora kinase genes in chronic myeloid leukemia to measure cell transformation
BMC Cancer (2022)
-
Disruption of direct 3D telomere–TRF2 interaction through two molecularly disparate mechanisms is a hallmark of primary Hodgkin and Reed–Sternberg cells
Laboratory Investigation (2017)
-
Evaluation of telomere length in human cardiac tissues using cardiac quantitative FISH
Nature Protocols (2017)
-
Insights into the molecular roles of heparan sulfate proteoglycans (HSPGs—syndecans) in autocrine and paracrine growth factor signaling in the pathogenesis of Hodgkin’s lymphoma
Tumor Biology (2016)
-
3D nuclear organization and genomic instability in cancer
BMC Proceedings (2013)