Abstract
Alterations in LINE-1 methylation are related to many diseases. The levels and patterns of LINE-1 hypomethylation were associated with a higher risk in developing several cancers, having a poorer prognosis and more aggressiveness. To evaluate the LINE-methylated status in psoriasis, LINE-1 methylation in various cells from patients with psoriasis, squamous cell carcinoma and normal controls were assessed by combined bisulfite restriction analysis of LINE-1. The results of the epigenetic changes for intragenic LINE-1 gene expression were also tested on two known expression microarrays. In patients with psoriasis, hypomethylation of LINE-1 and increase in %uCuC were prominent in the keratinocytes when compared with normal controls (P=0.014 and P=0.020, respectively). Alternatively, %uCmC was significantly lower in patients with severe psoriasis compared with mild psoriasis (P=0.022). The receiver-operating characteristic curve analysis indicated the high specificity and sensitivity of uCuC and uCmC in detecting psoriasis and severity of psoriasis. From expression array analysis, genes with LINE-1 were downregulated more than those genes without LINE-1 (P=3.84 × 10−27 and P=2.14 × 10−21, respectively). Modification in LINE-1 methylation may alter the gene expression resulting in a phenotypic change of the psoriatic skin. %uCuC and %uCmC may be used as biomarkers for psoriasis.
Similar content being viewed by others
Introduction
Non-long terminal repeats retrotransposons are transposable elements capable of insertion mutations that alter a gene’s expression, potentially rendering the gene unstable.1, 2, 3 Non-long terminal repeat retrotransposons constitute about 34% of the human genome.1 Non-long terminal repeat retrotransposons with the highest densities are long interspersed element-1 (LINE-1 or L1) (17%) and Alu (11%). The full-length LINE-1 and Alu are approximately 6 kilobases (kb) and 300 base pairs (bp) in length, respectively. LINE-1 contains an internal RNA polymerase II promoter and two open reading frames (ORF1 and ORF2) in 5′ untranslated region and a polyadenylation signal ending with oligo dA-rich tail of invariable length in 3′ untranslated region. Alu contains internal RNA polymerase III promoter and the element ends with oligo dA-rich tail of variable length. Owing to the retrotransposition events that have occurred during evolution, more than 500 000 copies of LINE-1 and 1 000 000 copies of Alu are widely inserted into the genome and as a result of this, undoubtedly produce numerous functional outcomes.2, 4, 5 Many LINE-1s control gene regulation by their 5′ untranslated region promoter either by the production of unique RNA sequences or via the intragenic LINE-1 RNAs that can repress host gene expression.6 Several studies have shown that DNA methylation is an important mechanism in silencing LINE-1 to control gene expression and to maintain genomic stability.7 Different methylation levels and patterns of LINE-1 at each loci were observed in various normal tissue types.8 Several studies have suggested that hypomethylation of LINE-1 and Alu are the causes for global hypomethylation and genomic instability in many malignancies and autoimmune diseases.5, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18 On the other hand, some reports showed that LINE-1 methylation levels were positively correlated with diseases in many loci at genome-wide levels as seen in cancer; hypermethylation of gene promoters are frequently discovered in patients with cancer.8, 19, 20 In addition, the degree of LINE-1 hypomethylation was directly associated with a higher risk for developing a more aggressive cancer with poorer prognosis.9, 12, 13, 21 Recently, a pattern of LINE-1 methylation was reported to be used as a biomarker for various types of cancers. uCuC pattern was found to be more efficient in distinguishing the cancers of the colon, liver, lung and nasopharynx from the normal controls.22 LINE-1 hypomethylation was also found in lymphocytes and synovial fibroblasts of autoimmune patients with systemic lupus erythematosus and rheumatoid arthritis, respectively.15, 16
Despite this, there are no data on LINE-1 and Alu in psoriasis, which is the most common chronic skin disease found worldwide. The disease is characterized by an abnormal proliferation and differentiation of keratinocytes and inflammation of the epidermis and dermis layers of the skin. Genetic and environmental factors are important in the phenotypic expression of the disease. Recently, epigenetic changes have been implicated in the pathogenesis of psoriasis. Several studies have reported that methylation changes in the promoter region of many genes such as SHP-1, ID4 and p16INK4a in the psoriatic skin and peripheral blood mononuclear cells (PBMCs), consequently affect the expression of mRNA of the genes.23, 24, 25, 26 It has been shown that in patients with psoriasis, 5-methylcytosine levels in the epidermis and PBMCs were significantly increased, whereas global histone H4 hypoacetylation was found only in PBMCs.27, 28 In addition, it has been reported that changes in the genome-wide CpG methylation in the skin of patients with psoriasis was correlated with anti-tumor necrotic factor- alpha therapy.29 Furthermore, genome-wide DNA methylation was associated with epigenetic dysregulation among monozygotic twins discordant for psoriasis.30 As there is no information on the methylation status of LINE-1 and Alu in psoriasis, this study examined and compared the DNA methylation levels and patterns of LINE-1 and Alu in keratinocytes and various other hematopoietic cells from patients with psoriasis, squamous cell carcinoma (SCC) and healthy controls by using combined bisulfite-restriction analysis (COBRA). Methylation patterns of LINE-1 were classified as either hypermethylated, hypomethylated or two partially methylated loci (mCmC, uCuC, mCuC and uCmC, respectively). The severity of psoriasis was assessed by examining the levels and patterns of LINE-1. The methylation status of LINE-1 at different loci was also studied. Genes containing LINE-1 were analyzed by the CU-DREAMX program.
Materials and methods
Patients and healthy controls
Twenty-nine chronic plaque-type psoriasis patients (16 males, 13 females), 8 SCC patients (3 males, 5 females) from the King Chulalongkorn Memorial Hospital and 35 normal, healthy subjects (8 males, 27 females) were enrolled in to the study. Patients with psoriatic arthritis or other autoimmune diseases were excluded from the study. All patients were free of systemic skin therapies for at least 4 weeks or topical skin therapies for at least 2 weeks prior to sample collection. For each psoriatic patient, the severity of the disease was classified according to the affected area and severity index (PASI) (<15=mild and ≥15=severe). Patients undergoing elective plastic surgery were recruited in to the study as normal controls. The study was approved by the ethical committee of the Institutional Review Board of the Faculty of Medicine, Chulalongkorn University. All participants provided informed consent. The demographic data of the patients are shown in Table 1.
Cell isolation
Paraffin-embedded epidermal skin from 13 psoriatic patients, 8 SCCs and 15 normal controls were isolated by using the PALM MicroLaser Microdissection System (P.A.L.M. MicroLaser Technologies AG, Burnried, Germany). The dissected epidermal skin was removed from the slide by using a cutter pulse and collected in to a microtube. PBMCs were isolated from heparinized venous blood by Ficoll-Hypaque (Amersham-Pharmacia Biotech, Uppsala, Sweden), a gradient centrifugation technique used to separate lymphocytes from the blood. Various cell populations (B cell, T cell and non-T non-B cell) were isolated from the PBMCs by using antibody-coated magnetic beads (Dynal Biotech, ASA, Oslo, Norway). The purity of each cell type was verified by flow cytometry.
DNA preparation and bisulfite modification
Heparinized blood samples were collected. All white blood cells and keratinocytes were isolated as previously described.16, 26 DNA from all samples were obtained by using QIAamp DNA mini kit (QIAGEN, Hilden, Germany). After that, 500 ng of DNA was treated by bisulfite using the EZ DNA methylation kit (Zymo Research, Orange, CA, USA) according to the manufacturer’s specifications. The bisulfite-treated DNA samples were stored at −20 °C until further analysis.
COBRA of LINE-1 and Alu
Levels of DNA methylation of LINE-1 and Alu were assessed by using COBRA as previously described.9, 16, 22, 31 For LINE-1 COBRA, 2 μl of modified DNA was amplified by using Hot Start PCR. Hot Start PCR was carried out at 95 °C for 15 min and later followed by 35 cycles of 95 °C for 1 min, 50 °C for 1 min, and 72 °C for 1 min. Final extension was carried out at 72 °C for 7 min. For Alu COBRA, 2 μl of modified DNA was amplified by using Hot Start PCR as follows: hot start at 95 °C for 5 min followed by 35 cycles of 95 °C for 45 s, 60 °C for 45 s, and 72 °C for 45 s, and final extension at 72 °C for 7 min. After amplification, 8 μl of the PCR products (160 bp for LINE-1 and 99 bp for Alu) were digested with the restriction enzymes TaqI and TasI (MBI Fermentas, Gen Burnie, MD, USA). Each reaction was incubated overnight at 65 °C and then run on 8% polyacrylamide gels to separate the DNA fragments. The gel was then stained with the SYBR green nucleic-acid stain. The intensity of the DNA fragments was measured by PhosphoImagerl using Image Quant software (Molecular Dynamics, GE Healthcare, Slough, UK). For interassay-variation normalization, DNA templates from HeLa, Jurkat and Daudi cell lines were used as positive controls in all of the experiments. As for Alu, the PCR products were digested with restriction enzymes TaqI and TasI (MBI Fermentas).
LINE-1and Alu methylation analysis
The status of the methylation at 2 CpG dinucleotides in the 5’ and 3’ of the sequence for LINE-1 loci was identified by COBRA and categorized into four groups. These four groups are: (i) unmethylation of both CpGs (uCuC) at LINE-1 loci; (ii) methylation of both CpGs (mCmC) at LINE-1 loci; (iii) partial methylation of CpGs (5’-methylated and 3’- unmethylated CpGs (mCuC)) at LINE-1 loci; and (iv) partial methylation of CpGs (5’-unmethylated and 3’-methylated CpGs (uCmC)) at LINE-1 loci. LINE-1 methylation levels and percentages of LINE-1 loci from each group were calculated from the percentages of the intensities obtained from the COBRA-digested LINE-1 products. DNA fragments from enzymatic digestion for COBRA LINE-1 were separated into five fragments: 160, 98, 80, 62 and 18 bp. The 18-bp fragment was not used in the following calculations. The 160-bp fragment was identified to be methylated pattern of mCuC. The 98-bp fragment was identified to be methylated pattern of uCuC was, whereas the 80-bp fragment was the mixture of mCmC and uCmC. Eventually, the 62-bp fragment was the mixture of uCuC and uCmC. The number of CpG dinucleotides was obtained by dividing each band intensity from the corresponding size of the double-stranded DNA fragment as follows: A=160 bp fragment intensity/160; B=98 bp fragment intensity/94; C=80 bp fragment intensity/79; and D=62 bp fragment intensity/62. The LINE-1 methylation levels were calculated using the number of CpG dinucleotides according to the following formulas: LINE-1 methylation level percentage=100 × (C+A)/(C+A+A+B+D); %mCuC)=100 × (A)/(((C−D+B)/2)+A+D); %uCmC=100 × (D−B)/((C−D+B)/2)+A+D; %uCuC=100 × B/(((C−D+B)/2)+A+D); %mCmC=100 × ((C−D+B)/2)/(((C−D+B)/2)+D+A)22 (Figure 1).
A schematic representation of combined bisulfite restriction analysis of LINE-1(COBRA-LINE-1). After DNA preparation and bisulfite modification, the PCR products were digested by Taq1 and Tas1 restriction enzymes to 160-, 98-, 62- and 18-bp fragments. The LINE-1 methylation levels were calculated as a percentage of the intensity of the digested methylation fragment divided by the sum of the undigested and digested fragment.
The Alu methylated and unmethylated bands were 57 and 78 bp, respectively. The percentages of the intensity of the Alu-methylated band were represented as the percentages of the methylation of Alu.
CU-CREAMX analysis
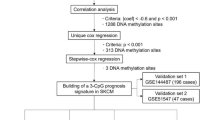
All LINE-1 data were extracted from the L1base (http://l1base.molgen.mpg.de).32 All microarray data on psoriasis were extracted from GEO dataset (http://www.ncbi.nlm.nih.gov/geo). For this study, the psoriasis GEO expression libraries used to assess the correlations between LINE-1 and psoriasis were GSE1335533 and GSE14905.34 The CU-DREAMX program, available at http://pioneer.netserv.chula.ac.th/~achatcha/cu-dream/, was used to measure this correlation. Any regulated gene with a P-value of <0.01 was considered significant.3
Statistical analysis
The methylation status was compared between groups by independent sample T-test (sig two-tailed) using the SPSS software for Windows 15.0 (SPSS Inc., Chicago, IL, USA). A P-value of<0.05 was considered to be significant. A receiver-operating characteristic (ROC) curve analysis was performed to assess whether LINE-1 and its patterns could distinguish patients with mild or severe psoriasis from the normal controls.
Results
Alterations in DNA methylation levels and patterns of LINE-1 in the psoriatic epidermis
From the COBRA-LINE-1 analysis, different levels of LINE-1 methylation and patterns of the epidermis from patients with psoriasis (n=13), SCC (n=8) and normal controls (n=15) were identified (Table 2). The levels of LINE-1 methylation in psoriatic (41.64%) and SCC (41.09%) epidermis were significantly lower than the normal epidermis (46.05%) (P=0.014 and P=0.010, respectively) (Table 2). As for the LINE-1 methylation pattern, the number of hypomethylated loci (uCuC) of LINE-1 in psoriasis (41.11%) was significantly higher than the normal controls (36.24%) (P=0.020). Unlike the epidermis, there were no significant differences between the patients with psoriasis and normal controls for LINE-1 methylation levels and patterns in various hematopoietic cell types (B, CD4+, CD8+ T lymphocytes and non-T, B cells) (Table 2).
As for the Alu analysis, there were no significant differences in the Alu methylation of the epidermis from patients with psoriasis (n=10), SCC (n=8) and normal controls (n=10). However, the Alu methylation levels in psoriatic non-T and B cells (30.26%) were lower compared with the normal controls (32.02%) (P=0.022) (Table 3).
LINE-1 methylation pattern as a biomarker for psoriasis
The ROC curve analysis was performed to explore the potential use of LINE-1 methylation pattern to distinguish psoriasis from the normal controls. The uCuC pattern was found to accurately detect psoriasis with a cutoff value of >39.67% (sensitivity 69.23 and specificity 73.33). The associations of LINE-1 methylation patterns and the different degree of severity of the disease are shown in Table 4. The degree of severity of psoriasis was classified as mild and severe when the PASI was <15 and ⩾15, respectively. Patterns with uCmC (severe vs mild=9.10 vs 18.8; P=0.022) and partial methylations (uCmC and mCuC) (29.91 vs 38.43; P=0.043) were lower in patients with severe psoriasis when compared with those with mild psoriasis. Next, the ROC curve analysis was utilized to explore the role of uCmC as a biomarker for the severity of psoriasis. The cutoff value for uCmC was ⩽10.87% which indicated that this pattern had a high specificity and sensitivity in detecting patients with severe psoriasis (sensitivity 83.33%; specificity 100.00%) (Figure 2).
ROC curve analysis of LINE-1 methylation in psoriasis. (a) The presence of LINE-1 methylation pattern uCuC was assessed in the keratinocytes from the patients with psoriasis and normal controls. The uCuC pattern was found to accurately detect psoriasis with a cutoff value of >39.67% (sensitivity 69.23 and specificity 73.33). (b) The LINE-1 methylation pattern uCmC was found to accurately detect severe psoriasis with a cutoff value of⩽10.87% (sensitivity=83.33% and specificity=100.00).
Downregulation of genes containing LINE-1 in psoriasis
Gene expression microarrays for psoriasis (GSE13355 and GSE14905) were examined for the presence of LINE-1 and its expression level by the CU-DREAMX program. Gene containing LINE-1s were categorized into four groups based on the location and possible controlling function of intragenic LINE-1 as 1454 of genes containing LINE-1, 336 of genes with sense LINE-1, 832 of genes with antisense LINE-1 and 286 of genes with both sense and antisense LINE-1. The χ2 test was used to determine the correlation of the genes containing intragenic LINE-1s and their expression in the skin from patient with psoriasis (Supplementary Table 1). The genes with LINE-1 from both arrays were downregulated (GSE13355, P=3.84 × 10−27, odd ratio=1.83 and GSE14905, P=2.14 × 10−21 odd ratio=1.79) (Table 5). Interestingly, the genes containing antisense LINE-1s were significantly more predominant in the downregulated genes in skin from patient with psoriasis than healthy individuals (GSE13355; P=7.25E-15, odd ratio=1.76 and GSE14905; P=1.47E-13, odd ratio=1.79) (Table 5). When these genes were further analyzed, it was discovered that these genes were involved in the pathogenesis of psoriasis. Interestingly, many of these genes were also known as tumor suppressor genes (Table 6).
Discussion
The genome-wide CpG hypomethylation, alterations of LINE-1 methylation and Alu hypomethylation with different patterns have been reported in various malignancies and cancers including SCC.3, 8, 10, 17, 18, 35, 36 In contrast to cancers, the genome-wide CpG hypermethylation of keratinocytes and PBMCs was reported in patients with psoriasis.27 Hence, the methylation data from cancer is not applicable for patients with psoriasis. This is further supported by the fact that the genomes of patients with psoriasis are more stable and have a different pathogenesis as those with cancer. For example, in psoriasis, inflammation and immunological factors play an important role in keratinocyte hyperproliferation and differentiation, whereas in cancer, this is not the case.
To date, there is no information on LINE-1 and Alu in patients with psoriasis. This study is the first of its kind to assess the methylation levels and patterns of LINE-1 and Alu in patients with psoriasis. This study did not detect any Alu methylations in the keratinocytes of the patients with psoriasis and therefore has concentrated on LINE-1 thereafter. Hypomethylation of LINE-1 was found in both the epidermal skin of patients with psoriasis and SCC. This discrepancy from the previous report may be due to the different technique used to assess methylation as well as the target DNA used. The use of global methylation may include other irrelevant genes so in this study, specific areas that were methylated were analyzed to be more accurate. By analyzing LINE-1, this study was able to assess in detail the different patterns and levels among various cell lines.
Further analysis of LINE-1 confirmed that there were different patterns on each locus studied. Previous studies have shown that patterns for methylations such as uCuC are far superior in detecting cancer than overall LINE-1 methylation.22 Similarly, this study also found that hypomethylated loci (uCuC) were associated with psoriasis. The presence of uCuC was significantly higher in patients with psoriasis than the normal controls and its ROC curve confirmed its use as a biomarker in accurately detecting psoriasis. The presence of other methylation patterns, uCmC and partial methylation, were significantly lower in patients with severe psoriasis. However, it should be noted that uCmC was the most specific and had the highest sensitivity in detecting patients with severe psoriasis compared with any other LINE-1 methylation patterns. Even the ROC curve analysis confirmed the accurate use of uCmC as a biomarker for severe psoriasis with a cutoff value of ⩽10.87% (sensitivity 83.33%; specificity 100.00%).
As for the results from the bioinformatics analysis, there were high prevalences of downregulated genes containing LINE-1 in patients with psoriasis. Most of the downregulated genes containing LINE-1 are involved in the biological processes that trigger psoriasis and the pathogenesis of the disease including tumor suppression, proliferation, apoptosis, cell differentiation and immune regulation (Table 6). This finding corroborates with the cancer data that hypermethylation of the promoter is frequently linked with transcriptional silencing of the tumor suppressor genes 19, 20, 21. It has been shown that downregulation of tumor suppressor genes in psoriatic epidermis will result in abnormal proliferation and differentiation of keratinocytes.23, 25, 26 It is possible that hypomethylation of LINE-1 downregulated tumor suppressor genes will cause acanthosis and hyperkeratosis in the epidermis of patients with psoriasis. Moreover, hypomethylation of LINE-1 may affect other genes involved in the induction of the disease and maintenance such as cell proliferation, apoptosis, differentiation and immune regulation.
Aside from that, hypomethylation of LINE-1 was associated with the downregulation of the genes containing LINE-1 in patients with psoriasis. It is possible that hypomethylation of intragenic-LINE-1 can repress the psoriatic gene as seen in patients with cancer.3 However, the mechanisms of LINE-1 hypomethylation and downregulation of the psoriatic genes are unclear. Additional studies are warranted to assess the functions of LINE-1. In conclusion, LINE-1 hypomethylation was found in patients with psoriasis and the genes containing LINE-1 are significantly downregulated in the epidermis. The LINE-1 methylation pattern uCuC was associated with psoriasis. The LINE-1 methylation pattern uCmC has a high sensitivity and specificity to detect severe psoriasis and therefore can be used as a biomarker for the disease.
References
Cordaux, R. & Batzer, M. A. The impact of retrotransposons on human genome evolution. Nat. Rev. Genet. 10, 691–703 (2009).
Kazazian, H. H. Jr & Moran, J. V. The impact of L1 retrotransposons on the human genome. Nat. Genet. 19, 19–24 (1998).
Aporntewan, C., Phokaew, C., Piriyapongsa, J., Ngamphiw, C., Ittiwut, C. & Tongsima, S. et al. Hypomethylation of intragenic LINE-1 represses transcription in cancer cells through AGO2. PLoS One 6, e17934 (2011).
Han, J. S., Szak, S. T. & Boeke, J. D. Transcriptional disruption by the L1 retrotransposon and implications for mammalian transcriptomes. Nature 429, 268–274 (2004).
Chisholm, K. M., Aubert, S. D., Freese, K. P., Zakian, V. A., King, M. C. & Welcsh, P. L. A genomewide screen for suppressors of Alu-mediated rearrangements reveals a role for PIF1. PLoS One 7, e30748 (2012).
Kitkumthorn, N. & Mutirangura, A. Long interspersed nuclear element-1 hypomethylation in cancer: biology and clinical applications. Clin. Epigenetics 2, 315–330 (2011).
Speek, M. Antisense promoter of human L1 retrotransposon drives transcription of adjacent cellular genes. Mol. Cell Biol. 21, 1973–1985 (2001).
Phokaew, C., Kowudtitham, S., Subbalekha, K., Shuangshoti, S. & Mutirangura, A. LINE-1 methylation patterns of different loci in normal and cancerous cells. Nucleic Acids Res. 36, 5704–5712 (2008).
Chalitchagorn, K., Shuangshoti, S., Hourpai, N., Kongruttanachok, N., Tangkijvanich, P. & Thong-ngam, D. et al. Distinctive pattern of LINE-1 methylation level in normal tissues and the association with carcinogenesis. Oncogene 23, 8841–8846 (2004).
Estecio, M. R., Gharibyan, V., Shen, L., Ibrahim, A. E., Doshi, K. & He, R. et al. LINE-1 hypomethylation in cancer is highly variable and inversely correlated with microsatellite instability. PLoS One 2, e399 (2007).
Perrin, D., Ballestar, E., Fraga, M. F., Frappart, L., Esteller, M. & Guerin, J. F. et al. Specific hypermethylation of LINE-1 elements during abnormal overgrowth and differentiation of human placenta. Oncogene 26, 2518–2524 (2007).
Shuangshoti, S., Hourpai, N., Pumsuk, U. & Mutirangura, A. Line-1 hypomethylation in multistage carcinogenesis of the uterine cervix. Asian Pac. J. Cancer Prev. 8, 307–309 (2007).
Pattamadilok, J., Huapai, N., Rattanatanyong, P., Vasurattana, A., Triratanachat, S. & Tresukosol, D. et al. LINE-1 hypomethylation level as a potential prognostic factor for epithelial ovarian cancer. Int. J. Gynecol. Cancer 18, 711–717 (2008).
Moore, L. E., Pfeiffer, R. M., Poscablo, C., Real, F. X., Kogevinas, M. & Silverman, D. et al. Genomic DNA hypomethylation as a biomarker for bladder cancer susceptibility in the Spanish Bladder Cancer Study: a case-control study. Lancet Oncol. 9, 359–366 (2008).
Karouzakis, E., Gay, R. E., Michel, B. A., Gay, S. & Neidhart, M. DNA hypomethylation in rheumatoid arthritis synovial fibroblasts. Arthritis Rheum. 60, 3613–3622 (2009).
Nakkuntod, J., Avihingsanon, Y., Mutirangura, A. & Hirankarn, N. Hypomethylation of LINE-1 but not Alu in lymphocyte subsets of systemic lupus erythematosus patients. Clin. Chim. Acta. 412, 1457–1461 (2011).
Lee, H. S., Kim, B. H., Cho, N. Y., Yoo, E. J., Choi, M. & Shin, S. H. et al. Prognostic implications of and relationship between CpG island hypermethylation and repetitive DNA hypomethylation in hepatocellular carcinoma. Clin. Cancer Res. 15, 812–820 (2009).
Bae, J. M., Shin, S. H., Kwon, H. J., Park, S. Y., Kook, M. C. & Kim, Y. W. et al. ALU and LINE-1 hypomethylations in multistep gastric carcinogenesis and their prognostic implications. Int. J. Cancer 131, 1323–1331 (2011).
Ehrlich, M. DNA methylation in cancer: too much, but also too little. Oncogene 21, 5400–5413 (2002).
Frigola, J., Sole, X., Paz, M. F., Moreno, V., Esteller, M. & Capella, G. et al. Differential DNA hypermethylation and hypomethylation signatures in colorectal cancer. Hum. Mol. Genet. 14, 319–326 (2005).
Saito, K., Kawakami, K., Matsumoto, I., Oda, M., Watanabe, G. & Minamoto, T. Long interspersed nuclear element 1 hypomethylation is a marker of poor prognosis in stage IA non-small cell lung cancer. Clin. Cancer Res. 16, 2418–2426 (2010).
Kitkumthorn, N., Tuangsintanakul, T., Rattanatanyong, P., Tiwawech, D. & Mutirangura, A. LINE-1 methylation in the peripheral blood mononuclear cells of cancer patients. Clin. Chim. Acta. 413, 869–874 (2012).
Chen, M., Chen, Z. Q., Cui, P. G., Yao, X., Li, Y. M. & Li, A. S. et al. The methylation pattern of p16INK4a gene promoter in psoriatic epidermis and its clinical significance. Br. J. Dermatol. 158, 987–993 (2008).
Chen, M., Cui, P. G., Yao, X., Cao, Y. H., Gong, J. Q. & Li, A. S. et al. [The methylation locus and frequency pattern on p16 INK4a gene promoter CpG in epidermis of patients with psoriasis]. Zhonghua Yi Xue Yi Chuan Xue Za Zhi 24, 674–676 (2007).
Ruchusatsawat, K., Wongpiyabovorn, J., Shuangshoti, S., Hirankarn, N. & Mutirangura, A. SHP-1 promoter 2 methylation in normal epithelial tissues and demethylation in psoriasis. J. Mol. Med. 84, 175–182 (2006).
Ruchusatsawat, K., Wongpiyabovorn, J., Protjaroen, P., Chaipipat, M., Shuangshoti, S. & Thorner, P. S. et al. Parakeratosis in skin is associated with loss of inhibitor of differentiation 4 via promoter methylation. Hum. Pathol. 42, 1878–1887 (2011).
Zhang, P., Su, Y., Chen, H., Zhao, M. & Lu, Q. Abnormal DNA methylation in skin lesions and PBMCs of patients with psoriasis vulgaris. J. Dermatol. Sci. 60, 40–42 (2010).
Zhang, P., Su, Y., Zhao, M., Huang, W. & Lu, Q. Abnormal histone modifications in PBMCs from patients with psoriasis vulgaris. Eur. J. Dermatol. 21, 552–557 (2011).
Roberson, E. D., Liu, Y., Ryan, C., Joyce, C. E., Duan, S. & Cao, L. et al. A subset of methylated CpG sites differentiate psoriatic from normal skin. J. Invest. Dermatol. 132, 583–592 (2012).
Gervin, K., Vigeland, M. D., Mattingsdal, M., Hammero, M., Nygard, H. & Olsen, A. O. et al. DNA methylation and gene expression changes in monozygotic twins discordant for psoriasis: identification of epigenetically dysregulated genes. PLoS Genet. 8, e1002454 (2012).
Jintaridth, P. & Mutirangura, A. Distinctive patterns of age-dependent hypomethylation in interspersed repetitive sequences. Physiol. Genomics 41, 194–200 (2010).
Penzkofer, T., Dandekar, T. & Zemojtel, T. L1Base: from functional annotation to prediction of active LINE-1 elements. Nucleic Acids Res. 33, D498–D500 (2005).
Nair, R. P., Duffin, K. C., Helms, C., Ding, J., Stuart, P. E. & Goldgar, D. et al. Genome-wide scan reveals association of psoriasis with IL-23 and NF-kappaB pathways. Nat. Genet. 41, 199–204 (2009).
Yao, Y., Richman, L., Morehouse, C., de los Reyes, M., Higgs, B. W. & Boutrin, A. et al. Type I interferon: potential therapeutic target for psoriasis? PLoS One 3, e2737 (2008).
Sunami, E., de Maat, M., Vu, A., Turner, R. R. & Hoon, D. S. LINE-1 hypomethylation during primary colon cancer progression. PLoS One 6, e18884 (2011).
Cash, H. L., Tao, L., Yuan, J. M., Marsit, C. J., Houseman, E. A. & Xiang, Y. B. et al. LINE-1 hypomethylation is associated with bladder cancer risk among nonsmoking Chinese. Int. J. Cancer 130, 1151–1159 (2011).
Acknowledgements
This work was supported by a research grant from the National Research University Project of CHE and the Ratchadaphiseksomphot Endowment Fund (HR1163A), Grant for Chula Research Scholar, Ratchadaphiseksomphot Endownment Fund, The 90th Year Anniversary of Chulalongkorn University (Ratchadaphiseksomphot Endowment Fund).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Yooyongsatit, S., Ruchusatsawat, K., Noppakun, N. et al. Patterns and functional roles of LINE-1 and Alu methylation in the keratinocyte from patients with psoriasis vulgaris. J Hum Genet 60, 349–355 (2015). https://doi.org/10.1038/jhg.2015.33
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2015.33
This article is cited by
-
The critical importance of epigenetics in autoimmune-related skin diseases
Frontiers of Medicine (2023)
-
Epigenetics in Non-tumor Immune-Mediated Skin Diseases
Molecular Diagnosis & Therapy (2021)
-
Epigenetics in psoriasis: perspective of DNA methylation
Molecular Genetics and Genomics (2021)
-
DNA methylation and inflammatory skin diseases
Archives of Dermatological Research (2020)
-
Epigenome-wide DNA methylation regulates cardinal pathological features of psoriasis
Clinical Epigenetics (2018)